



Biodiv Sci ›› 2014, Vol. 22 ›› Issue (4): 508-515. DOI: 10.3724/SP.J.1003.2014.14053 cstr: 32101.14.SP.J.1003.2014.14053
Special Issue: 土壤生物与土壤健康
• Original Papers • Previous Articles Next Articles
Jingbo Kan1, Lina Li2, Dong Qu2,*( ), Baoli Wang1,*(
), Baoli Wang1,*( )
)
Received:2014-03-14
Accepted:2014-05-16
Online:2014-07-20
Published:2014-07-24
Contact:
Qu Dong,Wang Baoli
Jingbo Kan, Lina Li, Dong Qu, Baoli Wang. Changes in bacterial abundance and community structure associated with flooding in paddy soil[J]. Biodiv Sci, 2014, 22(4): 508-515.
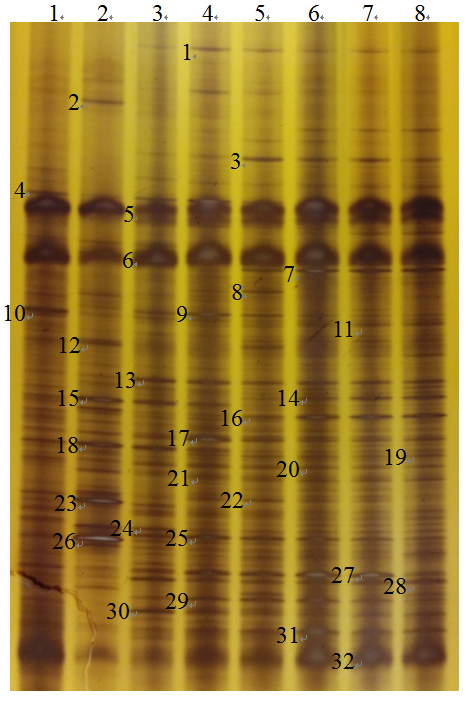
Fig. 2 DGGE profiles of bacterial community in different flooding time of paddy soil. Lane 1-8 showed the samples of flooding time of 1 h, 1 d, 5 d, 10 d, 20 d, 30 d, 40 d and 60 d, respectively.
| 淹水时间 Flooding time | 丰富度指数 Margalef index (dMa) | Shannon指数 Shannon-Wiener index (H') | 均匀度指数 Evenness index (E) | Simpson指数 Simpson index (Ds) |
|---|---|---|---|---|
| 1 h | 3.954 | 2.991 | 0.941 | 0.948 |
| 1 d | 3.655 | 2.962 | 0.932 | 0.945 |
| 5 d | 5.102 | 3.304 | 0.945 | 0.965 |
| 10 d | 5.371 | 3.368 | 0.955 | 0.965 |
| 20 d | 5.316 | 3.379 | 0.950 | 0.966 |
| 30 d | 5.657 | 3.382 | 0.959 | 0.965 |
| 40 d | 5.328 | 3.261 | 0.917 | 0.963 |
| 60 d | 4.914 | 3.228 | 0.931 | 0.963 |
Table 1 Diversity indices of bacterial community in different flooding periods of a paddy soil
| 淹水时间 Flooding time | 丰富度指数 Margalef index (dMa) | Shannon指数 Shannon-Wiener index (H') | 均匀度指数 Evenness index (E) | Simpson指数 Simpson index (Ds) |
|---|---|---|---|---|
| 1 h | 3.954 | 2.991 | 0.941 | 0.948 |
| 1 d | 3.655 | 2.962 | 0.932 | 0.945 |
| 5 d | 5.102 | 3.304 | 0.945 | 0.965 |
| 10 d | 5.371 | 3.368 | 0.955 | 0.965 |
| 20 d | 5.316 | 3.379 | 0.950 | 0.966 |
| 30 d | 5.657 | 3.382 | 0.959 | 0.965 |
| 40 d | 5.328 | 3.261 | 0.917 | 0.963 |
| 60 d | 4.914 | 3.228 | 0.931 | 0.963 |
| 淹水时间 Flooding time | 1 h | 1 d | 5 d | 10 d | 20 d | 30 d | 40 d | 60 d |
|---|---|---|---|---|---|---|---|---|
| 1 h | 100 | |||||||
| 1 d | 79.167 | 100 | ||||||
| 5 d | 70.175 | 70.175 | 100 | |||||
| 10 d | 67.857 | 62.069 | 92.537 | 100 | ||||
| 20 d | 61.017 | 57.627 | 85.294 | 89.855 | 100 | |||
| 30 d | 62.069 | 58.621 | 86.567 | 91.177 | 95.652 | 100 | ||
| 40 d | 61.017 | 57.627 | 85.294 | 86.957 | 91.429 | 95.652 | 100 | |
| 60 d | 57.143 | 57.143 | 83.087 | 84.849 | 86.567 | 90.909 | 95.522 | 100 |
Table 2 Similarity index (%) of bacterial community in different flooding periods of a paddy soil
| 淹水时间 Flooding time | 1 h | 1 d | 5 d | 10 d | 20 d | 30 d | 40 d | 60 d |
|---|---|---|---|---|---|---|---|---|
| 1 h | 100 | |||||||
| 1 d | 79.167 | 100 | ||||||
| 5 d | 70.175 | 70.175 | 100 | |||||
| 10 d | 67.857 | 62.069 | 92.537 | 100 | ||||
| 20 d | 61.017 | 57.627 | 85.294 | 89.855 | 100 | |||
| 30 d | 62.069 | 58.621 | 86.567 | 91.177 | 95.652 | 100 | ||
| 40 d | 61.017 | 57.627 | 85.294 | 86.957 | 91.429 | 95.652 | 100 | |
| 60 d | 57.143 | 57.143 | 83.087 | 84.849 | 86.567 | 90.909 | 95.522 | 100 |
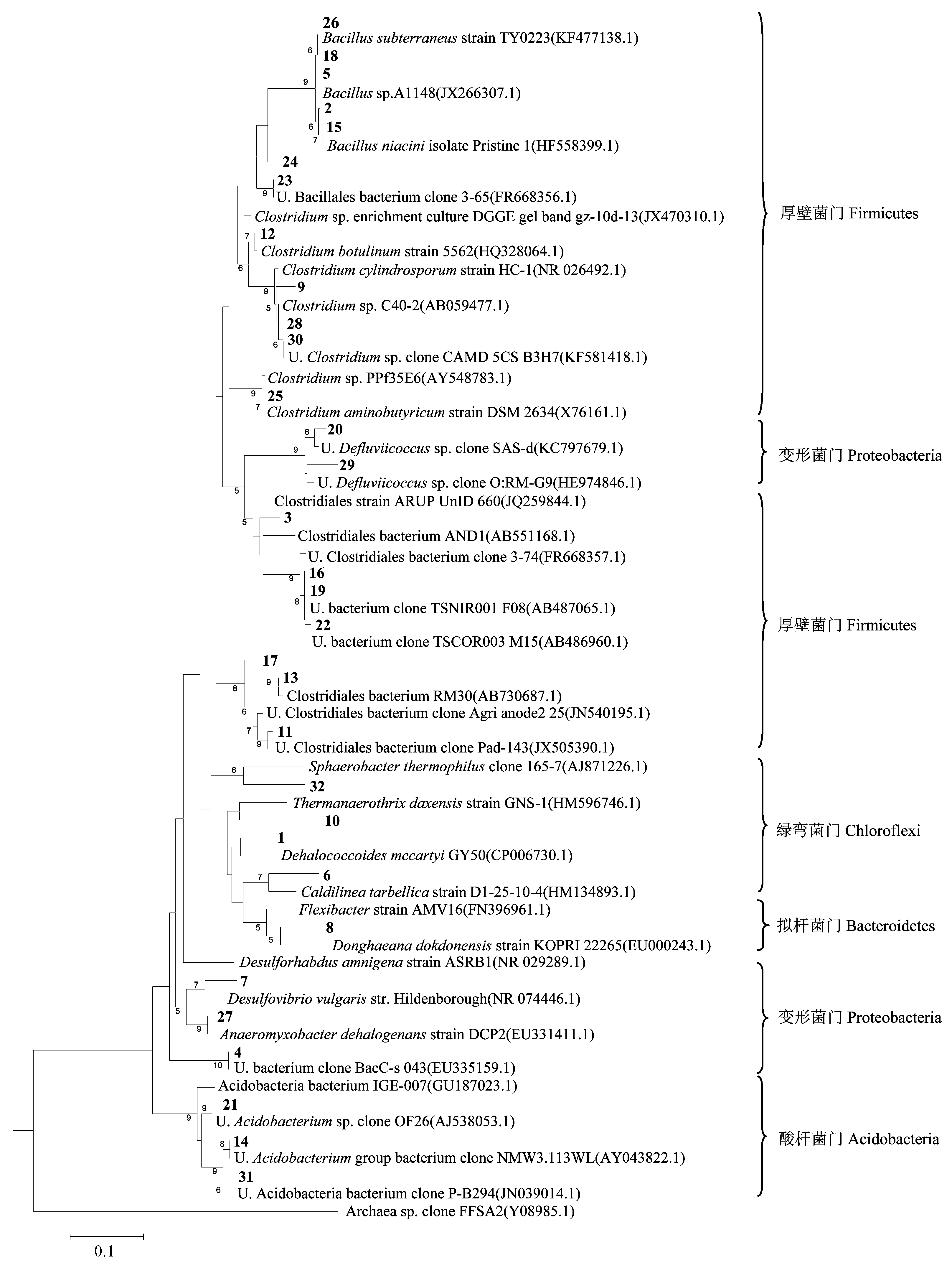
Fig. 4 Phylogenetic tree constructed with neighbor-joining methods based on 16S rRNA V3 gene sequences. Boldface type indicates bands number obtained in this study. The scale bar represents 10% estimated sequence divergence. U. stands for uncultured.
