



Biodiv Sci ›› 2026, Vol. 34 ›› Issue (1): 25386. DOI: 10.17520/biods.2025386 cstr: 32101.14.biods.2025386
• Special Feature: Methods for Ecological Data Analysis • Previous Articles Next Articles
Chunying Wu1,#( ), Viorel D. Popescu2,#(
), Viorel D. Popescu2,#( ), Yinqiu Ji1,*(
), Yinqiu Ji1,*( )(
)( )
)
Received:2025-10-01
Accepted:2026-01-20
Online:2026-01-20
Published:2026-02-03
Contact:
Yinqiu Ji
About author:First author contact:#Co-first authors
Supported by:Chunying Wu, Viorel D. Popescu, Yinqiu Ji. Hierarchical occupancy models as solutions to challenges in biodiversity assessment[J]. Biodiv Sci, 2026, 34(1): 25386.
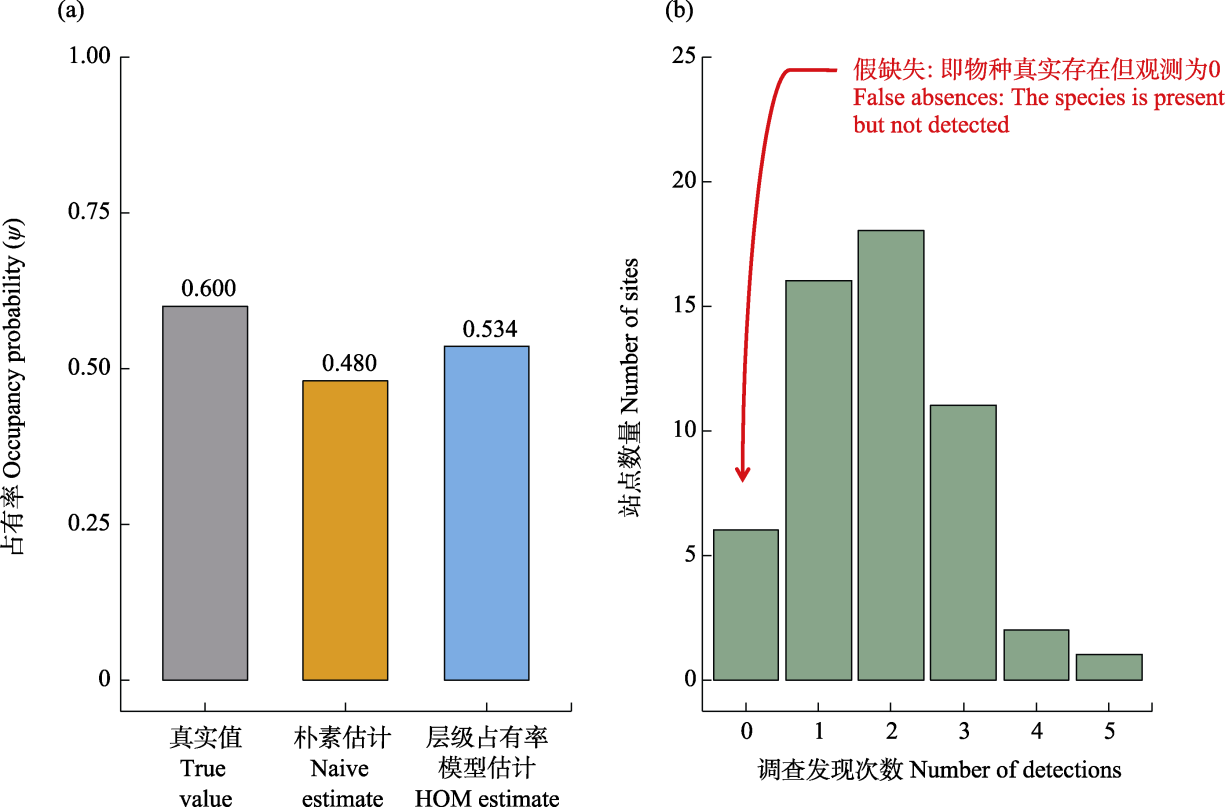
Fig. 2 Simulation of hierarchical occupancy model (HOM) performance in correcting for imperfect detection. (a) Comparison of occupancy probability (ψ) estimates among different methods (sample size n = 100). The naive estimate systematically underestimates occupancy by ignoring imperfect detection, whereas the HOM estimate is closer to the true value. (b) Distribution of detection frequency at sites where the species is truly present. The bar at “0 detections” indicated by the red arrow represents “false absences”, where the species exists at the site but remains undetected due to a detection probability P < 1. This phenomenon is the primary source of bias in occupancy estimation.
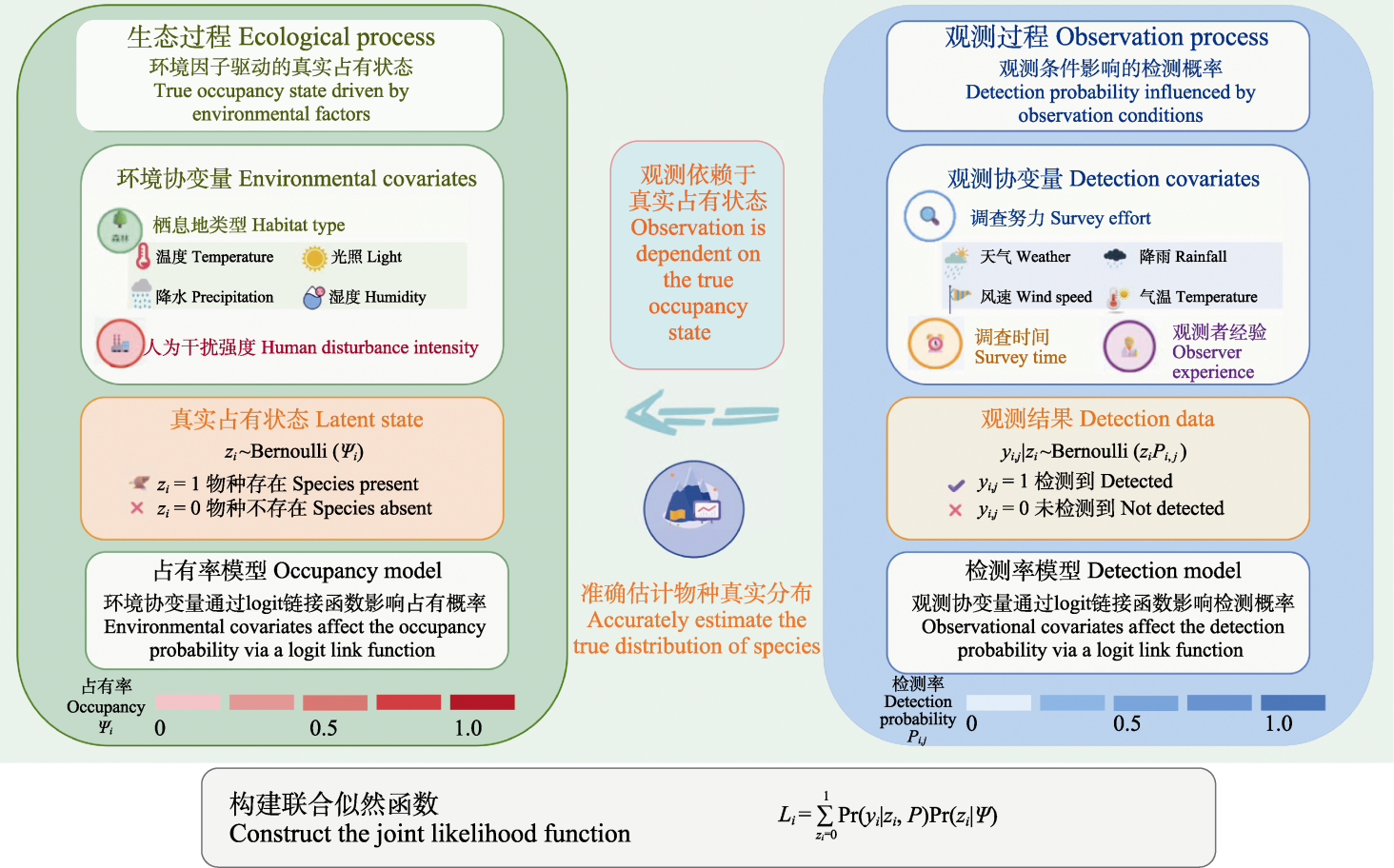
Fig. 3 Dual-level statistical framework of the hierarchical occupancy model. This figure illustrates the dual-level statistical framework of the hierarchical occupancy model, which centers on correcting for imperfect detection in species surveys by decoupling the ecological and observation processes. The ecological process on the left describes the true occupancy state (latent variable zi) driven by environmental factors; the occupancy probability ψ is influenced by environmental covariates—such as habitat type (e.g., forest cover, vegetation structure), abiotic factors, and human disturbance—and is modeled via a logit link function. The observation process on the right describes the detection probability P during field surveys, which is affected by observational covariates including survey effort (duration, intensity, and number of replicates), weather conditions, and observer experience, also linked through a logit function. Given that the observation y is strictly dependent on the true occupancy state z (meaning a species can only be detected if it is truly present), the model integrates these processes by marginalizing over the latent variable zi to construct a joint likelihood function. This approach mathematically accounts for false absences, thereby achieving an unbiased estimation of the true species distribution.
