



Biodiv Sci ›› 2026, Vol. 34 ›› Issue (1): 25369. DOI: 10.17520/biods.2025369 cstr: 32101.14.biods.2025369
• Special Feature: Methods for Ecological Data Analysis • Previous Articles Next Articles
Ziling Yan1,2( ), Xiaoyu Chen2(
), Xiaoyu Chen2( ), Meng Yao1,2,*(
), Meng Yao1,2,*( )(
)( )
)
Received:2025-09-15
Accepted:2025-12-09
Online:2026-01-20
Published:2026-01-21
Contact:
Meng Yao
Supported by:Ziling Yan, Xiaoyu Chen, Meng Yao. A comparative evaluation of bioinformatic pipelines for invertebrate biodiversity profiling via environmental DNA metabarcoding[J]. Biodiv Sci, 2026, 34(1): 25369.
| 处理流程 Pipeline | MOTU (Reads) | 门 Phylum | 纲 Class | 目 Order | 科 Family | 属 Genus | 种 Species | |
|---|---|---|---|---|---|---|---|---|
| UPARSE | BOLDigger | 1,548 (1,475,851) | 13 | 27 | 71 | 141 | 108 | 111 |
| BLASTN | 984 (1,349,864) | 11 | 22 | 46 | 107 | 92 | 98 | |
| Bayesian | 1,391 (1,689,419) | 11 | 19 | 34 | 66 | 71 | 81 | |
| Swarm | BOLDigger | 2,142 (1,765,422) | 14 | 31 | 81 | 148 | 109 | 112 |
| BLASTN | 1,541 (1,680,656) | 11 | 22 | 47 | 108 | 94 | 104 | |
| Bayesian | 2,207 (2,095,227) | 11 | 19 | 34 | 67 | 72 | 82 | |
| UNOISE | BOLDigger | 2,392 (1,950,275) | 14 | 31 | 81 | 153 | 108 | 111 |
| BLASTN | 1,567 (1,797,455) | 11 | 22 | 47 | 108 | 94 | 104 | |
| Bayesian | 2,158 (2,264,348) | 11 | 19 | 34 | 67 | 73 | 82 | |
| DADA2 | BOLDigger | 2,777 (3,329,666) | 14 | 32 | 83 | 164 | 118 | 120 |
| BLASTN | 1,818 (2,678,329) | 11 | 23 | 49 | 113 | 102 | 108 | |
| Bayesian | 2,683 (3,690,943) | 11 | 19 | 35 | 71 | 78 | 87 | |
Table 1 Invertebrate MOTU, reads and taxonomic coverage obtained from 12 pipelines. The taxonomic coverage here refers to the coverage breadth of the outputs from different pipelines across each taxonomic level.
| 处理流程 Pipeline | MOTU (Reads) | 门 Phylum | 纲 Class | 目 Order | 科 Family | 属 Genus | 种 Species | |
|---|---|---|---|---|---|---|---|---|
| UPARSE | BOLDigger | 1,548 (1,475,851) | 13 | 27 | 71 | 141 | 108 | 111 |
| BLASTN | 984 (1,349,864) | 11 | 22 | 46 | 107 | 92 | 98 | |
| Bayesian | 1,391 (1,689,419) | 11 | 19 | 34 | 66 | 71 | 81 | |
| Swarm | BOLDigger | 2,142 (1,765,422) | 14 | 31 | 81 | 148 | 109 | 112 |
| BLASTN | 1,541 (1,680,656) | 11 | 22 | 47 | 108 | 94 | 104 | |
| Bayesian | 2,207 (2,095,227) | 11 | 19 | 34 | 67 | 72 | 82 | |
| UNOISE | BOLDigger | 2,392 (1,950,275) | 14 | 31 | 81 | 153 | 108 | 111 |
| BLASTN | 1,567 (1,797,455) | 11 | 22 | 47 | 108 | 94 | 104 | |
| Bayesian | 2,158 (2,264,348) | 11 | 19 | 34 | 67 | 73 | 82 | |
| DADA2 | BOLDigger | 2,777 (3,329,666) | 14 | 32 | 83 | 164 | 118 | 120 |
| BLASTN | 1,818 (2,678,329) | 11 | 23 | 49 | 113 | 102 | 108 | |
| Bayesian | 2,683 (3,690,943) | 11 | 19 | 35 | 71 | 78 | 87 | |
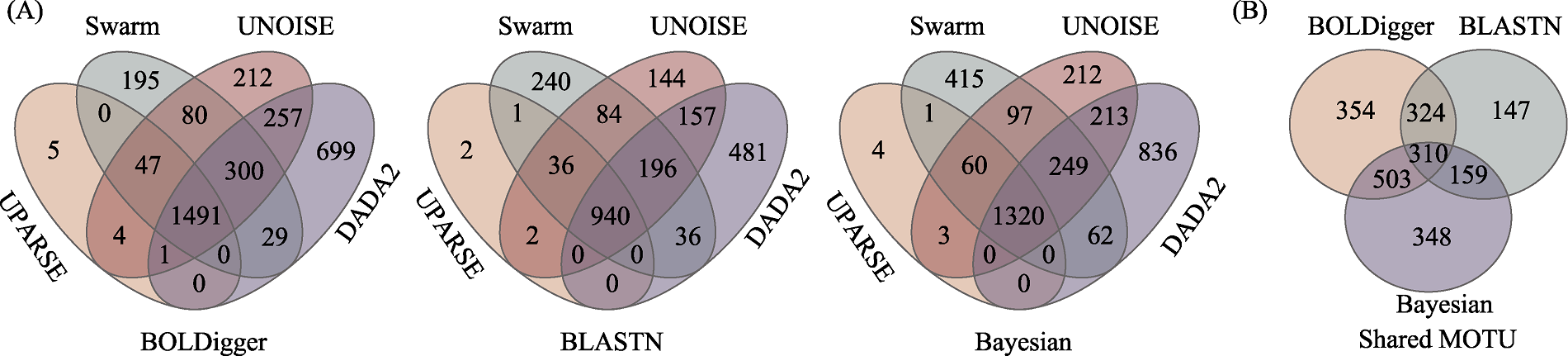
Fig. 3 Venn diagrams of the number of invertebrate MOTU detected using different clustering or denoising methods (A) and the comparison of MOTU jointly detected using different clustering or denoising methods across different taxonomic assignment methods (B)
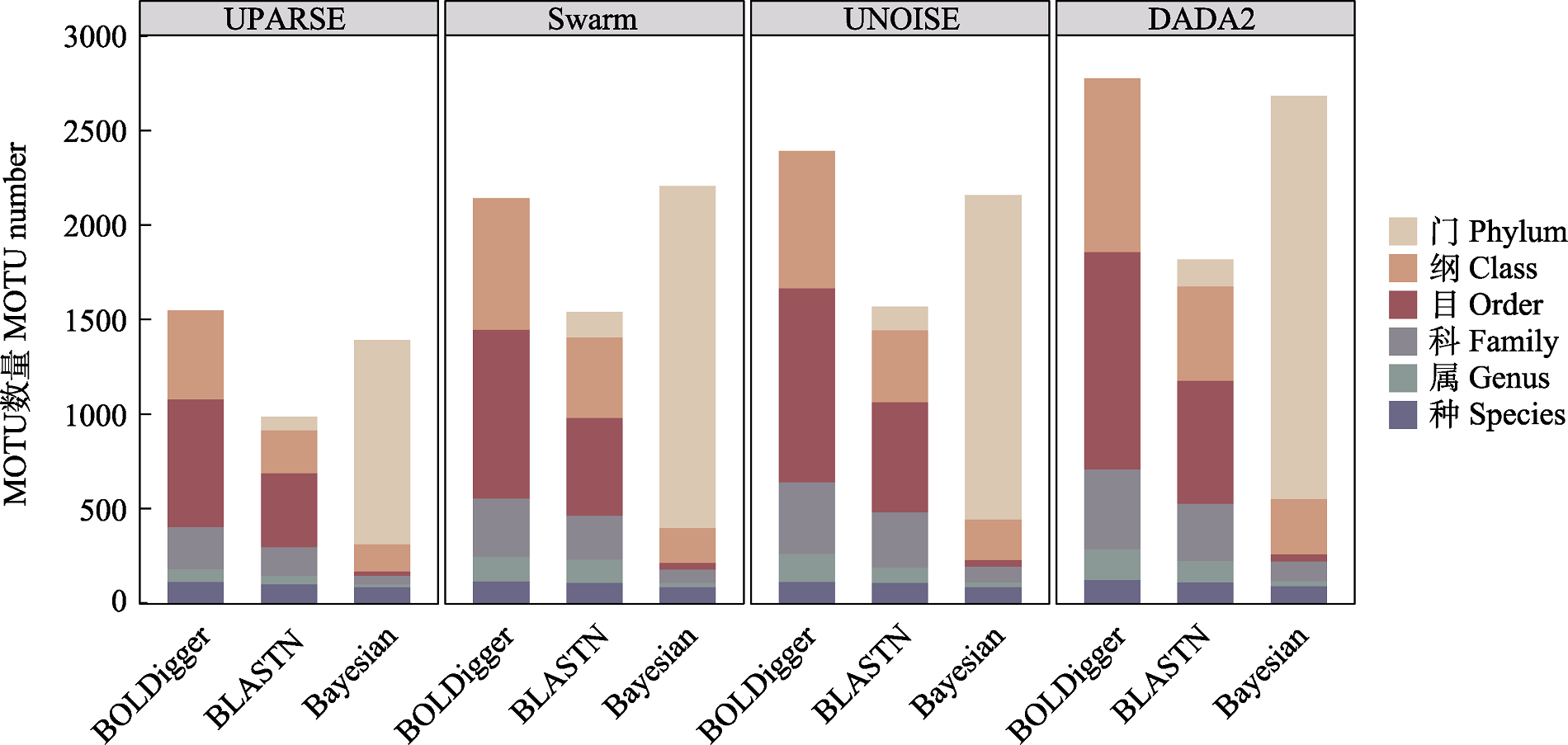
Fig. 4 The taxonomic resolution of invertebrate MOTU detected using different pipelines. The taxonomic resolution here refers to the taxonomic precision of the identification results from different pipelines.
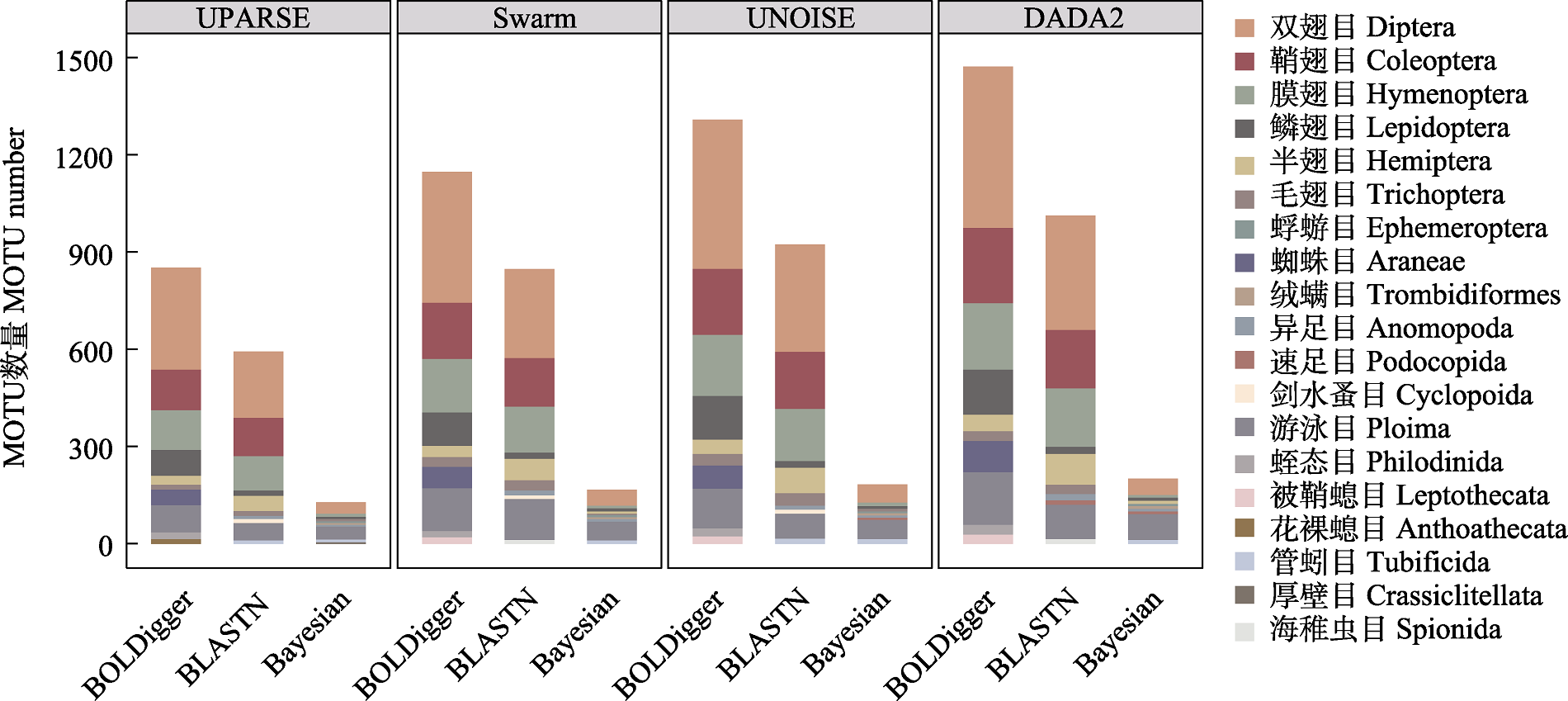
Fig. 7 The number of MOTU belong to major invertebrate orders detected using different pipelines. Only the top 10 orders with the largest number of MOTU are shown.
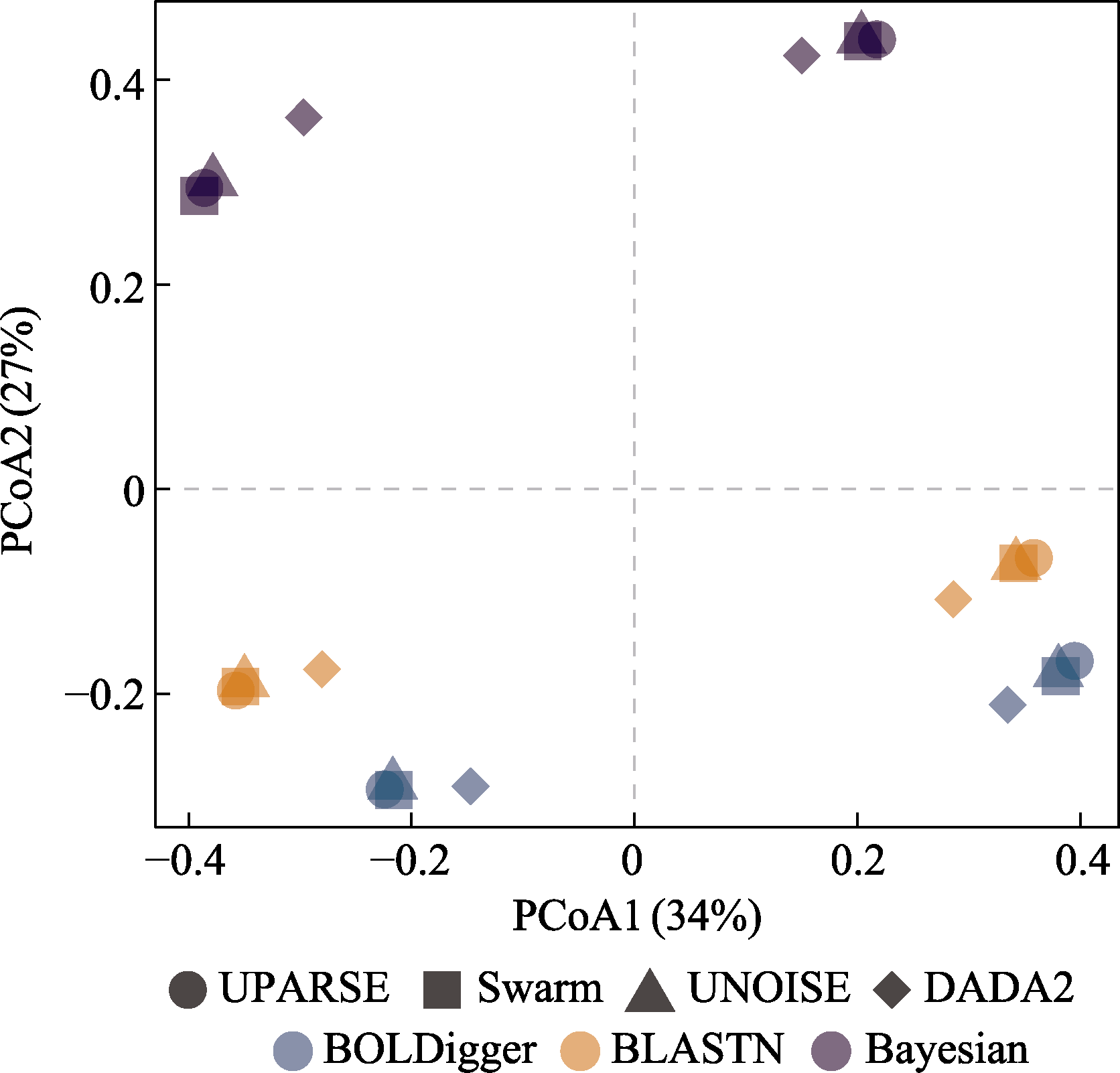
Fig. 8 PCoA analysis of invertebrate MOTU detected using different pipelines based on Jaccard dissimilarity index. The symbols with the same shape and color correspond to the results of eDNA samples from rivers and lakes.
| [1] |
Alberdi A, Aizpurua O, Gilbert MTP, Bohmann K (2018) Scrutinizing key steps for reliable metabarcoding of environmental samples. Methods in Ecology and Evolution, 9, 134-147.
DOI URL |
| [2] |
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic Local Alignment Aearch Tool. Journal of Molecular Biology, 215, 403-410.
DOI PMID |
| [3] |
Antich A, Palacin C, Wangensteen OS, Turon X (2021) To denoise or to cluster, that is not the question: Optimizing pipelines for COI metabarcoding and metaphylogeography. BMC Bioinformatics, 22, 177.
DOI PMID |
| [4] | Bayer PE, Bennett A, Nester G, Corrigan S, Raes EJ, Cooper M, Ayad ME, McVey P, Kardailsky A, Pearce J, Fraser MW, Goncalves P, Burnell S, Rauschert S (2025) A comprehensive evaluation of taxonomic classifiers in marine vertebrate eDNA studies. Molecular Ecology Resources, 25, e14107. |
| [5] |
Bhat AH, Prabhu P, Balakrishnan K (2019) A critical analysis of state-of-the-art metagenomics OTU clustering algorithms. Journal of Biosciences, 44, 148.
DOI |
| [6] |
Bokulich NA, Kaehler BD, Rideout JR, Dillon M, Bolyen E, Knight R, Huttley GA, Caporaso JG (2018) Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome, 6, 90.
DOI |
| [7] | Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, …, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang YL, Zhu QY, Knight R, Caporaso JG (2019) Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology, 37, 852-857. |
| [8] |
Boyer F, Mercier C, Bonin A, Le Bras Y, Taberlet P, Coissac E (2016) Obitools: A unix-inspired software package for DNA metabarcoding. Molecular Ecology Resources, 16, 176-182.
DOI PMID |
| [9] |
Brandt MI, Trouche B, Quintric L, Günther B, Wincker P, Poulain J, Arnaud-Haond S (2021) Bioinformatic pipelines combining denoising and clustering tools allow for more comprehensive prokaryotic and eukaryotic metabarcoding. Molecular Ecology Resources, 21, 1904-1921.
DOI PMID |
| [10] | Buchner D, Leese F (2020) BOLDigger—A Python package to identify and organise sequences with the Barcode of Life Data systems. Metabarcoding and Metagenomics, 4, e53535. |
| [11] |
Bunholi IV, Foster NR, Casey JM (2023) Environmental DNA and RNA in aquatic community ecology: Toward methodological standardization. Environmental DNA, 5, 1133-1147.
DOI URL |
| [12] |
Cahyani NKD, Anggoro AW, Al Malik MD, Subhan B, Sani LMI, Madduppa H (2024) Inventorizing marine biodiversity using eDNA data from Indonesian coral reefs: Comparative high throughput analysis using different bioinformatic pipelines. Marine Biodiversity, 54, 39.
DOI |
| [13] |
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: High-resolution sample inference from Illumina amplicon data. Nature Methods, 13, 581-583.
DOI PMID |
| [14] |
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: Architecture and applications. BMC Bioinformatics, 10, 421.
DOI PMID |
| [15] |
Chen XY, Yan ZL, Li S, Yao M (2025) Advancing aquatic biodiversity assessments of invertebrates using eDNA metabarcoding: A systematic evaluation of primers for marine and freshwater communities. Methods in Ecology and Evolution, 16, 2408-2430.
DOI URL |
| [16] |
Chernyshev AV, Kajihara H (2019) Comparative muscular morphology in archinemertea (Nemertea: Palaeonemertea). Zoomorphology, 138, 193-207.
DOI |
| [17] |
Coissac E (2012) OligoTag: A program for designing sets of tags for next-generation sequencing of multiplexed samples. Methods in Molecular Biology, 888, 13-31.
DOI PMID |
| [18] |
Cruaud P, Vigneron A, Fradette MS, Charette SJ, Rodriguez MJ, Dorea CC, Culley AI (2017) Open the SterivexTM casing: An easy and effective way to improve DNA extraction yields. Limnology and Oceanography: Methods, 15, 1015-1020.
DOI URL |
| [19] | De Grave S, Pentcheff ND, Ahyong ST, Chan TY, Crandall KA, Dworschak PC, Felder DL, Feldmann RM, Fransen CHJM, Goulding LYD, Lemaitre R, Low MEY, Martin JW, Ng PKL, Schweitzer CE, Tan SH, Tshudy D, Wetzer R (2009) A classification of living and fossil genera of decapod crustaceans. Raffles Bulletin of Zoology, 21, 1-109. |
| [20] |
De Santiago A, Pereira TJ, Mincks SL, Bik HM (2022) Dataset complexity impacts both MOTU delimitation and biodiversity estimates in eukaryotic 18S rRNA metabarcoding studies. Environmental DNA, 4, 363-384.
DOI URL |
| [21] |
Deiner K, Bik HM, Mächler E, Seymour M, Lacoursière- Roussel A, Altermatt F, Creer S, Bista I, Lodge DM, de Vere N, Pfrender ME, Bernatchez L (2017) Environmental DNA metabarcoding: Transforming how we survey animal and plant communities. Molecular Ecology, 26, 5872-5895.
DOI PMID |
| [22] |
Dumont HJ, Munuswamy N (1997) The potential of freshwater Anostraca for technical applications. Hydrobiologia, 358, 193-197.
DOI |
| [23] |
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics, 26, 2460-2461.
DOI PMID |
| [24] |
Edgar RC (2013) UPARSE: Highly accurate OTU sequences from microbial amplicon reads. Nature Methods, 10, 996-998.
DOI PMID |
| [25] | Edgar RC (2016) UNOISE2: Improved error-correction for Illumina 16S and ITS amplicon sequencing. bioRxiv preprint. https://doi.org/10.1101/081257. (accessed on 2025-07-13) |
| [26] | Eisenhauer N, Hines J (2021) Invertebrate biodiversity and conservation. Current Biology, 31, R1214-R1218. |
| [27] |
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R (1994) DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology, 3, 294-299.
PMID |
| [28] | Hakimzadeh A, Asbun AA, Albanese D, Bernard M, Buchner D, Callahan B, Caporaso JG, Curd E, Djemiel C, Durling MB, Elbrecht V, Gold Z, Gweon HS, Hajibabaei M, Hildebrand F, Mikryukov V, Normandeau E, Özkurt E, Palmer JM, Pascal G, Porter TM, Straub D, Vasar M, Větrovský T, Zafeiropoulos H, Anslan S (2024) A pile of pipelines: An overview of the bioinformatics software for metabarcoding data analyses. Molecular Ecology Resources, 24, e13847. |
| [29] |
Hao XL, Jiang R, Chen T (2011) Clustering 16S rRNA for OTU prediction: A method of unsupervised Bayesian clustering. Bioinformatics, 27, 611-618.
DOI PMID |
| [30] | Jackson MA, Bell JT, Spector TD, Steves CJ (2016) A heritability-based comparison of methods used to cluster 16S rRNA gene sequences into operational taxonomic units. PeerJ, 4, e2341. |
| [31] | Kopylova E, Navas-Molina JA, Mercier C, Xu ZZ, Mahé F, He Y, Zhou H-W, Rognes T, Caporaso JG, Knight R (2016) Open-source sequence clustering methods improve the state of the art. mSystems, 1, e00003-15. |
| [32] |
Leray M, Yang JY, Meyer CP, Mills SC, Agudelo N, Ranwez V, Boehm JT, Machida RJ (2013) A new versatile primer set targeting a short fragment of the mitochondrial COI region for metabarcoding metazoan diversity: Application for characterizing coral reef fish gut contents. Frontiers in Zoology, 10, 34.
DOI PMID |
| [33] | Li M, Wei TT, Shi BY, Hao XY, Xu HG, Sun HY (2019) Biodiversity monitoring of freshwater benthic macroinvertebrates using environmental DNA. Biodiversity Science, 27, 480-490. (in Chinese with English abstract) |
|
[李萌, 尉婷婷, 史博洋, 郝希阳, 徐海根, 孙红英 (2019) 环境DNA技术在淡水底栖大型无脊椎动物多样性监测中的应用. 生物多样性, 27, 480-490.]
DOI |
|
| [34] |
Li ZY, Zhao WC, Jiang Y, Wen YJ, Li M, Liu L, Zou KS (2024) New insights into biologic interpretation of bioinformatic pipelines for fish eDNA metabarcoding: A case study in Pearl River estuary. Journal of Environmental Management, 368, 122136.
DOI URL |
| [35] |
Lu Q, Zhang S-Y, Du JQ, Liu Q, Dong CX, Zhao JD, Wang YF, Yao M (2023) Multi-group biodiversity distributions and drivers of metacommunity organization along a glacial-fluvial-limnic pathway on the Tibetan Plateau. Environmental Research, 220, 115236.
DOI URL |
| [36] | Lynn DH (2008) The Ciliated Protozoa: Characterization, Classification, and Guide to the Literature, 3rd edn. Springer, New York. |
| [37] | Macé B, Hocdé R, Marques V, Guerin PE, Valentini A, Arnal V, Pellissier L, Manel S (2022) Evaluating bioinformatics pipelines for population-level inference using environmental DNA. Environmental DNA, 4, 674-686. |
| [38] |
Macher T-H, Beermann AJ, Arle J, Foerster J, Greyer M, Mora D, Koschorreck J, Rolauffs P, Rother A, Schüler S, Zimmermann J, Hering D, Leese F (2025) Fit for purpose? Evaluating benthic invertebrate DNA metabarcoding for ecological status class assessment in streams under the Water Framework Directive. Water Research, 272, 122987.
DOI URL |
| [39] |
Mahé F, Czech L, Stamatakis A, Quince C, de Vargas C, Dunthorn M, Rognes T (2021) Swarm v3: Towards tera-scale amplicon clustering. Bioinformatics, 38, 267-269.
DOI PMID |
| [40] |
Malica J, Rączka G, Turczański K, Andrzejewska A, Skorupski M, Urbanowski CK, Kamczyc J (2024) Impact of land use history and soil properties on soil mite communities (Acari, Mesostigmata) inhabiting stands growing on post-agricultural land. Land Degradation & Development, 35, 1776-1791.
DOI URL |
| [41] | Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet Journal, 17, 10-12. |
| [42] |
Mathon L, Valentini A, Guérin PE, Normandeau E, Noel C, Lionnet C, Boulanger E, Thuiller W, Bernatchez L, Mouillot D, Dejean T, Manel S (2021) Benchmarking bioinformatic tools for fast and accurate eDNA metabarcoding species identification. Molecular Ecology Resources, 21, 2565-2579.
DOI PMID |
| [43] | Mercier C, Boyer F, Bonin A, Coissac É (2013) SUMATRA and SUMACLUST: Fast and exact comparison and clustering of sequences. |
| [44] |
Múrria C, Wangensteen OS, Somma S, Väisänen L, Fortuño P, Arnedo MA, Prat N (2024) Taxonomic accuracy and complementarity between bulk and eDNA metabarcoding provides an alternative to morphology for biological assessment of freshwater macroinvertebrates. Science of the Total Environment, 935, 173243.
DOI URL |
| [45] | Nearing JT, Douglas GM, Comeau AM, Langille MGI (2018) Denoising the Denoisers: An independent evaluation of microbiome sequence error-correction approaches. PeerJ, 6, e5364. |
| [46] | Nilsen T, Snipen L-G, Angell IL, Keeley NB, Majaneva S, Pettersen R, Rudi K (2024) Swarm and UNOISE outperform DADA2 and Deblur for denoising high-diversity marine seafloor samples. ISME Communications, 4, ycae071. |
| [47] | Pawłowski J, Apothéloz-Perret-Gentil L, Mächler E, Altermatt F (2020) Apothéloz-Perret-Gentil L, Mächler E, Altermatt F (2020) Environmental DNA applications for biomonitoring and bioassessment in aquatic ecosystems. Guidelines. Federal Office for the Environment, Bern. Environmental Studies. no. 2010: 71 pp. https://doi.org/10.5167/uzh-187800. (accessed on 2025-07-26) |
| [48] |
Petit-Marty N, Casas L, Saborido-Rey F (2023) State-of-the-art of data analyses in environmental DNA approaches towards its applicability to sustainable fisheries management. Frontiers in Marine Science, 10, 1061530.
DOI URL |
| [49] | R Core Team (2023) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. (accessed on 2025-09-04) |
| [50] | Ratnasingham S, Hebert PDN (2007) BOLD: The Barcode of Life Data System (http://www.barcodinglife.org). Molecular Ecology Notes, 7, 355-364. |
| [51] | Rogers DC (2013) Anostraca Catalogus (Crustacea: Branchiopoda). The Raffles Bulletin of Zoology, 61, 525-546. |
| [52] | Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: A versatile open source tool for metagenomics. PeerJ, 4, e2584. |
| [53] |
Rosen GL, Reichenberger ER, Rosenfeld AM (2011) NBC: The Naïve Bayes Classification tool webserver for taxonomic classification of metagenomic reads. Bioinformatics, 27, 127-129.
DOI PMID |
| [54] |
Sfecci E, Lacour T, Amade P, Mehiri M, Sfecci E, Lacour T, Amade P, Mehiri M (2016) Polycyclic guanidine alkaloids from Poecilosclerida marine sponges. Marine Drugs, 14, 77.
DOI URL |
| [55] | Tekle YI, Smith AR, McGinnis M, Ghebezadik S, Patel P (2025) A new Paramoeba isolate from Florida exhibits a microtubule-bound endosymbiont closely associated with the host nucleus. Journal of Eukaryotic Microbiology, 72, e70011. |
| [56] |
Turon X, Antich A, Palacín C, Præbel K, Wangensteen OS (2020) From metabarcoding to metaphylogeography: Separating the wheat from the chaff. Ecological Applications, 30, e02036.
DOI URL |
| [57] |
Vourka A, Karaouzas I, Parmakelis A (2023) River benthic macroinvertebrates and environmental DNA metabarcoding: A scoping review of eDNA sampling, extraction, amplification and sequencing methods. Biodiversity and Conservation, 32, 4221-4238.
DOI |
| [58] |
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Applied and Environmental Microbiology, 73, 5261-5267.
DOI URL |
| [1] | Yun’ao Li, Wenfu Zhang, Guigang Zhao, Chunyan Yang, Xiangqing Chen, Shengdong Yuan, Min Cao, Wang Cai, Jie Yang. Application of airborne eDNA for terrestrial animal diversity monitoring: A case study of 20-ha forest dynamics plot in Xishuangbanna, Yunnan, China [J]. Biodiv Sci, 2025, 33(6): 24318-. |
| [2] | Wen Peng, Zeshuai Deng, Wenbao Zheng, Lingxuan Gong, Yufeng Zeng, Hao Meng, Jun Chen, Daode Yang. Application of eDNA technology in amphibian surveys: A case study of Hunan Mangshan National Nature Reserve [J]. Biodiv Sci, 2025, 33(6): 24552-. |
| [3] | Liu Zhiyu, Ji Xin, Sui Guohui, Yang Ding, Li Xuankun. Invertebrate diversity in buffalo grass and weedy lawns at Beijing Capital International Airport [J]. Biodiv Sci, 2025, 33(4): 24456-. |
| [4] | Yixiu Kou, Zhaohong Weng, Fenfen Ji, Kit Yue Kwan, Yangjie Xie, Jiaqiao Wang, Hangzhao Pan, Yunting Zhao, Kun Ye. Applications of environmental DNA techniques in monitoring endangered aquatic animals [J]. Biodiv Sci, 2025, 33(11): 24574-. |
| [5] | Yihui Jiang, Yue Liu, Xu Zeng, Zheying Lin, Nan Wang, Jihao Peng, Ling Cao, Cong Zeng. Fish diversity and connectivity in six national marine protected areas in the East China Sea [J]. Biodiv Sci, 2024, 32(6): 24128-. |
| [6] | Zhiyuan Dong, Linlin Chen, Naipeng Zhang, Li Chen, Debin Sun, Yanmei Ni, Baoquan Li. Response of fish diversity to hydrological connectivity of typical tidal creek system in the Yellow River Delta based on environmental DNA metabarcoding [J]. Biodiv Sci, 2023, 31(7): 23073-. |
| [7] | Keyi Wu, Wenda Ruan, Difeng Zhou, Qingchen Chen, Chengyun Zhang, Xinyuan Pan, Shang Yu, Yang Liu, Rongbo Xiao. Syllable clustering analysis-based passive acoustic monitoring technology and its application in bird monitoring [J]. Biodiv Sci, 2023, 31(1): 22370-. |
| [8] | Liping Liu, Ruifeng Song, Fu Zhang, Xiuxiang Zhang, Guixiang Peng, Zhiyuan Tan. Diversity of endophytic diazotrophs isolated from Oryza alta [J]. Biodiv Sci, 2020, 28(8): 1018-1025. |
| [9] | Li Yiyuan, C. Molik David, E. Pfrender Michael. EPPS, a metabarcoding bioinformatics pipeline using Nextflow [J]. Biodiv Sci, 2019, 27(5): 567-575. |
| [10] | Liu Shanlin. DNA barcoding and emerging reference construction and data analysis technologies [J]. Biodiv Sci, 2019, 27(5): 526-533. |
| [11] | Erhu Gao, Jiekun He, Zhichen Wang, Yang Xu, Xiaoping Tang, Haisheng Jiang. China’s zoogeographical regionalization based on terrestrial vertebrates [J]. Biodiv Sci, 2017, 25(12): 1321-1330. |
| [12] | Chengye Hu, Yuyue Shui, Kuo Tian, Liang Li, Hulin Qin, Chuncao Zhang, Mengmeng Ji, Bonian Shui. Functional group classification and niche identification of major fish species in the Qixing Islands Marine Reserve, Zhejiang Province [J]. Biodiv Sci, 2016, 24(2): 175-184. |
| [13] | Lifu Sun, Kequan Pei, Yanhua Zhang, Jun Zhao, Guoting Yang, Guofu Qin, Yushuang Song, Ruiqing Song. Genetic diversity of Armillaria gallica isolates from China and Europe revealed with ISSR analysis [J]. Biodiv Sci, 2012, 20(2): 224-230. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn