



生物多样性 ›› 2011, Vol. 19 ›› Issue (6): 770-778. DOI: 10.3724/SP.J.1003.2011.09149
所属专题: 中国的海洋生物多样性
收稿日期:2011-08-23
接受日期:2011-12-10
出版日期:2011-11-20
发布日期:2011-12-19
通讯作者:
阚金军
作者简介:*E-mail: jkan@stroudcenter.org基金资助:Received:2011-08-23
Accepted:2011-12-10
Online:2011-11-20
Published:2011-12-19
Contact:
Jinjun Kan
Supported by:摘要:
咸淡水的混合和重要营养盐与有机物的再循环, 使得河口成为地球上生产力较高而动态变化明显的水生生态系统。一个典型的河口区断面中, 细菌群落包含了一些从淡水到海洋的过渡类型: 例如α-变形菌(Alphaproteobacteria)、β-变形菌(Betaproteobacteria)、γ-变形菌(Gammaproteobacteria)、蓝细菌(Cyanobacteria)[聚球藻(Synechococcus)]、拟杆菌(Bacteroidetes)、放线细菌(Actinobacteria)和疣微菌(Verrucomicrobia)等。此外, 河口也包含其独特的细菌群落: SAR11组、玫瑰杆菌属(Roseobacter)、SAR86和放线细菌(Actinobacteria)的一些进化亚枝(subclades), 表明海湾或者大型温带河口区细菌类群具有区域生态适应性。以研究较多的美国切萨皮克湾(Chesapeake Bay)为例, 其细菌群落呈现出显著的季节性变化和周期性的年际变化特征; 这些变化除了受水的滞留时间和细菌生长速度影响外, 还可能受其他许多环境因子的影响。其中叶绿素a和水温变化的影响最大, 其他环境因子如溶解氧、铵态氮、亚硝酸盐和硝酸盐以及病毒的丰度也有影响。近年来, 基于群落水平的基因组学(genomics)和后基因组学(postgenomics)(转录组学和蛋白质组学)技术应用于研究自然条件下微生物群落错综复杂的基因多样性和表达, 提供了揭示水环境中微生物群落组成和新功能基因的途径。
阚金军, 孙军 (2011) 河口细菌群落多样性及其控制因素:以切萨皮克湾为例. 生物多样性, 19, 770-778. DOI: 10.3724/SP.J.1003.2011.09149.
Jinjun Kan, Jun Sun (2011) Bacterial community biodiversity in estuaries and its controlling factors: a case study in Chesapeake Bay. Biodiversity Science, 19, 770-778. DOI: 10.3724/SP.J.1003.2011.09149.
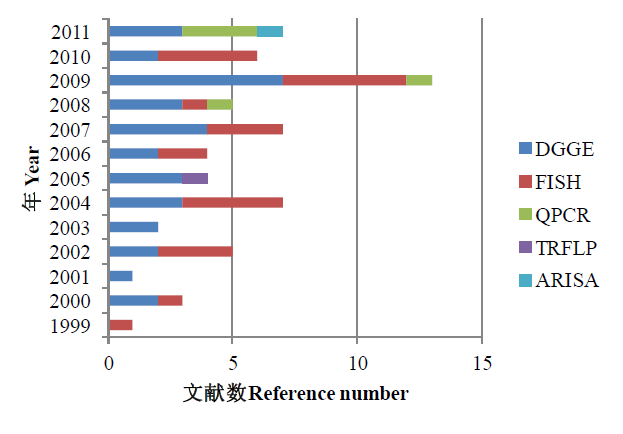
图1 近年来国际主要河口细菌群落的分子生物学检测。 DGGE: 变性梯度凝胶电泳; FISH: 荧光原位杂交; QPCR: 实时荧光定量核酸扩增检测系统; TRFLP: 末端限制性片段长度多样性; ARISA: 核糖体间隔基因自动分析。(检索时间2011年11月20日)
Fig. 1 The molecular techniques for detection diversity of estuary bacteria communities around the world in 1999-2011. DGGE, Denaturing gradient gel electrophoresis; FISH, Fluorescence in situ hybridization; QPCR, Real-time polymerase chain reaction; TRFLP, Terminal restriction fragment length polymorphism; ARISA, Automated ribosomal intergenic spacer analysis. (Searching on November 20th 2011)
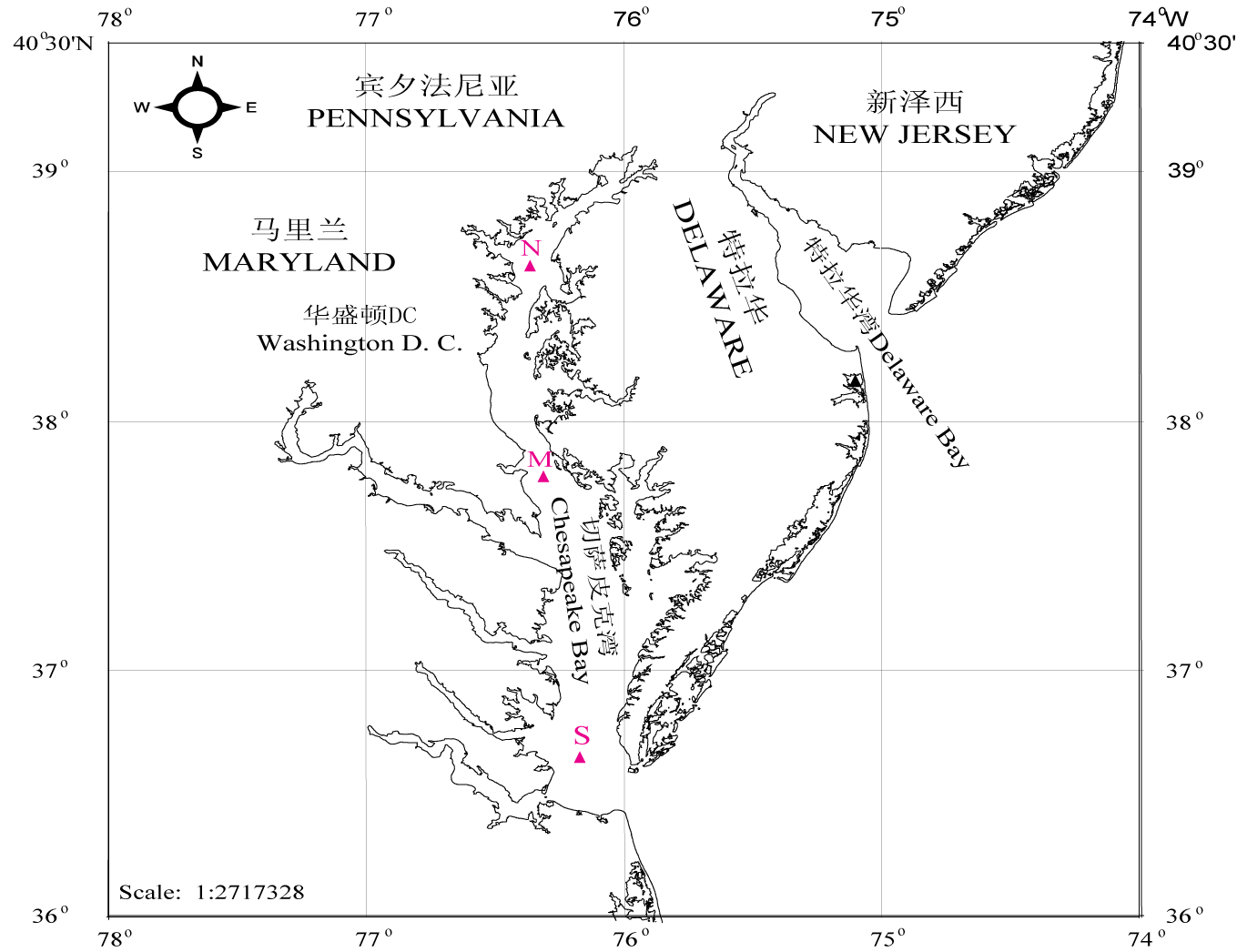
图2 切萨皮克湾采样站位。 N, 北部站; M, 中部站; S, 南部站。
Fig. 2 Map of the Chesapeake Bay showing sampling stations. N, M, and S represent north, middle and south Bay stations respectively.
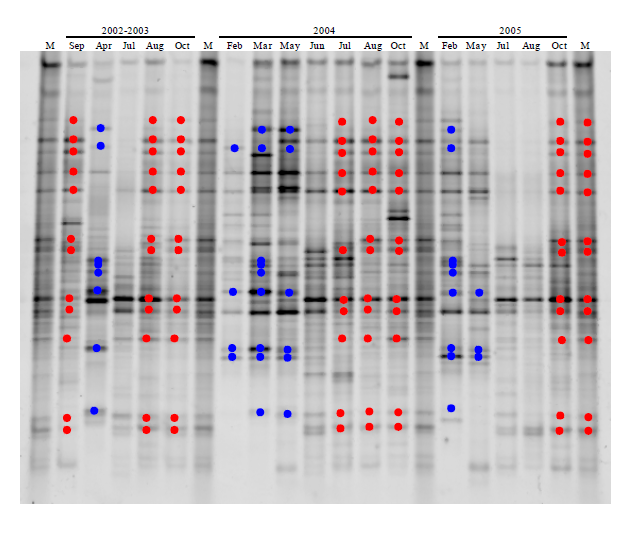

图4 2002-2005年切萨皮克湾中部细菌群落指纹图谱。 红色和蓝色分别表示夏季和冬季。M: 标记
Fig. 4 Fingerprints of Chesapeake Bay bacterial communities during 2002-2005 (middle Bay). Red and blue highlighted the bands appeared in summer and winter, respectively. M, marker.
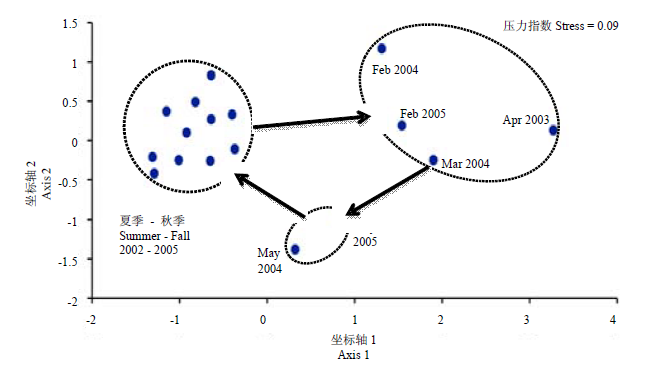
图5 切萨皮克湾细菌指纹图谱(自图4)的MDS分析
Fig. 5 Multidimensional scaling (MDS) analysis by SAS software package (SAS, 1992) of Chesapeake Bay bacterial DGGE fingerprints from Fig. 4
| [1] |
Amann RI, Krumholz L, Stahl DA (1990) Fluorescent-oligonucleotide probing of whole cells for determinative, phylogenetic, and environmental studies in microbiology. Journal of Bacteriology, 172, 762-770.
URL PMID |
| [2] |
Béjà O, Aravind L, Koonin EV, Suzuki MT, Hadd A, Nguyen LP, Jovanovich SB, Gates CM, Feldman RA, Spudich JL, Spudich EN, DeLong EF (2000a) Bacterial rhodopsin: evidence for a new type of phototrophy in the sea. Science, 289, 1902-1906.
URL PMID |
| [3] |
Béjà O, Suzuki MT, Koonin EV, Aravind L, Hadd A, Nguyen LP, Villacorta R, Amjadi M, Garrigues C, Jovanovich SB, Feldman RA, DeLong EF (2000b) Construction and analysis of bacterial artificial chromosome libraries from a marine microbial assemblage. Environmental Microbiology, 2, 516-529.
DOI URL PMID |
| [4] |
Béjà O, Spudich EN, Spudich JL, Leclerc M, DeLong EF (2001) Proteorhodopsin phototrophy in the ocean. Nature, 411, 786-789.
DOI URL PMID |
| [5] | Benson DA, Karsch-Mizrachi I, Lipman DJ, Ostell J, Sayers EW (2011) GenBank. Nucleic Acids Research, 39(Suppl. 1), D32-D37. |
| [6] | Bouvier TC, del Giorgio PA (2002) Compositional changes in free-living bacterial communities along a salinity gradient in two temperate estuaries. Limnology and Oceanography, 47, 453-470. |
| [7] |
Button DK, Schut F, Quang P, Martin R, Robertson BR (1993) Viability and isolation of marine bacteria by dilution culture: theory, procedures, and initial results. Applied and Environmental Microbiology, 59, 881-891.
URL PMID |
| [8] |
Cho JC, Giovannoni SJ (2004) Cultivation and growth characteristics of a diverse group of oligotrophic marine Gammaproteobacteria. Applied and Environmental Microbiology, 70, 432-440.
URL PMID |
| [9] |
Cho JC, Vergin KL, Morris RM, Giovannoni SJ (2004) Lentisphaera araneosa gen. nov., sp. nov., a transparent exopolymer producing marine bacterium, and the description of a novel bacterial phylum, Lentisphaerae. Environmental Microbiology, 6, 611-621.
URL PMID |
| [10] |
Conway T, Schoolnik GK (2003) Microarray expression profiling: capturing a genome-wide protrait of the transcriptome. Molecular Microbiology, 47, 879-889.
URL PMID |
| [11] |
Crump BC, Armbrust EV, Baross JA (1999) Phylogenetic analysis of particle-attached and free-living bacterial communities in the Columbia River, its estuary, and the adjacent coastal ocean. Applied and Environmental Microbiology, 65, 3192-3204.
URL PMID |
| [12] | Crump BC, Hobbie JE (2005) Synchrony and seasonality in bacterioplankton diversity of two temperate rivers. Limnology and Oceanography, 50, 1718-1729. |
| [13] |
Crump BC, Hopkinson CS, Sogin ML, Hobbie JE (2004) Microbial biogeography along an estuarine salinity gradient: combined influences of bacterial growth and residence time. Applied and Environmental Microbiology, 70, 1494-1505.
URL PMID |
| [14] | Crump BC, Peterson BJ, Raymond PA, Amon RMW, Rinehart A, McClelland JW, Holmes RM (2009) Circumpolar synchrony in big river bacterioplankton. Proceedings of the National Academy of Sciences, USA, 106, 21208-21212. |
| [15] | Daley RJ, Hobbie JE (1975) Direct counts of aquatic bacteria by a modified epifluorescent technique. Limnology and Oceanography, 20, 875-882. |
| [16] | de la Torre JR, Christianson LM, Béjà O, Suzuki MT, Karl DM, Heidelberg J, DeLong EF (2003) Proteorhodopsin genes are distributed among divergent marine bacterial taxa. Proceedings of the National Academy of Sciences,USA, 100, 12830-12835. |
| [17] | del Giorgio PA, Bouvier TC (2002) Linking the physiological and phylogenetic succession in free-living bacterial communities along an estuarine salinity gradient. Limnology and Oceanography, 47, 471-486. |
| [18] |
DeLong EF (2005) Microbial community genomics in the ocean. Nature Reviews Microbiology, 3, 459-469.
DOI URL PMID |
| [19] |
DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, Frigaard N-U, Martinez A, Sullivan MB, Edwards R, Brito BR, Chisholm SW, Karl DM (2006) Community genomics among stratified microbial assemblages in the ocean's interior. Science, 311, 496-503.
URL PMID |
| [20] |
Eymann C, Homuth G, Scharf C, Hecker M (2002) Bacillus subtilis functional genomics: global characterization of the stringent response by proteome and transcriptome analysis. Journal of Bacteriology, 184, 2500-2520.
DOI URL PMID |
| [21] | Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S (2006) Annually reoccurring bacterial communities are predictable from ocean conditions. Proceedings of the National Academy of Sciences, USA, 103, 13104-13109. |
| [22] |
Giovannoni SJ, Britschgi TB, Moyer CL, Field KG (1990) Genetic diversity in Sargasso Sea bacterioplankton. Nature, 345, 60-63.
URL PMID |
| [23] | Giovannoni SJ, Rappé MS (2000) Evolution, diversity, and molecular ecology of marine prokaryotes. In: Microbial Ecology of the Oceans (ed. Kirchman DL), pp. 47-84. Wiley-Liss, New York. |
| [24] | Glibert PM, Conley DJ, Fisher TR, Harding LWJ, Malone TC (1995) Dynamics of the 1990 winter/spring bloom in Chesapeake Bay. Marine Ecology: Progress Series, 122, 27-43. |
| [25] |
Hahn MW, Lunsdorf H, Wu Q, Schauer M, Höfle MG, Boenigk J, Stadler P (2003) Isolation of novel ultramicrobacterial classified as Actinobacteria from five freshwater habitats in Europe and Asia. Applied and Environmental Microbiology, 69, 1442-1451.
DOI URL PMID |
| [26] |
Heidelberg JF, Heidelberg KB, Colwell RR (2002) Seasonality of Chesapeake Bay bacterioplankton species. Applied and Environmental Microbiology, 68, 5488-5497.
DOI URL PMID |
| [27] | Henriques IS, Alves A, Tacao M, Almeida A, Cunha A, Correia A (2006) Seasonal and spatial variability of free-living bacterial community composition along an estuarine gradient (Ria de Aveiro, Portugal). Estuarine Coastal and Shelf Science, 68, 139-148. |
| [28] | Hollibaugh JT, Wong PS, Murrell MC (2000) Similarity of particle-associated and free-living bacterial communities in northern San Francisco Bay, California. Aquatic Microbial Ecology, 21, 103-109. |
| [29] | Hoppe HG (1976) Determination and properties of actively metabolizing heterotrophic bacteria in the sea, investigated by the means of micro-autoradiography. Marine Biology, 36, 291-302. |
| [30] | Jannasch HW, Jones GE (1959) Bacterial populations in sea water as determined by different methods of enumeration. Limnology and Oceanography, 4, 128-139. |
| [31] |
Janssen PH, Yates PS, Grinton BE, Taylor PM, Sait M (2002) Improved culturability of soil bacteria and isolation in pure culture of novel members of the divisions Acidobacteria, Actinobacteria, Proteobacteria, and Verrucomicrobia. Applied and Environmental Microbiology, 68, 2391-2396.
URL PMID |
| [32] | Kan J, Chen F (2004) Co-monitoring bacterial and dinoflagellates communities by denaturing gradient gel electrophoresis (DGGE) and SSU rDNA sequencing during a dinoflagellate bloom. Acta Oceanologica Sinica, 23, 483-492. |
| [33] | Kan J, Crump BC, Wang K, Chen F (2006a) Bacterioplankton community in Chesapeake Bay: Predictable or random assemblages. Limnology and Oceanography, 51, 2157-2169. |
| [34] | Kan J, Wang K, Chen F (2006b) Temporal variation and detection limit of an estuarine bacterioplankton community analyzed by denaturing gradient gel electrophoresis (DGGE). Aquatic Microbial Ecology, 42, 7-18. |
| [35] |
Kan J, Suzuki M, Wang K, Evans SE, Chen F (2007) High temporal but low spatial heterogeneity of bacterioplankton in the Chesapeake Bay. Applied and Environmental Microbiology, 73, 6776-6789.
DOI URL PMID |
| [36] | Kan J, Evans SE, Chen F, Suzuki MT (2008) Novel estuarine bacterioplankton in rRNA operon libraries from the Chesapeake Bay. Aquatic Microbial Ecology, 51, 55-66. |
| [37] | Kan J, Hanson TE, Chen F (2011) Synchronicity between population structure and proteome profiles: a metaproteomic analysis of Chesapeake Bay bacterial communities. In: Handbook of Molecular Microbial Ecology, Volume I: Metagenomics and Complementary Approaches (ed. de Bruijn FJ), pp. 637-644. Wiley-Blackwell, Hoboken. |
| [38] |
Kan J, Hanson TE, Ginter JM, Wang K, Chen F (2005) Metaproteomic analysis of Chesapeake Bay microbial communities. Saline Systems, 1, 7.
DOI URL PMID |
| [39] |
Kent AD, Jones SE, Yannarell AC, Graham JM, Lauster GH, Kratz TK, Triplett EW (2004) Annual patterns in bacterioplankton community variability in a humic lake. Microbial Ecology, 48, 550-560.
DOI URL PMID |
| [40] |
Kirchman DL, Rich JH (1997) Regulation of bacterial growth rates by dissolved organic carbon and temperature in the equatorial Pacific Ocean. Microbial Ecology, 33, 11-20.
DOI URL PMID |
| [41] | Lopez MF (1999) Proteome analysis I. Gene products are where the biological action is. Journal of Chromatography B, 722, 191-202. |
| [42] | Malone TC, Kemp WM, Ducklow HW, Boynton WR, Tuttle JH, Jonas RB (1986) Lateral variation in the production and fate of phytoplankton in a partially stratified estuary. Marine Ecology-Progress Series, 32, 149-160. |
| [43] |
Martinez A, Tyson GW, DeLong EF (2009) Widespread known and novel phosphonate utilization pathways in marine bacteria revealed by functional screening and metagenomic analyses. Environmental Microbiology, 12, 222-238.
DOI URL PMID |
| [44] | Monsen NE, Cloern JE, Lucas LV, Monismith SG (2002) A comment on the use of flushing time, residence time, and age as transport time scales. Limnology and Oceanography, 47, 1545-1553. |
| [45] |
Murray AE, Preston CM, Massana R, Taylor LT, Blakis A, Wu K, DeLong EF (1998) Seasonal and spatial variability of bacterial and archaeal assemblages in the coastal waters near Anvers Island, Antarctica. Applied and Environmental Microbiology, 64, 2585-2595.
DOI URL PMID |
| [46] | Newton RJ, Griffin LE, Bowles KM, Meile C, Gifford S, Givens CE, Howard EC, King E, Oakley CA, Reisch CR, Rinta-Kanto JM, Sharma S, Sun S, Varaljay V, Vila-Costa M, Westrich JR, Moran MA (2010) Genome characteristics of a generalist marine bacterial lineage. ISME Journal, 4, 784-798. |
| [47] | Nixon SW, Ammerman JW, Atkinson LP, Berounsky VM, Billen G, Boicourt WC, Boynton WR, Church TM, Ditoro DM, Elmgren R, Garber JH, Giblin AE, Jahnke RA, Owens NJP, Pilson MEQ, Seitzinger SP (1996) The fate of nitrogen and phosphorus at the land sea margin of the North Atlantic Ocean. Biogeochemistry, 35, 141-180. |
| [48] |
Olsen GJ, Lane DJ, Giovannoni J, Pace NR (1986) Microbial ecology and evolution: a ribosomal RNA approach. Annual Review of Microbiology, 40, 337-365.
URL PMID |
| [49] | Pace NR, Stahl DA, Olsen GJ, Lane DJ (1985) Analyzing natural microbial populations by rRNA sequences. American Society of Microbiology News, 51, 4-12. |
| [50] |
Pernthaler J, Glöckner FO, Unterholzner S, Alfreider A, Psenner R, Amann R (1998) Seasonal community and population dynamics of pelagic bacteria and archaea in a high mountain lake. Applied and Environmental Microbiology, 64, 4299-4306.
DOI URL PMID |
| [51] |
Petersohn A, Brigulla M, Haas S, Hoheisel JD, Volker U, Hecker M (2001) Global analysis of the general stress response of Bacillus subtilis. Journal of Bacteriology, 183, 5617-5631.
URL PMID |
| [52] |
Pinhassi M, Sala M, Havskum H, Peters F, Guadayol O, Malits A, Marrase C (2004) Changes in bacterioplankton composition under different phytoplankton regimens. Applied and Environmental Microbiology, 70, 6753-6766.
URL PMID |
| [53] | Pomeroy LR, Wiebe WJ (2001) Temperature and substrates as interactive limiting factors for marine heterotrophic bacteria. Aquatic Microbial Ecology, 23, 187-204. |
| [54] | Porter KG, Feig YS (1980) The use of DAPI for identifying and counting aquatic microflora. Limnology and Oceanography, 25, 943-948. |
| [55] |
Ram RJ, VerBerkmoes NC, Thelen MP, Tyson GW, Baker BJ, Blake II RC, Shah M, Hettich RL, Banfield JF (2005) Community proteomics of a natural microbial biofilm. Science, 308, 1915-1920.
DOI URL PMID |
| [56] |
Rappé MS, Cannon SA, Vergin KL, Giovannoni SJ (2002) Cultivation of the ubiquitous SAR11 marine bacterioplankton clade. Nature, 418, 630-633.
URL PMID |
| [57] |
Riemann L, Steward GF, Azam F (2000) Dynamics of bacterial community composition and activity during a mesocosm diatom bloom. Applied and Environmental Microbiology, 66, 578-587.
DOI URL PMID |
| [58] |
Riesenfeld CS, Schloss PD, Handelsman J (2004) Metagenomics: genomic analysis of microbial communities. Annual Review of Genetics, 38, 525-552.
DOI URL PMID |
| [59] | SAS Institute Inc (1992) SAS User's Guide. SAS Institute Inc. |
| [60] |
Sekiguchi H, Koshikawa H, Hiroki M, Murakami S, Xu K, Watanabe M, Nakahara T, Zhu M, Uchiyama H (2002) Bacterial distribution and phylogenetic diversity in the Changjiang estuary before the construction of the Three Gorges Dam. Microbial Ecology, 43, 82-91.
URL PMID |
| [61] | Selje N, Simon M (2003) Composition and dynamics of particle-associated and free-living bacterial communities in the Weser estuary, Germany. Aquatic Microbial Ecology, 30, 221-237. |
| [62] | Shiah F-K, Ducklow HW (1994) Temperature regulation of heterotrophic bacterioplankton abundance, production, and specific growth rate. Limnology and Oceanography, 39, 1243-1258. |
| [63] |
Simon C, Daniel R (2011) Metagenomic analyses: past and future trends. Applied and Environmental Microbiology, 77, 1153-1161.
URL PMID |
| [64] | Smith EM, Kemp WM (2003) Planktonic and bacterial respiration along an estuarine gradient: response to carbon and nutrient enrichment. Aquatic Microbial Ecology, 30, 251-261. |
| [65] | Sowell SM, Wilhelm LJ, Norbeck AD, Lipton MS, Nicora CD, Barofsky DF, Carlson CA, Smith RD, Giovanonni SJ (2008) Transport functions dominate the SAR11 metaproteome at low-nutrient extremes in the Sargasso Sea. ISME Journal, 3, 93-105. |
| [66] |
Staley JT, Konopka A (1985) Measurement of in situ activities of nonphotosynthetic microorganisms in aquatic and terrestrial habitats. Annual Review of Microbiology, 39, 321-346.
URL PMID |
| [67] |
Stevenson, BS, Eichorst SA, Wertz JT, Schmidt TM, Breznak JA (2004) New strategies for cultivation and detection of previously uncultured microbes. Applied and Environmental Microbiology, 70, 4748-4755.
URL PMID |
| [68] | Sun J, Feng YY, Zhang YH, Hutchins DA (2007) Fast microzooplankton grazing on fast-growing, low-biomass phytoplankton ― A case study in spring in the Chesapeake Bay, Delaware Inland Bays and the Delaware Bay. Hydrobiology, 589, 127-139. |
| [69] |
Temperton B, Gilbert JA, Quinn JP, McGrath JW (2011) Novel analysis of oceanic surface water metagenomes suggests importance of polyphosphate metabolism in oligotrophic environments. PLoS ONE, 6, e16499. doi: 10.1371/journal. pone.0016499
URL PMID |
| [70] |
Tringe SG, von Mering C, Kobayashi A, Salamov AA, Chen K, Chang HW, Podar M, Short JM, Mathur EJ, Detter JC, Bork P, Hugenholtz P, Rubin EM (2005) Comparative metagenomics of microbial communities. Science, 308, 554-557.
DOI URL PMID |
| [71] | Troussellier M, Schäfer H, Batailler N, Bernard L, Courties C, Lebaron P, Muyzer G, Servais P, Vives-Rego J (2002) Bacterial activity and genetic richness along an estuarine gradient (Rhone River plume, France). Aquatic Microbial Ecology, 28, 13-24. |
| [72] |
Tyson GW, Chapman J, Hugenholtz P, Allen EE, Ram RJ, Richardson PM, Solovyev VV, Rubin EM, Rokhsar DS, Banfield JF (2004) Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature, 428, 37-43.
DOI URL PMID |
| [73] |
Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA, Wu D, Paulsen I, Nelson KE, Nelson W, Fouts DE, Levy S, Knap AH, Lomas MW, Nealson K, White O, Peterson J, Hoffman J, Parsons R, Baden-Tillson H, Pfannkoch C, Rogers Y, Smith HO (2004) Environmental genome shotgun sequencing of the Sargasso Sea. Science, 304, 66-74.
URL PMID |
| [74] | Wikner J, Hagström A (1991) Annual study of bacteri- oplankton community dynamics. Limnology and Oceanography, 36, 1313-1324. |
| [75] |
Wilmes P, Bond PL (2004) The application of two-dimensional polyacrylamide gel electrophoresis and downstream analyses to a mixed community of prokaryotic microorganisms. Environmental Microbiology, 6, 911-920.
DOI URL PMID |
| [76] |
ZoBell CE (1946) Marine Microbiology. Chronica Botanica, Waltham, Mass.
DOI URL PMID |
| [1] | 李庆多, 栗冬梅. 全球蝙蝠巴尔通体流行状况分析[J]. 生物多样性, 2023, 31(9): 23166-. |
| [2] | 冯晨, 张洁, 黄宏文. 统筹植物就地保护与迁地保护的解决方案: 植物并地保护(parallel situ conservation)[J]. 生物多样性, 2023, 31(9): 23184-. |
| [3] | 齐海玲, 樊鹏振, 王跃华, 刘杰. 中国北方六省区胡桃的遗传多样性和群体结构[J]. 生物多样性, 2023, 31(8): 23120-. |
| [4] | 景昭阳, 程可光, 舒恒, 马永鹏, 刘平丽. 全基因组重测序方法在濒危植物保护中的应用[J]. 生物多样性, 2023, 31(5): 22679-. |
| [5] | 熊飞, 刘红艳, 翟东东, 段辛斌, 田辉伍, 陈大庆. 基于基因组重测序的长江上游瓦氏黄颡鱼群体遗传结构[J]. 生物多样性, 2023, 31(4): 22391-. |
| [6] | 蒲佳佳, 杨平俊, 戴洋, 陶可欣, 高磊, 杜予州, 曹俊, 俞晓平, 杨倩倩. 长江下游外来生物福寿螺的种类及其种群遗传结构[J]. 生物多样性, 2023, 31(3): 22346-. |
| [7] | 何艺玥, 刘玉莹, 张富斌, 秦强, 曾燏, 吕振宇, 杨坤. 梯级水利工程背景下的嘉陵江干流蛇鮈群体遗传多样性和遗传结构[J]. 生物多样性, 2023, 31(11): 23160-. |
| [8] | 朱瑞良, 马晓英, 曹畅, 曹子寅. 中国苔藓植物多样性研究进展[J]. 生物多样性, 2022, 30(7): 22378-. |
| [9] | 孙维悦, 舒江平, 顾钰峰, 莫日根高娃, 杜夏瑾, 刘保东, 严岳鸿. 基于保护基因组学揭示荷叶铁线蕨的濒危机制[J]. 生物多样性, 2022, 30(7): 21508-. |
| [10] | 陶克涛, 白东义, 图格琴, 赵若阳, 安塔娜, 铁木齐尔·阿尔腾齐米克, 宝音德力格尔, 哈斯, 芒来, 韩海格. 基于基因组SNPs对东亚家马不同群体遗传多样性的评估[J]. 生物多样性, 2022, 30(5): 21031-. |
| [11] | 崔静, 徐明芳, 章群, 李瑶, 曾晓舒, 李莎. 基于3种线粒体标记探讨中日沿海角木叶鲽遗传多样性差异[J]. 生物多样性, 2022, 30(5): 21485-. |
| [12] | 翟俊杰, 赵慧峰, 商光申, 孙振钧, 张玉峰, 王兴. 蚯蚓基因组学的研究进展: 基于全基因组及线粒体基因组[J]. 生物多样性, 2022, 30(12): 22257-. |
| [13] | 孙军, 宋煜尧, 施义锋, 翟键, 燕文卓. 近十年中国海洋生物多样性研究进展[J]. 生物多样性, 2022, 30(10): 22526-. |
| [14] | 刘山林, 邱娜, 张纾意, 赵竹楠, 周欣. 基因组学技术在生物多样性保护研究中的应用[J]. 生物多样性, 2022, 30(10): 22441-. |
| [15] | 栗冬梅, 杨卫红, 李庆多, 韩茜, 宋秀平, 潘虹, 冯云. 巴尔通体在滇西南蝙蝠中高度流行并具有丰富的遗传变异特征[J]. 生物多样性, 2021, 29(9): 1245-1255. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
备案号:京ICP备16067583号-7
Copyright © 2022 版权所有 《生物多样性》编辑部
地址: 北京香山南辛村20号, 邮编:100093
电话: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn