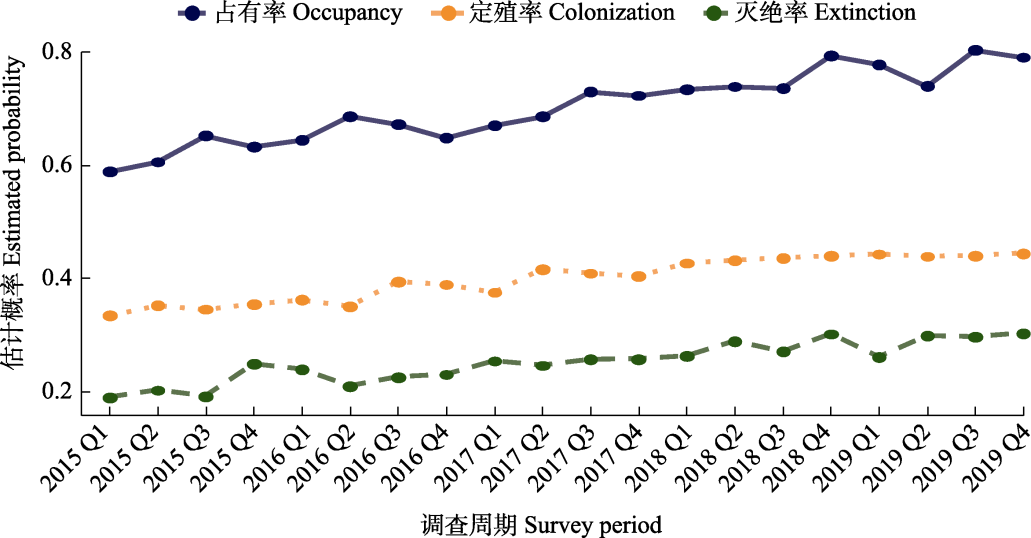
Fig. 4 Schematic diagram of multi-season occupancy dynamics and state transition trends. This diagram illustrates the quarterly dynamics (where Q1-Q4 represent the first to fourth quarters of each year, respectively) of a species based on simulated data between 2015 and 2019: The occupancy rate (ψ, blue solid line) is generally maintained between 0.6-0.8, indicating a high persistent probability of presence in the study area. The colonization probability (γ, orange dotted line) is stable at 0.3-0.4 with a slight upward trend, reflecting a relatively strong capacity for range expansion. The extinction probability (ε, green dashed line) is maintained at 0.1-0.2, suggesting potential habitat disturbance or environmental pressure. This figure is created using simulated data and serves to illustrate the application of the multi-season hierarchical occupancy model (MSOM) in revealing species dynamics. Please see Appendix 2 for the R code.
| 类别 Category | 核心假设 Core assumption | 描述 Description |
|---|---|---|
| 生态过程 Ecological process | 闭合假设 Closure assumption | 在调查期间, 站点占有状态无变化(单季节模型); 动态模型放松此假设(MacKenzie et al., |
| 生态过程 Ecological process | 独立站点 Site independence | 站点间占有率条件独立, 或通过随机效应建模空间相关 Occupancy is conditionally independent among sites, or spatial correlation is accounted for through random effects |
| 观测过程 Observation process | 无假阳性 No false positives | 检测到即真实存在; eDNA等场景需扩展模型处理假阳性(Guillera-Arroita et al., |
| 观测过程 Observation process | 检测独立 Independent detections | 重复调查间检测事件独立 Detection events are independent across replicate surveys |
| 参数估计 Parameter estimation | 局部识别 Local identifiability | 重复调查足够多以分离ψ和P (通常j ≥ 3) A sufficient number of replicate surveys are required to distinguish ψ from P (typically j ≥ 3) |
Table 1 Core assumptions of hierarchical occupancy models
| 类别 Category | 核心假设 Core assumption | 描述 Description |
|---|---|---|
| 生态过程 Ecological process | 闭合假设 Closure assumption | 在调查期间, 站点占有状态无变化(单季节模型); 动态模型放松此假设(MacKenzie et al., |
| 生态过程 Ecological process | 独立站点 Site independence | 站点间占有率条件独立, 或通过随机效应建模空间相关 Occupancy is conditionally independent among sites, or spatial correlation is accounted for through random effects |
| 观测过程 Observation process | 无假阳性 No false positives | 检测到即真实存在; eDNA等场景需扩展模型处理假阳性(Guillera-Arroita et al., |
| 观测过程 Observation process | 检测独立 Independent detections | 重复调查间检测事件独立 Detection events are independent across replicate surveys |
| 参数估计 Parameter estimation | 局部识别 Local identifiability | 重复调查足够多以分离ψ和P (通常j ≥ 3) A sufficient number of replicate surveys are required to distinguish ψ from P (typically j ≥ 3) |
| 类别 Category | 核心内容 Core item | 描述 Description |
|---|---|---|
| 模型选择 Model selection | AICc/WAIC/LOO-CV | 用于比较模型拟合与复杂度; AICc适用于小样本(Burnham & Anderson, |
| 模型选择 Model selection | 后验预测检查 Posterior predictive check (PPC) | 贝叶斯框架下评估模型拟合(Gelman et al., |
| 模型验证 Model selection | 交叉验证 Cross-validation (CV) | 使用k折交叉验证评估预测性能 Evaluating predictive performance using k-fold cross-validation (k-fold CV) |
| 残差诊断 Residual diagnostics | DHARMa残差 DHARMa residuals | 模拟残差检查拟合优度, 适用于层级模型(Hartig, |
Table 2 Framework for model selection and validation in hierarchical occupancy models
| 类别 Category | 核心内容 Core item | 描述 Description |
|---|---|---|
| 模型选择 Model selection | AICc/WAIC/LOO-CV | 用于比较模型拟合与复杂度; AICc适用于小样本(Burnham & Anderson, |
| 模型选择 Model selection | 后验预测检查 Posterior predictive check (PPC) | 贝叶斯框架下评估模型拟合(Gelman et al., |
| 模型验证 Model selection | 交叉验证 Cross-validation (CV) | 使用k折交叉验证评估预测性能 Evaluating predictive performance using k-fold cross-validation (k-fold CV) |
| 残差诊断 Residual diagnostics | DHARMa残差 DHARMa residuals | 模拟残差检查拟合优度, 适用于层级模型(Hartig, |
| [1] | Beger M, Grantham HS, Pressey RL, Wilson KA, Peterson EL, Dorfman D, Mumby PJ, Possingham HP (2010) Conservation planning in a changing world: Multispecies approaches to prioritizing the global protection of biodiversity. BioScience, 60, 52-62. |
| [2] |
Belmont J, Martino S, Illian J, Rue H (2024) Spatio-temporal occupancy models with INLA. Methods in Ecology and Evolution, 15, 2087-2100.
DOI URL |
| [3] |
Brambilla M, Caprio E, Assandri G, Scridel D, Bassi E, Bionda R, Celada C, Falco R, Bogliani G, Pedrini P, Rolando A, Chamberlain D (2017) A spatially explicit definition of conservation priorities according to population resistance and resilience, species importance and level of threat in a changing climate. Diversity and Distributions, 23, 727-738.
DOI URL |
| [4] | Burnham KP, Anderson DR (2002) Model Selection and Multimodel Inference: A Practical Information-Theoretic Approach, 2nd edn. Springer, New York. |
| [5] | Ceballos G, Ehrlich PR, Barnosky AD, García A, Pringle RM, Palmer TM (2015) Accelerated modern human-induced species losses: Entering the sixth mass extinction. Science Advances, 1, e1400253. |
| [6] | Chen IC, Hill JK, Ohlemüller R, Roy DB, Thomas CD (2011) Rapid range shifts of species associated with high levels of climate warming. Science, 333, 1024-1026. |
| [7] |
Crist TO, Veech JA (2006) Additive partitioning of rarefaction curves and species-area relationships: Unifying α-, β- and γ-diversity with sample size and habitat area. Ecology Letters, 9, 923-932.
DOI URL |
| [8] |
Devarajan K, Morelli TL, Tenan S (2020) Multi-species occupancy models: Review, roadmap, and recommendations. Ecography, 43, 1612-1624.
DOI URL |
| [9] | Dorazio RM, Gotelli NJ, Ellison AM (2011) Modern methods of estimating biodiversity from presence-absence surveys. In: BiodiversityLoss in a Changing Planet (ed. Grillo O), pp.71-102. InTech, Rijeka. |
| [10] |
Dorazio RM, Royle JA, Söderström B, Glimskär A (2006) Estimating species richness and accumulation by modeling species occurrence and detectability. Ecology, 87, 842-854.
PMID |
| [11] |
Doser JW, Leuenberger W, Sillett TS, Hallworth MT, Zipkin EF (2022) Integrated community occupancy models: A framework to assess occurrence and biodiversity dynamics using multiple data sources. Methods in Ecology and Evolution, 13, 919-932.
DOI URL |
| [12] |
Ficetola GF, Pansu J, Bonin A, Coissac E, Giguet-Covex C, De Barba M, Gielly L, Lopes CM, Boyer F, Pompanon F, Rayé G, Taberlet P (2015) Replication levels, false presences and the estimation of the presence/absence from eDNA metabarcoding data. Molecular Ecology Resources, 15, 543-556.
DOI URL |
| [13] |
Ficetola GF, Taberlet P (2023) Towards exhaustive community ecology via DNA metabarcoding. Molecular Ecology, 32, 6320-6329.
DOI PMID |
| [14] | Gelman A, Hill J (2007) Data Analysis Using Regression and Multilevel/Hierarchical Models. Cambridge University Press, Cambridge. |
| [15] | Gelman A, Hwang J, Vehtari A (2014) Understanding predictive information criteria for Bayesian models. Statistics and Computing, 24, 997-1016. |
| [16] |
Gronau QF, Sarafoglou A, Matzke D, Ly A, Boehm U, Marsman M, Leslie DS, Forster JJ, Wagenmakers EJ, Steingroever H (2017) A tutorial on bridge sampling. Journal of Mathematical Psychology, 81, 80-97.
DOI PMID |
| [17] |
Guillera-Arroita G, Lahoz-Monfort JJ, van Rooyen AR, Weeks AR, Tingley R (2017) Dealing with false-positive and false-negative errors about species occurrence at multiple levels. Methods in Ecology and Evolution, 8, 1081-1091.
DOI URL |
| [18] | Hamer AJ, Schloegel LM, Rossauer B, Anyomi K, Stockwell MP, Mahony MJ, Clulow S, Banks SC (2021) Multi-species occupancy models reveal community assembly and wetland restoration in amphibians. Journal of Applied Ecology, 58, 607-616. |
| [19] |
Hansen LM, Pfeifer M (2019) Machine learning in remote sensing: A comprehensive review. ISPRS Journal of Photogrammetry and Remote Sensing, 152, 166-177.
DOI URL |
| [20] | Hartig F (2024) DHARMa: Residual Diagnostics for Hierarchical (Multi-level/Mixed) Regression Models. R Package Version 0.4.7. https://CRAN.R-project.org/package=DHARMa. (accessed on 2025-09-02) |
| [21] | Hepler SA, Erhardt RJ (2021) A spatiotemporal model for multivariate occupancy data. Environmetrics, 32, e2657. |
| [22] | Hepler SA, Yang BQ (2024) Occupancy models with autocorrelated detection heterogeneity. Environmental and Ecological Statistics, 31, 777-800. |
| [23] | IPBES (2019) Global Assessment Report on Biodiversity and Ecosystem Services. IPBES Secretariat, Bonn, Germany. |
| [24] |
Ji YQ, Baker CCM, Popescu VD, Wang JX, Wu CY, Wang ZY, Li YH, Wang L, Hua CL, Yang ZX, Yang CY, Xu CCY, Diana A, Wen QZ, Pierce NE, Yu DW (2022) Measuring protected-area effectiveness using vertebrate distributions from leech iDNA. Nature Communications, 13, 1555.
DOI PMID |
| [25] | Ji YQ, Diana A, Li XY, Matechou E, Griffin JE, Liu SW, Luo MJ, Wu CY, Bai R, Yao CY, Yin TT, Dong F, Wu F, Wang K, Yu ZB, Chen XY, Jiang XL, Che J, Yu DW, Popescu VD (2025) High quality, granular, timely, trustworthy and efficient vertebrate species distribution data across a 30,000 km2 protected area complex. Ecology Letters, 28, e70302. |
| [26] | Kelleher S, Guillera-Arroita G, Elith J, Briscoe NJ (2025) Twenty years of dynamic occupancy models: A review of applications and look to the future. Ecography, 48, e07757. |
| [27] | Kéry M, Royle JA (2015) Applied Hierarchical Modeling in Ecology: Analysis of Distribution, Abundance and Species Richness in R and BUGS (Vol. 1): Prelude and Static Models. Academic Press, New York. |
| [28] | Kéry M, Royle JA (2020) Applied Hierarchical Modeling in Ecology: Analysis of Distribution, Abundance and Species Richness in R and BUGS (Vol. 2): Dynamic and Advanced Models. Academic Press, New York. |
| [29] |
Kéry M, Royle JA, Plattner M, Dorazio RM (2009) Species richness and occupancy estimation in communities subject to temporary emigration. Ecology, 90, 1279-1290.
PMID |
| [30] |
Kleiven EF, Barraquand F, Gimenez O, Henden JA, Ims RA, Soininen EM, Yoccoz NG (2023) A dynamic occupancy model for interacting species with two spatial scales. Journal of Agricultural, Biological and Environmental Statistics, 28, 466-482.
DOI |
| [31] |
Lipsey MK, Naugle DE, Nowak J, Lukacs PM (2017) Extending utility of hierarchical models to multi-scale habitat selection. Diversity and Distributions, 23, 783-793.
DOI URL |
| [32] |
MacKenzie DI, Nichols JD, Hines JE, Knutson MG, Franklin AB (2003) Estimating site occupancy, colonization, and local extinction when a species is detected imperfectly. Ecology, 84, 2200-2207.
DOI URL |
| [33] | MacKenzie DI, Nichols JD, Lachman GB, Droege S, Royle JA, Langtimm CA (2002) Estimating site occupancy rates when detection probabilities are less than one. Ecology, 83, 2248-2255. |
| [34] | MacKenzie DI, Nichols JD, Royle JA, Pollock KH, Bailey LL, Hines JE (2017) Occupancy Estimation and Modeling: Inferring Patterns and Dynamics of Species Occurrence, 2nd edn. Academic Press, London. |
| [35] |
MacKenzie DI, Nichols JD, Seamans ME, Gutiérrez RJ (2009) Modeling species occurrence dynamics with multiple states and imperfect detection. Ecology, 90, 823-835.
PMID |
| [36] |
MacKenzie DI, Royle JA (2005) Designing occupancy studies: General advice and allocating survey effort. Journal of Applied Ecology, 42, 1105-1114.
DOI URL |
| [37] |
Margules CR, Pressey RL (2000) Systematic conservation planning. Nature, 405, 243-253.
DOI |
| [38] |
Miller DA, Nichols JD, McClintock BT, Grant EHC, Bailey LL, Weir LA (2011) Improving occupancy estimation when two types of observational error occur: Non-detection and species misidentification. Ecology, 92, 1422-1428.
PMID |
| [39] | Moilanen E, Wilson KA, Possingham HP (2009) Spatial Conservation Prioritization. Oxford University Press, Oxford. |
| [40] |
Moll RJ, Kilshaw K, Montgomery RA, Abade L, Campbell RD, Harrington LA, Millspaugh JJ, Birks JDS, MacDonald DW (2016) Clarifying habitat niche width using broad-scale, hierarchical occupancy models: A case study with a recovering mesocarnivore. Journal of Zoology, 300, 177-185.
DOI URL |
| [41] |
Nichols JD, Williams BK (2006) Monitoring for conservation. Trends in Ecology & Evolution, 21, 668-673.
DOI URL |
| [42] | Popescu VD, de Valpine P, Butsie DC (2016) Biological realism of wildlife population estimates in data-poor systems. Ecological Applications, 26, 1429-1444. |
| [43] |
Rahman H, Moore JF, Connor T, Amin R, Evans MJ, Steinmetz R, Royle JA, Belant JL (2021) Assessing mammalian diversity in northeastern Bangladesh using multi-species occupancy models. Biodiversity and Conservation, 30, 15-34.
DOI |
| [44] |
Rota CT, Ferreira MAR, Kays RW, Forrester TD, Kalies EL, McShea WJ, Parsons AW, Millspaugh JJ (2016) A multispecies occupancy model for two or more interacting species. Methods in Ecology and Evolution, 7, 1164-1173.
DOI URL |
| [45] | Royle JA, Dorazio RM (2008) Hierarchical Modeling and Inference in Ecology:The Analysis of Data from Populations, Metapopulations and Communities. Academic Press, San Diego. |
| [46] |
Rue H, Martino S, Chopin N (2009) Approximate Bayesian inference for latent Gaussian models by using integrated nested Laplace approximations. Journal of the Royal Statistical Society Series B: Statistical Methodology, 71, 319-392.
DOI URL |
| [47] |
Saadi I, Farooq B, Mustafa A, Teller J, Cools M (2018) An efficient hierarchical model for multi-source information fusion. Expert Systems with Applications, 110, 352-362.
DOI URL |
| [48] | Smith AM, Goldberg CS (2020) Occupancy in dynamic systems: Multi-scale and false-positive environmental DNA monitoring. Ecology, 101, e02945. |
| [49] | Sollmann R, Eaton MJ, Link WA, Mulondo P, Ayebare S, Prinsloo S, Plumptre AJ, Johnson DS (2021) A Bayesian Dirichlet process community occupancy model to estimate community structure and species similarity. Ecological Applications, 31, e02249. |
| [50] | Thuiller W, Guéguen M, Renaud J, Karger DN, Zimmermann NE (2019) Climate change and species distribution: From theory to practice. Global Change Biology, 25, 1126-1141. |
| [51] |
Tittensor DP, Walpole M, Hill SLL, Boyce DG, Britten GL, Burgess ND, Butchart SHM, Leadley PW, Regan EC, Alkemade R, Baumung R, Bellard C, Bouwman L, Bowles-Newark NJ, Chenery AM, Cheung WWL, Christensen V, Cooper HD, Crowther AR, Dixon MJR, Galli A, Gaveau V, Gregory RD, Gutierrez NL, Hirsch TL, Höft R, Januchowski-Hartley SR, Karmann M, Krug CB, Leverington FJ, Loh J, Lojenga RK, Malsch K, Marques A, Morgan DHW, Mumby PJ, Newbold T, Noonan-Mooney K, Pagad SN, Parks BC, Pereira HM, Robertson T, Rondinini C, Santini L, Scharlemann JPW, Schindler S, Sumaila UR, Teh LSL, van Kolck J, Visconti P, Ye YM (2014) A mid-term analysis of progress toward international biodiversity targets. Science, 346, 241-244.
DOI PMID |
| [52] |
Tobler MW, Zúñiga Hartley A, Carrillo-Percastegui SE, Powell GVN (2015) Spatiotemporal hierarchical modelling of species richness and occupancy using camera trap data. Journal of Applied Ecology, 52, 413-421.
DOI URL |
| [53] | Vehtari A, Gelman A, Gabry J (2017) Practical Bayesian model evaluation using leave-one-out cross-validation and WAIC. Statistics and Computing, 27, 1413-1432. |
| [54] | Wilson KA, Underwood EC, Morrison SA, Klausmeyer KR, Murdoch WW, Reyers B, Wardell-Johnson G, Marquet PA, Albert AS, Simons AJ (2007) Prioritizing global conservation efforts. Nature, 446, 109-112. |
| [55] |
Zheng J (2023) Impact of transportation infrastructure on amphibians. Theoretical and Natural Science, 6, 426-430.
DOI URL |
| [56] |
Zipkin EF, DeWan A, Andrew Royle J (2009) Impacts of forest fragmentation on species richness: A hierarchical approach to community modelling. Journal of Applied Ecology, 46, 815-822.
DOI URL |
| [1] | Shurong Tian, Ying Wei, Fen Xiao, Yunyun Zhou, Yixing Xie, Cheng Wang, Fen Song, Zhiqiang Liang, Xiaojie Gui. Population dynamics and conservation strategies of Andrias davidianus in Hunan Zhangjiajie Giant Salamander National Nature Reserve, China [J]. Biodiv Sci, 2025, 33(7): 24581-. |
| [2] | Fengxiang Zhou, Xixia Lu, Liming Yong, Qianhui Zeng, Liangliang Yang, Ping Li, Yuke Zhang, Xianyan Wang. Research progress on acoustic monitoring of cetaceans [J]. Biodiv Sci, 2025, 33(7): 24556-. |
| [3] | Nong Qiaoyi, Cao Jun, Cheng Wenda, Peng Yanqiong. Comparative study of monitoring methods for Apoidea resources and diversity [J]. Biodiv Sci, 2025, 33(4): 25057-. |
| [4] | Jinyi Luo, Ji Zhang, Yanling Bi, Zhe Chen, Ruidong Wu. Identifying potential protected areas by integrating multi-faceted conservation features: A case study of Dali Bai Autonomous Prefecture [J]. Biodiv Sci, 2025, 33(11): 25080-. |
| [5] | Zhenzhen Li, Mengtian Du, Yuanxin Zhu, Dawei Wang, Zhilin Li, Tianming Wang. A practical guide for estimating the density of unmarked populations using camera traps [J]. Biodiv Sci, 2023, 31(3): 22422-. |
| [6] | Yifei Sun, Shizheng Wang, Jiawei Feng, Tianming Wang. Diel and seasonal variability of the forest soundscape in the Northeast China Tiger and Leopard National Park [J]. Biodiv Sci, 2023, 31(1): 22523-. |
| [7] | Jian Zhang, Hongzhi Kong, Xiaolei Huang, Shenglei Fu, Liangdong Guo, Qinghua Guo, Fumin Lei, Zhi Lü, Yurong Zhou, Keping Ma. Thirty key questions for biodiversity science in China [J]. Biodiv Sci, 2022, 30(10): 22609-. |
| [8] | Xing Chen, Tianpei Guan, Wenle Jiang, Dandan Li, Kong Yang, Sheng Li. Distribution and population status of bovine species in China based on bibliometric analysis [J]. Biodiv Sci, 2021, 29(5): 668-679. |
| [9] | Guoping Yang, Tao Wu, Yunfen Geng, Xiaoshuang Li, Jiabo Hao, Chunming Yuan. Effects of habitat fragmentation on population structure and dynamics of the endangered plant Pterospermum kingtungense [J]. Biodiv Sci, 2021, 29(4): 449-455. |
| [10] | Tianming Wang, Limin Feng, Haitao Yang, Lei Bao, Hongfang Wang, Jianping Ge. An introduction to Long-term Tiger-Leopard Observation Network based on camera traps in Northeast China [J]. Biodiv Sci, 2020, 28(9): 1059-1066. |
| [11] | Xueyou Li, Wenqiang Hu, Changzhe Pu, Quan Li, Qiupeng Yu, Zhechang Hu, William V. Bleisch, Xuelong Jiang. Camera-trapping monitoring platform for mammals and pheasants in the Longitudinal Range and Gorge Region of Southwest China: Protocol, progress and future outlook [J]. Biodiv Sci, 2020, 28(9): 1090-1096. |
| [12] | Yanlin Liu, Dazhao Song, Beibei Liu, Fan Xia, Yuelong Chen, Yiqing Wang, Qiaowen Huang. Overview of the Camera-trapping Platform for Felid Species in China: Data integration by a conservation NGO [J]. Biodiv Sci, 2020, 28(9): 1067-1074. |
| [13] | Sheng Li, William J. McShea, Dajun Wang, Xiaoli Shen, Hongliang Bu, Tianpei Guan, Fang Wang, Xiaodong Gu, Xiaofeng Zhang, Haohong Liao. Construction progress of the Camera-trapping Network for the Mountains of Southwest China [J]. Biodiv Sci, 2020, 28(9): 1049-1058. |
| [14] | Zhifeng Xu, Wen Zhong, Dongkang Zhang, Hongying Hu. Diversity of butterfly communities in Jimusaer County, Xinjiang [J]. Biodiv Sci, 2020, 28(8): 993-1002. |
| [15] | Fang Hong, Ying Xiang, Chaoyang Chen, Liangxian Sun, Chunshou Luo, Guofang Jiang. Community diversity and faunal composition of butterflies in the Longqishan Nature Reserve [J]. Biodiv Sci, 2020, 28(8): 1003-1007. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn