



生物多样性 ›› 2018, Vol. 26 ›› Issue (12): 1289-1295. DOI: 10.17520/biods.2018121 cstr: 32101.14.biods.2018121
杨计平1, 李策1,2, 陈蔚涛1, 李跃飞1, 李新辉1,*( )
)
收稿日期:2018-04-16
接受日期:2018-04-16
出版日期:2018-12-20
发布日期:2019-02-11
通讯作者:
李新辉
作者简介:# 共同第一作者
基金资助:
Jiping Yang1, Ce Li1,2, Weitao Chen1, Yuefei Li1, Xinhui Li1,*( )
)
Received:2018-04-16
Accepted:2018-04-16
Online:2018-12-20
Published:2019-02-11
Contact:
Li Xinhui
About author:# 同等贡献作者 Contributed equally to this work
摘要:
鳤(Ochetobius elongatus)曾经是我国许多河流的重要经济鱼类, 然而由于环境污染和人为干扰等原因, 鳤的资源量萎缩严重, 种群处于极度濒危的境地。目前, 常规采样方法获得鳤样本的难度较大, 致使相关研究难以开展。本研究通过仔鱼采集和成鱼采集的手段, 测定两个线粒体基因和两个核基因, 对西江中下游鳤的遗传多样性和种群动态历史进行研究。结果显示, 西江中下游鳤的遗传多样性处于较低水平并且出现了衰退的迹象, 可能经历了遗传瓶颈效应。此外, 种群动态历史分析发现, 西江中下游鳤在后更新世期间(0.06和0.13百万年前)经历了种群扩张, 刚好处于中更新世(0.78-0.126百万年前)冰期褪去之后, 表明更新世气候的波动影响了西江中下游鳤的种群动态。作为鳤可能的重要产卵场, 西江中下游部分江段可以考虑建立鳤的自然保护区, 用于保育和修复鳤的种质资源。
杨计平, 李策, 陈蔚涛, 李跃飞, 李新辉 (2018) 西江中下游鳤的遗传多样性与种群动态历史. 生物多样性, 26, 1289-1295. DOI: 10.17520/biods.2018121.
Jiping Yang, Ce Li, Weitao Chen, Yuefei Li, Xinhui Li (2018) Genetic diversity and population demographic history of Ochetobius elongatus in the middle and lower reaches of the Xijiang River. Biodiversity Science, 26, 1289-1295. DOI: 10.17520/biods.2018121.
| 采样点 Sample site | 采样日期 Sampling time | 样本量 Sample size | 经度 Longitude | 纬度 Latitude |
|---|---|---|---|---|
| 广西桂平 Guiping | 2017.09 | 1 | 110°05°54.78°° E | 23°24°08.25°° N |
| 广西梧州 Wuzhou | 2017.09 | 1 | 111°17°15.32°° E | 23°28°07.94°° N |
| 广东肇庆 Zhaoqing | 2011.05 | 22 | 112°23°40.13°° E | 23°05°06.24°° N |
| 广东肇庆 Zhaoqing | 2013.05 | 28 | 112°23°40.13°° E | 23°05°06.24°° N |
表1 本研究的采样信息
Table 1 Sample information of the present study
| 采样点 Sample site | 采样日期 Sampling time | 样本量 Sample size | 经度 Longitude | 纬度 Latitude |
|---|---|---|---|---|
| 广西桂平 Guiping | 2017.09 | 1 | 110°05°54.78°° E | 23°24°08.25°° N |
| 广西梧州 Wuzhou | 2017.09 | 1 | 111°17°15.32°° E | 23°28°07.94°° N |
| 广东肇庆 Zhaoqing | 2011.05 | 22 | 112°23°40.13°° E | 23°05°06.24°° N |
| 广东肇庆 Zhaoqing | 2013.05 | 28 | 112°23°40.13°° E | 23°05°06.24°° N |
| 引物名 Primer | 引物序列 Primer sequence (5′-3′) | 退火温度 Annealed temperature (℃) | 参考文献 Reference | |
|---|---|---|---|---|
| Cytb | L14724 | GACTTGAAA AACCACCGTTG | 58-64 | Xiao et al, 2001 |
| H15915 | CTCCGATCTCCGGATTACAAGAC | |||
| ND2 | AFND2L | AAGCTYTYGGGCCCATACC | 58 | 黎瑞宝, 2013① |
| AFND2R | TCCYGCTTAGGGCTTTGAAGG | |||
| myh6 | myh6_F459 | CATMTTYTCCATCTCAGATAATGC | 53 | Chen et al, 2008 |
| myh6_R1325 | ATTCTCACCACCATCCAGTTGAA | |||
| RAG2 | RAG2-f2a | AARCGCTCMTGTCCMACTGG | 55 | Lovejoy & Collette, 2001 |
| RAG2-R6a | TGRTCCARGCAGAAGTACTTG |
表2 本研究使用的引物信息
Table 2 Primer information used in the present study
| 引物名 Primer | 引物序列 Primer sequence (5′-3′) | 退火温度 Annealed temperature (℃) | 参考文献 Reference | |
|---|---|---|---|---|
| Cytb | L14724 | GACTTGAAA AACCACCGTTG | 58-64 | Xiao et al, 2001 |
| H15915 | CTCCGATCTCCGGATTACAAGAC | |||
| ND2 | AFND2L | AAGCTYTYGGGCCCATACC | 58 | 黎瑞宝, 2013① |
| AFND2R | TCCYGCTTAGGGCTTTGAAGG | |||
| myh6 | myh6_F459 | CATMTTYTCCATCTCAGATAATGC | 53 | Chen et al, 2008 |
| myh6_R1325 | ATTCTCACCACCATCCAGTTGAA | |||
| RAG2 | RAG2-f2a | AARCGCTCMTGTCCMACTGG | 55 | Lovejoy & Collette, 2001 |
| RAG2-R6a | TGRTCCARGCAGAAGTACTTG |
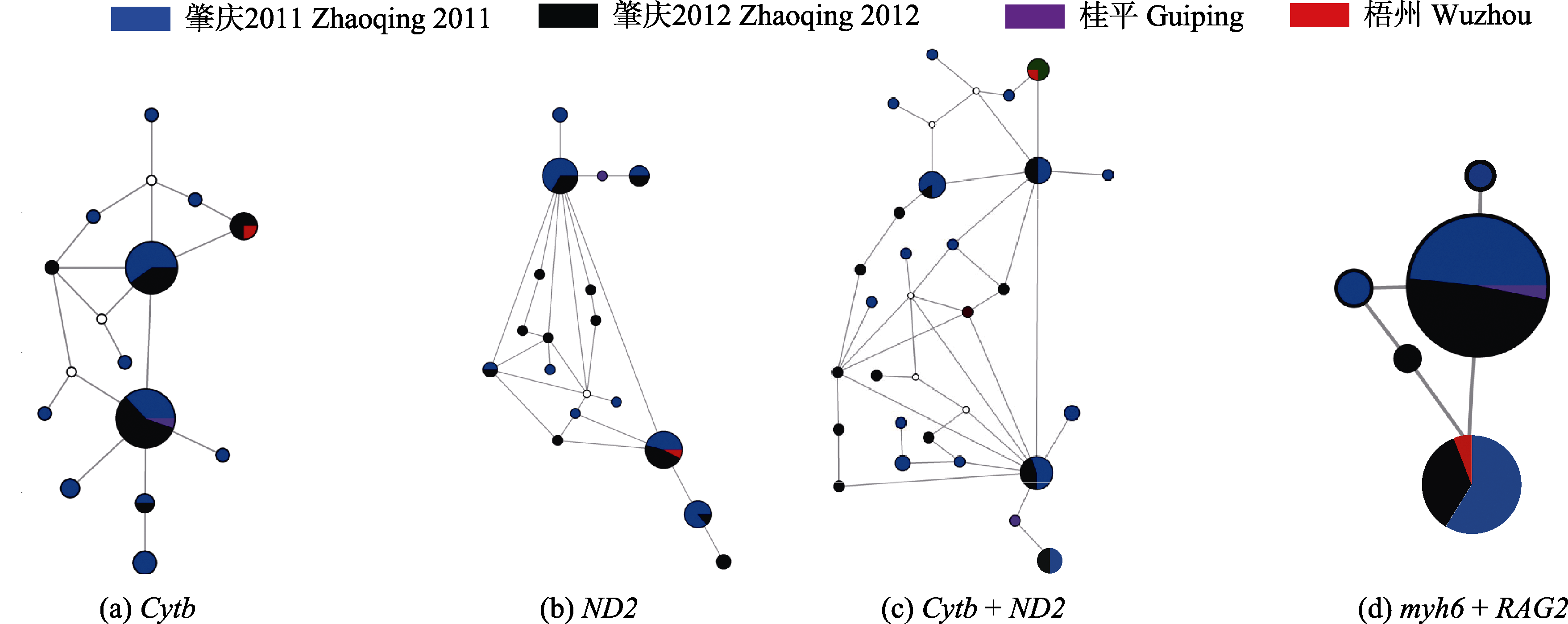
图1 基于不同基因和基因组合的单倍型/等位基因中介网状图。白色圆圈表示缺失单倍型。
Fig. 1 Median-joining networks of haplotype and allele based on different genes and gene groups. White circles indicate missing haplotype.
| 单倍型数/个体数 Number of haplotype/ sample size | 单倍型多态性 Haplotype diversity | 近期核苷酸多态性 Current nucleotide diversity (θπ, %) | 历史核苷酸多态性 Historical nucleotide diversity (θw, %) | Tajima’s D 检验 Tajima’s D test | Fu’s Fs检验 Fu’s Fs test | 最优模型 Optimal model | |
|---|---|---|---|---|---|---|---|
| Cytb | 13/52 | 0.784 | 0.197 | 0.470 | -1.83** | -5.43** | HKY+I |
| ND2 | 17/52 | 0.869 | 0.239 | 0.272 | -0.35 | -8.16*** | F81+I+G |
| Cytb + ND2 | 26/52 | 0.940 | 0.219 | 0.366 | -1.33 | -14.54*** | - |
| myh6 | 2/52 | 0.462 | 0.060 | 0.029 | - | - | HKY |
| RAG2 | 3/52 | 0.147 | 0.022 | 0.077 | - | - | F81 |
| myh6 + RAG2 | 5/52 | 0.546 | 0.040 | 0.055 | - | - | - |
表3 鳤遗传多样性指数、中性检验和最优核苷酸替代模型
Table 3 Genetic diversity indices, neutrality tests and optimal nucleotide substitution model of Ochetobius elongatus
| 单倍型数/个体数 Number of haplotype/ sample size | 单倍型多态性 Haplotype diversity | 近期核苷酸多态性 Current nucleotide diversity (θπ, %) | 历史核苷酸多态性 Historical nucleotide diversity (θw, %) | Tajima’s D 检验 Tajima’s D test | Fu’s Fs检验 Fu’s Fs test | 最优模型 Optimal model | |
|---|---|---|---|---|---|---|---|
| Cytb | 13/52 | 0.784 | 0.197 | 0.470 | -1.83** | -5.43** | HKY+I |
| ND2 | 17/52 | 0.869 | 0.239 | 0.272 | -0.35 | -8.16*** | F81+I+G |
| Cytb + ND2 | 26/52 | 0.940 | 0.219 | 0.366 | -1.33 | -14.54*** | - |
| myh6 | 2/52 | 0.462 | 0.060 | 0.029 | - | - | HKY |
| RAG2 | 3/52 | 0.147 | 0.022 | 0.077 | - | - | F81 |
| myh6 + RAG2 | 5/52 | 0.546 | 0.040 | 0.055 | - | - | - |
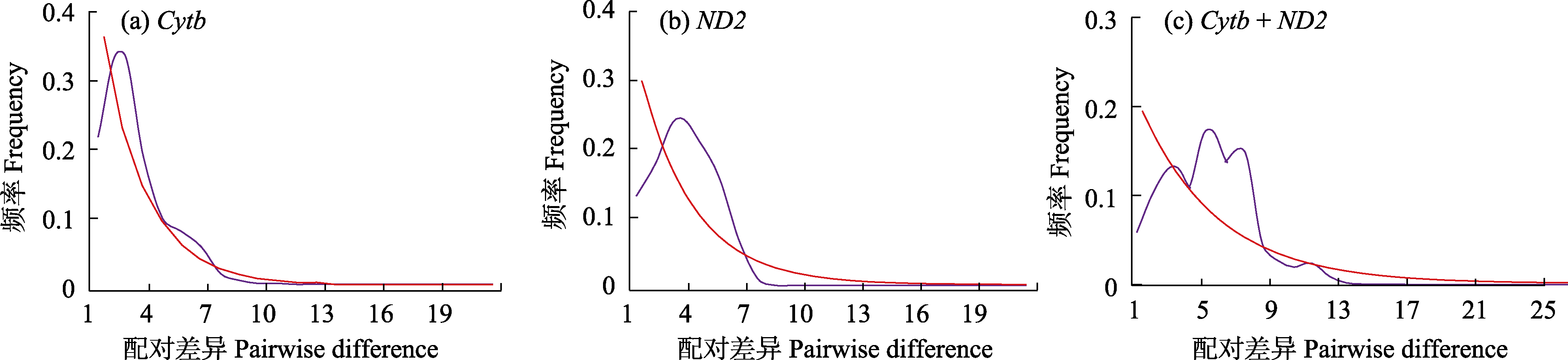
图2 基于不同基因和基因组合的错配分布分析。红色和紫色线条分别表示观测值和期望值。
Fig. 2 Mismatch distribution analysis based on different genes and gene groups. Red and purple lines represent observed value and expected values, respectively.
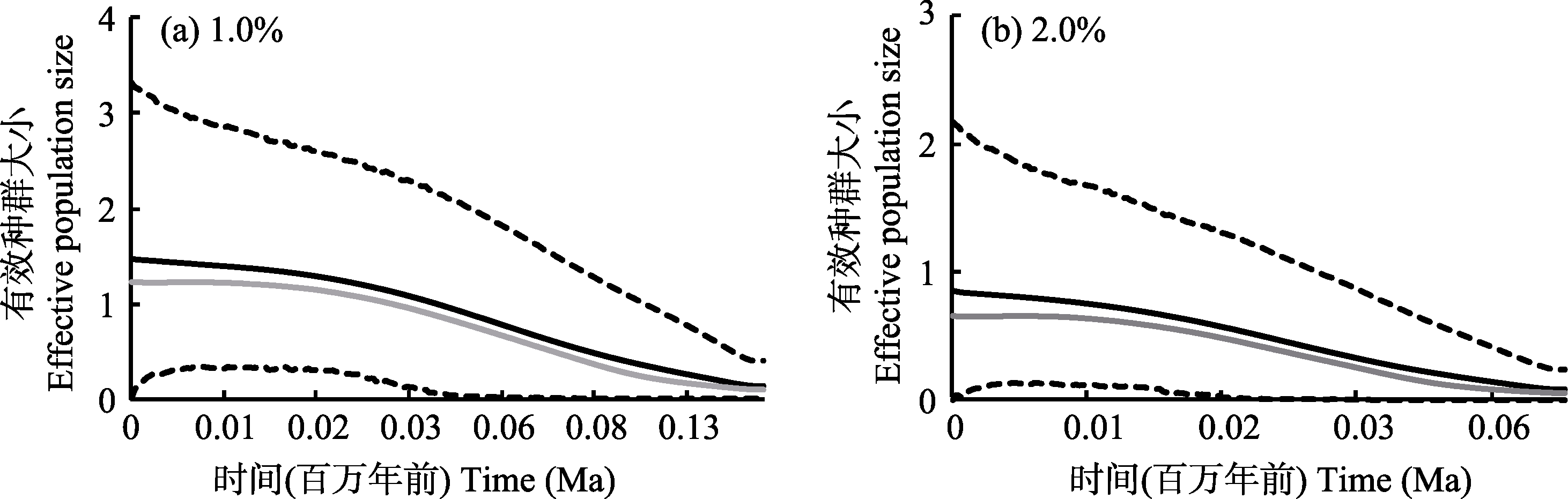
图3 拓展贝叶斯天际线点图(EBSP)。黑色实线表示有效种群大小的平均值, 灰色实线表示有效种群大小的中位数值, 虚线表示95%置信区间的有效种群大小。
Fig. 3 Extended Bayesian Skyline plots. Black lines represent mean values of effective population size, gray lines represent median values of effective population size, and dashed lines represent 95% confidence interval of effective population size.
| 1 | Araujo-Lima CARM, Oliveira EC (1998) Transport of larval fish in the Amazon. Journal of Fish Biology, 53, 297-306. |
| 2 | Bandelt HJ, Forster P, Rohl A (1999) Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution, 16, 37-48. |
| 3 | Chen WJ, Miya M, Saitoh K, Mayden RL (2008) Phylogenetic utility of two existing and four novel nuclear gene loci in reconstructing Tree of Life of ray-finned fishes: The order Cypriniformes (Ostariophysi) as a case study. Gene, 423, 125-134. |
| 4 | Chen YY, Cao WX, Zheng CY (1986) Ichthyofauna of the Zhujiang River with a discussion on zoogeographical divisions for freshwater fishes. Acta Hydrobiologica Sinica, 10, 228-234. (in Chinese with English abstract) |
| [陈宜瑜, 曹文宣, 郑慈英 (1986) 珠江的鱼类区系及其动物地理区划的讨论. 水生生物学报, 10, 228-234.] | |
| 5 | Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evolutionary Biology, 7, 214. |
| 6 | Drummond AJ, Rambaut A, Shapiro B, Pybus OG (2005) Bayesian coalescent inference of past population dynamics from molecular sequences. Molecular Biology and Evolution, 22, 1185-1192. |
| 7 | Durand JD, Tsigenopoulos CS, Unlu E, Berrebi P (2002) Phylogeny and biogeography of the family Cyprinidae in the Middle East inferred from cytochrome b DNA—Evolutionary significance of this region. Molecular Phylogenetics and Evolution, 25, 91-100. |
| 8 | Edgar RC (2004) MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research, 32, 1792-1797. |
| 9 | Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources, 10, 564-567. |
| 10 | Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics, 147, 915-925. |
| 11 | Gascoyne M, Benjamin GJ, Schwarcz HP, Ford DC (1979) Sea-level lowering during the Illinoian glaciation: Evidence from a Bahama “blue hole”. Science, 205, 806-808. |
| 12 | Grant WS, Bowen BW (1998) Shallow population histories in deep evolutionary lineages of marine fishes: Insights from sardines and anchovies and lessons for conservation. Journal of Heredity, 89, 415-426. |
| 13 | Hewitt GM (2004) Genetic consequences of climatic oscillations in the Quaternary. Philosophical Transactions of the Royal Society B-Biological Sciences, 359, 183-195. |
| 14 | Jiang ZG, Jiang JP, Wang YZ, Zhang E, Zhang YY, Li LL, Xie F, Cai B, Cao L, Zheng GM, Dong L, Zhang ZW, Ding P, Luo ZH, Ding CQ, Ma ZJ, Tang SH, Cao WX, Li CW, Hu HJ, Ma Y, Wu Y, Wang YX, Zhou KY, Liu SY, Chen YY, Li JT, Feng ZJ, Wang Y, Wang B, Li C, Song XL, Cai L, Zang CX, Zeng Y, Meng ZB, Fang HX, Ping XG (2016) Red List of China’s Vertebrates. Biodiversidy Science, 24, 500-551. |
| [蒋志刚, 江建平, 王跃招, 张鹗, 张雁云, 李立立, 谢锋, 蔡波, 曹亮, 郑光美, 董路, 张正旺, 丁平, 罗振华, 丁长青, 马志军, 汤宋华, 曹文宣, 李春旺, 胡慧建, 马勇, 吴毅, 王应祥, 周开亚, 刘少英, 陈跃英, 李家堂冯祚建, 王燕, 王斌, 李成, 宋雪琳, 蔡蕾, 臧春鑫, 曾岩, 孟智斌, 方红霞, 平晓鸽 (2016) 中国脊椎动物红色名录. 生物多样性, 24, 500-551.] | |
| 15 | Ketmaier V, Bianco PG, Cobollia M, Krivokapic M, Caniglia R, De Matthaeis E (2004) Molecular phylogeny of two lineages of Leuciscinae cyprinids (Telestes and Scardinius) from the peri-Mediterranean area based on cytochrome b data. Molecular Phylogenetics and Evolution, 32, 1061-1071. |
| 16 | Li J, Li XH, Jia XP, Li YF, He MF, Tan XC, Wang C, Jiang WX (2010) Evolvement and diversity of fish community in Xijiang River. Journal of Fishery Sciences of China, 17, 298-311. (in Chinese with English abstract) |
| [李捷, 李新辉, 贾晓平, 李跃飞, 何美峰, 谭细畅, 王超, 蒋万祥 (2010) 西江鱼类群落多样性及其演变. 中国水产科学, 17, 298-311.] | |
| 17 | Li SF, Lü GQ, Bernatchez L (1998) Diversity of mitochondrial DNA in the populations of silver carp, bighead carp, grass carp and black carp in the middle- and lower reaches of the Yangtze River. Acta Zoologica Sinica, 44, 82-93. (in Chinese with English abstract) |
| [李思发, 吕国庆, Bernatchez L (1998) 长江中下游鲢鳙草青四大家鱼线粒体DNA多样性分析. 动物学报, 44, 82-93.] | |
| 18 | Li YF, Li XH, Tan XC, Li J, Wang C (2012) Occurrence of larval Elopichthys bambusa and its relationship with hydrological conditions in the middle and lower reaches of Pearl River. Journal of Fisheries of China, 36(4), 15-22. (in Chinese with English abstract) |
| [李跃飞, 李新辉, 谭细畅, 李捷, 王超 (2012) 珠江中下游鳡鱼苗的发生及其与水文环境的关系. 水产学报, 36, 15-22.] | |
| 19 | Liang ZS, Mo RL, Chen FC (1985) Species identification and spawning type of common fish species in Xijiang River during early phase. In: The Report of Fishery Research in Pearl River Basin. Compilation Committee of Fishery Resources Survey of Pearl River Basin, Guangzhou. (in Chinese) |
| [梁秩燊, 莫瑞林, 陈福才 (1985) 西江常见鱼类早期发育的分类鉴定及其产卵类型. 见: 珠江水系渔业资源调查研究报告. 珠江水系渔业资源调查编委会, 广州.] | |
| 20 | Librado P, Rozas J (2009) DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25, 1451-1452. |
| 21 | Lovejoy NR, Collette BB (2001) Phylogenetic relationships of new world needlefishes (Teleostei: Belonidae) and the biogeography of transitions between marine and freshwater habitats. Copeia, 2001, 324-338. |
| 22 | Meyer A (1993) Evolution of Mitochondrial DNA in Fishes. Elsevier, The Hague. |
| 23 | Nesler TP, Muth RT, Wasowicz AF (1988) Evidence for baseline flow spikes as spawning cues for Colorado squawfish in the Yampa River, Colorado. In: American Fisheries Society Symposium, pp. 68-79. Bethesda, Maryland. |
| 24 | Nylander JAA (2004) MrModeltest v2. Evolutionary Biology Centre, Uppsala University, Uppsala. |
| 25 | Rambaut A, Drummond A (2007) Tracer v1.4.. (accessed on 2018-3-2). |
| 26 | Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Molecular Biology and Evolution, 9, 552-569. |
| 27 | Tajima F (1989) The effect of change in population size on DNA polymorphism. Genetics, 123, 597-601. |
| 28 | Tao W, Zou M, Wang X, Gan X, Mayden RL, He S (2010) Phylogenomic analysis resolves the formerly intractable adaptive diversification of the endemic clade of east Asian Cyprinidae (Cypriniformes). PLoS ONE, 5, e13508. |
| 29 | Tyus HM, Haines GB (1991) Distribution, habitat use, and growth of age-0 Colorado squawfish in the Green River basin, Colorado and Utah. Transactions of the American Fisheries Society, 120, 79-89. |
| 30 | Wang X, Li J, He S (2007) Molecular evidence for the monophyly of East Asian groups of Cyprinidae (Teleostei: Cypriniformes) derived from the nuclear recombination activating gene 2 sequences. Molecular Phylogenetics and Evolution, 42, 157-170. |
| 31 | Wu WJ, Peng M, Wang DP, Shi J, Li YS, Han YQ, Lei JJ, He AY (2015) Comparison of mitochondrial D-Loop and Cyt b sequences of Hypophthalmichthys molitrix. Open Journal of Fisheries Research, 2(4), 67-73. (in Chinese with English abstract) |
| [吴伟军, 彭敏, 王大鹏, 施军, 李育森, 韩耀全, 雷建军, 何安尤 (2015) 红水河鲢线粒体D-Loop和Cyt b基因序列分析. 水产研究, 2(4), 67-73.] | |
| 32 | Xiao W, Zhang Y, Liu H (2001) Molecular systematics of Xenocyprinae (Teleostei: Cyprinidae): Taxonomy, biogeography, and coevolution of a special group restricted in East Asia. Molecular Phylogenetics and Evolution, 18, 163-173. |
| 33 | Yue PQ (2000) Fauna Sinica, Osteichthyes, Cypriniformes II Science Press, Beijing. (in Chinese) |
| [乐佩琦 (2000) 中国动物志. 硬骨鱼纲, 鲤形目, 中卷. 科学出版社, 北京.] | |
| 34 | Zheng CY (1989) The Ichthyography of the Pearl River. Science Press, Beijing. (in Chinese) |
| [郑慈英 (1989) 珠江鱼类志. 科学出版社. 北京.] | |
| 35 | Zitek A, Schmutz S, Unfer G, Ploner A (2004) Fish drift in a Danube sidearm-system: I. Site-, inter- and intraspecific patterns. Journal of Fish Biology, 65, 1319-1338. |
| [1] | 干靓 刘巷序 鲁雪茗 岳星. 全球生物多样性热点地区大城市的保护政策与优化方向[J]. 生物多样性, 2025, 33(5): 24529-. |
| [2] | 曾子轩 杨锐 黄越 陈路遥. 清华大学校园鸟类多样性特征与环境关联[J]. 生物多样性, 2025, 33(5): 24373-. |
| [3] | 周昊, 王茗毅, 张楚格, 肖治术, 欧阳芳. 昆虫旅馆在独栖蜂多样性保护中的现状与挑战[J]. 生物多样性, 2025, 33(5): 24472-. |
| [4] | 祝晓雨, 王晨灏, 王忠君, 张玉钧. 城市绿地生物多样性研究进展与展望[J]. 生物多样性, 2025, 33(5): 25027-. |
| [5] | 王欣, 鲍风宇. 基于鸟类多样性提升的南滇池国家湿地公园生态修复效果分析[J]. 生物多样性, 2025, 33(5): 24531-. |
| [6] | 明玥, 郝培尧, 谭铃千, 郑曦. 基于城市绿色高质量发展理念的中国城市生物多样性保护与提升研究[J]. 生物多样性, 2025, 33(5): 24524-. |
| [7] | 易木荣, 卢萍, 彭勇, 汤勇, 许久恒, 尹浩萍, 张路杨, 翁晓东, 底明晓, 雷隽, 卢宸祺, 曹如君, 戴年华, 占德洋, 童媚, 楼智明, 丁永刚, 柴静, 车静. 北潦河金家水支流江西大鲵野外种群现状及栖息地评估[J]. 生物多样性, 2025, 33(4): 24145-. |
| [8] | 王太, 宋福俊, 张永胜, 娄忠玉, 张艳萍, 杜岩岩. 河西走廊内陆河水系鱼类多样性及资源现状[J]. 生物多样性, 2025, 33(4): 24387-. |
| [9] | 李沫潼, 何拓, 李薇, 廖菁, 曾岩. 从CITES的术语看野生动植物国际贸易监管规则[J]. 生物多样性, 2025, 33(4): 24545-. |
| [10] | 张晶晶, 黄文彬, 陈奕廷, 杨泽鹏, 柯伟业, 彭昭杰, 魏世超, 张志伟, 胡怡思, 余文华, 周文良. 广东南澎列岛海洋生态国家级自然保护区造礁石珊瑚多样性及分布特征[J]. 生物多样性, 2025, 33(4): 24424-. |
| [11] | 卢晓强, 董姗姗, 马月, 徐徐, 邱凤, 臧明月, 万雅琼, 李孪鑫, 于赐刚, 刘燕. 前沿技术在生物多样性研究中的应用现状、挑战与展望[J]. 生物多样性, 2025, 33(4): 24440-. |
| [12] | 郭雨桐, 李素萃, 王智, 解焱, 杨雪, 周广金, 尤春赫, 朱萨宁, 高吉喜. 全国自然保护地对国家重点保护野生物种的覆盖度及其分布状况[J]. 生物多样性, 2025, 33(3): 24423-. |
| [13] | 赵维洋, 王伟, 马冰然. 其他有效的区域保护措施(OECMs)研究进展与展望[J]. 生物多样性, 2025, 33(3): 24525-. |
| [14] | 周志华, 金效华, 罗颖, 李迪强, 岳建兵, 刘芳, 何拓, 李希, 董晖, 罗鹏. 中国林草部门落实《昆明-蒙特利尔全球生物多样性框架》的机制、成效分析及建议[J]. 生物多样性, 2025, 33(3): 24487-. |
| [15] | 刘立, 臧明月, 马月, 万雅琼, 胡飞龙, 卢晓强, 刘燕. 央地协同推动国家生物多样性战略和行动计划执行的措施、进展与展望[J]. 生物多样性, 2025, 33(3): 24532-. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
备案号:京ICP备16067583号-7
Copyright © 2026 版权所有 《生物多样性》编辑部
地址: 北京香山南辛村20号, 邮编:100093
电话: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn
![]()