| [1] | Cerli C, Liu Q, Hanke A, Kaiser K, Kalbitz K (2013) Lignin decomposition and microbial community in paddy soils: effects of alternating redox conditions. In: General Assembly Conference Abstracts (ed. Reichart GJ), Vol.15, p 13896. European Geosciences Union, Vienna. |
| [2] | Charlesworth B (1971) Selection in density-regulated populations.Ecology, 52, 469-474. |
| [3] | Chen X, Zhang LM, Shen JP, Xu Z, He JZ (2010) Soil type determines the abundance and community structure of ammonia-oxidizing bacteria and archaea in flooded paddy soils.Journal of Soils and Sediments, 10, 1510-1516. |
| [4] | Eichorst SA, Breznak JA, Schmidt TM (2007) Isolation and characterization of soil bacteria that define Terriglobus gen. nov., in the phylum Acidobacteria.Applied and Environmental Microbiology, 73, 2708-2717. |
| [5] | Goecks J, Nekrutenko A, Taylor J (2010) Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences.Genome Biology, 11, R86. |
| [6] | Gordon H, Haygarth PM, Bardgett RD (2008) Drying and rewetting effects on soil microbial community composition and nutrient leaching.Soil Biology and Biochemistry, 40, 302-311. |
| [7] | Higashioka Y, Kojima H, Watanabe M, Fukui M (2013) Desulfatitalea tepidiphila gen. nov., sp. nov., a sulfate-reducing bacterium isolated from tidal flat sediment.International Journal of Systematic and Evolutionary Microbiology, 63, 761-765. |
| [8] | Hill TC, Walsh KA, Harris JA (2003) Using ecological diversity measures with bacterial communities.FEMS Microbiol Ecol, 43, 1-11 |
| [9] | Ji B, Gimenez G, Barbe V, Vacherie B, Rouy Z, Amrani A, Fardeau ML, Bertin P, Alazard D, Leroy S, Talla E, Ollivier B, Dolla A, Pradel N (2013) Complete genome sequence of the piezophilic, mesophilic, sulfate-reducing bacterium Desulfovibrio hydrothermalis AM13T.Genome Announcements, 1(1), e00226. |
| [10] | Katoh M, Iwata A, Shaku I, Nakajima Y, Matsuya K, Kimura M (2003) Impact of water percolation on nutrient leaching from an irrigated paddy field in Japan.Soil Use and Management, 19, 298-304. |
| [11] | Kudo K, Yamaguchi N, Makino T, Ohtsuka T, Kimura K, Dong DT, Amachi S (2013) Release of arsenic from soil by a novel dissimilatory arsenate-reducing bacterium, Anaeromyxobacter sp. strain PSR-1.Applied and Environmental Microbiology, 79, 4635-4642. |
| [12] | Lee HJ, Kim SY, Kim PJ, Madsen EL, Jeon CO (2014) Methane emission and dynamics of methanotrophic and methanogenic communities in a flooded rice field ecosystem.FEMS Microbiology Ecology, 88, 195-212. |
| [13] | Li HJ, Peng JJ, Weber KA, Zhu YG (2011) Phylogenetic diversity of Fe (III)-reducing microorganisms in rice paddy soil: enrichment cultures with different short-chain fatty acids as electron donors.Journal of Soils and Sediments, 11, 1234-1242. |
| [14] | Liu L, Peng YK, Zheng XH, Xiao L, Yang LY (2010) Vertical structure of bacterial and archaeal communities within the sediment of a eutrophic lake as revealed by culture-independent methods.Journal of Freshwater Ecology, 25, 565-573. |
| [15] | Lüdemann H, Arth I, Liesack W (2000) Spatial changes in the bacterial community structure along a vertical oxygen gradient in flooded paddy soil cores.Applied and Environmental Microbiology, 66, 754-762. |
| [16] | Noll M, Matthies D, Frenzel P, Derakshani M, Liesack W (2005) Succession of bacterial community structure and diversity in a paddy soil oxygen gradient.Environmental Microbiology, 7, 382-395. |
| [17] | Rappé MS, Giovannoni SJ (2003) The uncultured microbial majority.Annual Reviews in Microbiology, 57, 369-394. |
| [18] | Reim A, Lüke C, Krause S, Pratscher J, Frenzel P (2012) One millimetre makes the difference: high-resolution analysis of methane-oxidizing bacteria and their specific activity at the oxic-anoxic interface in a flooded paddy soil.The ISME Journal, 6, 2128-2139. |
| [19] | Rizzo A, Boano F, Revelli R, Ridolfi L (2013) Role of water flow in modeling methane emissions from flooded paddy soils.Advances in Water Resources, 52, 261-274. |
| [20] | Sar P, Kazy SK, Paul D, Sarkar A (2013) Metal bioremediation by thermophilic microorganisms. In: Thermophilic Microbes in Environmental and Industrial Biotechnology(ed. Satyanarayana T, Littlechild J, Kawarabayasi Y), pp. 171-201. Springer, the Netherlands. |
| [21] | Shu W, Pablo GP, Jun Y, Danfeng H (2012). Abundance and diversity of nitrogen-fixing bacteria in rhizosphere and bulk paddy soil under different duration of organic management.World Journal of Microbiology and Biotechnology, 28, 493-503. |
| [22] | Silva AF, Carvalho G, Oehmen A, Lousada-Ferreira M, van Nieuwenhuijzen A, Reis MA, Crespo MTB (2012) Microbial population analysis of nutrient removal-related organisms in membrane bioreactors.Applied Microbiology and Biotechnology, 93, 2171-2180. |
| [23] | Sorokin DY, Lücker S, Vejmelkova D, Kostrikina NA, Kleerebezem R, Rijpstra WIC, Damsté JSS, Paslier DL, Muyzer G, Wagner M, Van Loosdrecht MCM, Daims H (2012) Nitrification expanded: discovery, physiology and genomics of a nitrite-oxidizing bacterium from the phylum Chloroflexi.The ISME Journal, 6, 2245-2256. |
| [24] | Tuo XH (拓晓骅), Zhu H (朱辉), Wang BL (王保莉), Qu D (曲东) (2012) Changing characteristics of Geobacteraceae community structure in paddy soil during flooding incubation.Journal of Agro-Environment Science(农业环境科学学报), 31, 1165-1171. (in Chinese with English abstract) |
| [25] | Wang BL (王保莉), Huang S (黄森), Liu H (刘浩), Qu D (曲东) (2013) Characteristics of Bacillus communities in flooded paddy soil.Journal of Agro-Environment Science(农业环境科学学报), 32, 1021-1027. (in Chinese with English abstract) |
| [26] | Wang Y (王英), Teng QH (滕齐辉), Cui ZL (崔中利), Sun B (孙波), Cao H (曹慧), Hu F (胡锋) (2007) Diversity and spatial distribution of bacteria in non-tillage paddy fields.Acta Pedologica Sinica(土壤学报), 44, 137-143. (in Chinese with English abstract) |
| [27] | Wu MN, Qin HL, Chen Z, Wu JS, Wei WX (2011) Effect of long-term fertilization on bacterial composition in rice paddy soil.Biology and Fertility of Soils, 47, 397-405. |
| [28] | Yi WJ, You JH, Zhu C, Wang BL, Qu D (2013) Diversity, dynamic and abundance of Geobacteraceae species in paddy soil following slurry incubation.European Journal of Soil Biology, 56, 11-18. |
| [29] | Zhou W, Kageyama K, Li F, Yuasa A (2007) Monitoring of microbiological water quality by real-time PCR.Environmental Technology, 28, 545-553. |
| [1] | Xiao-Qing Wu Meihui Zhang Suting Ge Manshu Li Kun Song Guochun Shen Jian Zhang. Spatiotemporal Dynamics of Woody Plant Species Diversity and Aboveground Biomass during Near-Natural Forest Reconstruction in Shanghai: A Case Study from the Eco-Island in Minhang District [J]. Biodiv Sci, 2025, 33(5): 24444-. |
| [2] | Mingyi Zhang, Xiaomei Wang, Yanxin Zheng, Nan Wu, Donghao Li, Enyuan Fan, Na Li, Xiujuan Shan, Tao Yu, Chunnuan Zhao, Bo Li, Shuai Xu, Yuping Wu, Liqun Ren. Resource status and habitat function of typical oyster reef areas in the Yellow River Estuary [J]. Biodiv Sci, 2025, 33(4): 24208-. |
| [3] | Tong Miao, Wang Huan, Zhang Wenshuang, Wang Chao, Song Jianxiao. Distribution characteristics of antibiotic resistance genes in soil bacterial communities exposed to heavy metal pollution [J]. Biodiv Sci, 2025, 33(3): 24101-. |
| [4] | Jia Zhenni, Zhang Yicen, Du Yanjun, Ren Haibao. Influences of disturbances on successional dynamics of species diversity in mid- subtropical forests [J]. Biodiv Sci, 2025, 33(2): 24078-. |
| [5] | Li Yanpeng, Chen Jie, Lu Chunyang, Xu Han. Community characteristics of a 64-ha secondary forest dynamics plot in a tropical montane rainforest in Jianfengling, Hainan [J]. Biodiv Sci, 2025, 33(2): 24445-. |
| [6] | Shiyu Wei, Tianjiao Song, Jiayi Luo, Yan Zhang, Zixuan Zhao, Jingwen Ru, Hua Yi, Yanbing Lin. Altitudinal distribution patterns of soil bacterial communities in the Huoditang coniferous forests of the Qinling Mountains [J]. Biodiv Sci, 2024, 32(9): 24180-. |
| [7] | Jing Chen, Bingchang Zhang, Yanjin Liu, Jie Wu, Kang Zhao, Jiao Ming. Diversity of Leptolyngbya-like cyanobacteria in biocrusts in desert area [J]. Biodiv Sci, 2024, 32(9): 24186-. |
| [8] | Xiang Gao, Shufang Pan, Zhengzheng Sun, Jixiao Li, Tianyu Gao, Lu Dong, Ning Wang. Distribution and activity rhythm of small Indian civet (Viverricula indica) in Fenghuang Hill and Qi’ao Island, Zhuhai, Guangdong [J]. Biodiv Sci, 2024, 32(8): 24045-. |
| [9] | Yongqiang Shi, Qingshan Luan, Xiujuan Shan, Chao Wei, Yongsong Zhao, Cece Sun, Xianshi Jin. Annual changes in zooplankton biodiversity in the southern waters of Changdao [J]. Biodiv Sci, 2024, 32(7): 23428-. |
| [10] | Yanpeng Li, Lijun Pan, Jie Chen, Han Xu, Lixin Yang. Response and influencing factors of leaf functional traits to forest succession in subtropical mixed plantations [J]. Biodiv Sci, 2024, 32(7): 24049-. |
| [11] | Teng Wang, Chunhou Li, Guanghua Wang, Jinfa Zhao, Juan Shi, Hongyu Xie, Yong Liu, Yu Liu. Species composition and succession of coral reef fishes on Qilianyu Island, Xisha Islands [J]. Biodiv Sci, 2024, 32(6): 23481-. |
| [12] | Weiqiang Xu, Qiang Su. Exploring the interplay of fractal model and species abundance distribution: A case study of shellfish and insect [J]. Biodiv Sci, 2024, 32(4): 23410-. |
| [13] | Yanmei Ni, Li Chen, Zhiyuan Dong, Debin Sun, Baoquan Li, Xumin Wang, Linlin Chen. Community structure of macrobenthos and ecological health evaluation in the restoration area of the Yellow River Delta wetland [J]. Biodiv Sci, 2024, 32(3): 23303-. |
| [14] | Yaoqi Chen, Jingjing Guo, Guojun Cai, Yili Ge, Yu Liao, Zheng Dong, Hui Fu. Evolution characteristics of submerged macrophyte community diversity in the middle and lower reaches of the Yangtze River in the past seventy years (1954-2021) [J]. Biodiv Sci, 2024, 32(3): 23319-. |
| [15] | Jiaxin Wei, Zhiguo Jiang, Linsen Yang, Huanhuan Xiong, Jiaojiao Jin, Fanglin Luo, Jiehua Li, Hao Wu, Yaozhan Xu, Xiujuan Qiao, Xinzeng Wei, Hui Yao, Huiliang Yu, Jingyuan Yang, Mingxi Jiang. Community composition and structure in a 25 ha mid-subtropical mountain deciduous broad-leaved forest dynamics plot in Shennongjia, Hubei, China [J]. Biodiv Sci, 2024, 32(3): 23338-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn