



Biodiv Sci ›› 2014, Vol. 22 ›› Issue (1): 3-20. DOI: 10.3724/SP.J.1003.2014.13170 cstr: 32101.14.SP.J.1003.2014.13170
Special Issue: 基因组和生物多样性; 物种形成与系统进化
• Orginal Article • Previous Articles Next Articles
Limin Lu1,2, Miao Sun1,2, Jingbo Zhang1,2, Honglei Li1,2, Li Lin1,2, Tuo Yang1,2, Min Chen1,2, Zhiduan Chen1,*( )
)
Received:2013-07-22
Accepted:2014-01-06
Online:2014-01-20
Published:2014-02-10
Contact:
Chen Zhiduan
Limin Lu,Miao Sun,Jingbo Zhang,Honglei Li,Li Lin,Tuo Yang,Min Chen,Zhiduan Chen. Tree of life and its applications[J]. Biodiv Sci, 2014, 22(1): 3-20.
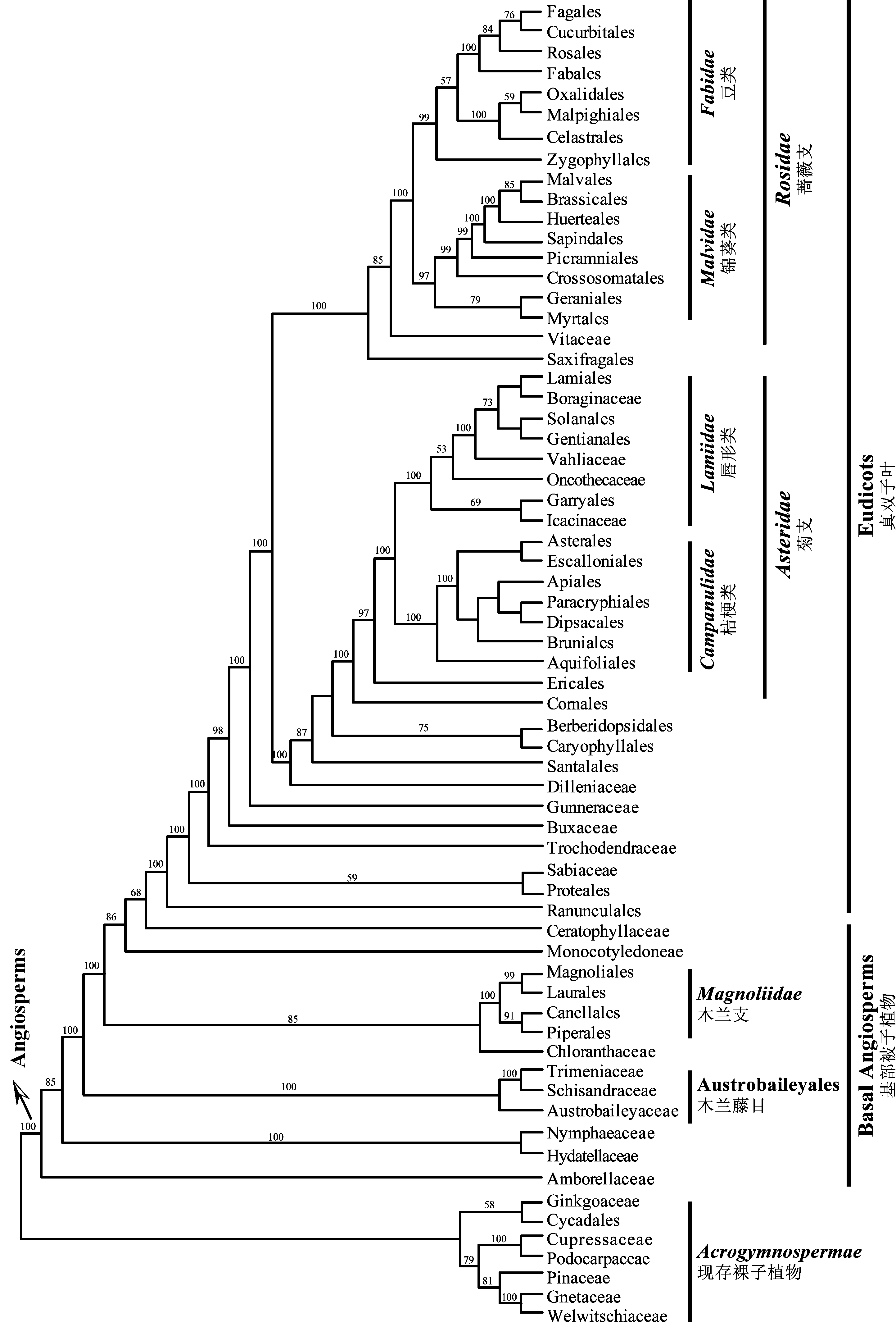
Fig. 1 Phylogenetic relationships of the angiosperms at the ordinal level (based on Soltis et al., 2011). Names of the families, orders and other major clades follow APG III (2009) and Cantino et al. (2007). Numbers above branches are Maximum Likelihood bootstrap values.
| 冲突原因阐述 Description | 解决方案 Solutions | |
|---|---|---|
| 软冲突(假冲突) Soft incongruence | ||
| 人为因素 Artificial factors | 数据不足 Insufficient data | 下一代测序技术; 系统发育基因组学; 取样代表性; 基因选择; 进化模型的选择 Next-generation sequencing technologies; phylogenomics; Taxon sampling; DNA regions selection; selection of substitution models |
| 取样偏差 Biased sampling | ||
| 基因选择不当 Sub-optimized gene selection | ||
| 测序错误 Sequencing errors | ||
| 序列因素 Sequencing factors | 碱基组成成分偏差 Compositional bias | 第三密码子排除; 氨基酸序列建树; RY编码; 快速进化位点移除; 一致网络分析法 3rd codon position exclusion; amino acid tree; RY coding; removing fast-evolving nucleotide sites; consensus networks |
| 长枝吸引 Long-branch attraction | ||
| 进化速率异质性 Evolutionary rate heterogeneity | ||
| 进化饱和 Evolutionary saturation | ||
| 硬冲突 Hard incongruence | ||
| 生物过程 Biological factors | 快速辐射分化 Rapid diversification | 最小遗传距离法; 融合法; 基因树简约法; 网状进化网络分析法 Minimum genetic distance method; coalescence-based methods; gene tree parsimony; reticulation networks |
| 杂交/渐渗 Hybridization/Introgression | ||
| 不完全谱系筛选 Incomplete lineage sorting | ||
| 基因水平转移 Horizontal gene transfer | ||
| 并系基因 Paralogous genes | ||
| 基因重复和/或丢失 Gene duplication and/or gene loss | ||
| 基因重组 Gene recombination |
Table 1 Causes and resolutions for phylogenetic incongruences
| 冲突原因阐述 Description | 解决方案 Solutions | |
|---|---|---|
| 软冲突(假冲突) Soft incongruence | ||
| 人为因素 Artificial factors | 数据不足 Insufficient data | 下一代测序技术; 系统发育基因组学; 取样代表性; 基因选择; 进化模型的选择 Next-generation sequencing technologies; phylogenomics; Taxon sampling; DNA regions selection; selection of substitution models |
| 取样偏差 Biased sampling | ||
| 基因选择不当 Sub-optimized gene selection | ||
| 测序错误 Sequencing errors | ||
| 序列因素 Sequencing factors | 碱基组成成分偏差 Compositional bias | 第三密码子排除; 氨基酸序列建树; RY编码; 快速进化位点移除; 一致网络分析法 3rd codon position exclusion; amino acid tree; RY coding; removing fast-evolving nucleotide sites; consensus networks |
| 长枝吸引 Long-branch attraction | ||
| 进化速率异质性 Evolutionary rate heterogeneity | ||
| 进化饱和 Evolutionary saturation | ||
| 硬冲突 Hard incongruence | ||
| 生物过程 Biological factors | 快速辐射分化 Rapid diversification | 最小遗传距离法; 融合法; 基因树简约法; 网状进化网络分析法 Minimum genetic distance method; coalescence-based methods; gene tree parsimony; reticulation networks |
| 杂交/渐渗 Hybridization/Introgression | ||
| 不完全谱系筛选 Incomplete lineage sorting | ||
| 基因水平转移 Horizontal gene transfer | ||
| 并系基因 Paralogous genes | ||
| 基因重复和/或丢失 Gene duplication and/or gene loss | ||
| 基因重组 Gene recombination |
| [1] | Alfaro ME, Santini F, Brock C, Alamillo H, Dornburg A, Rabosky DL, Carnevale G, Harmon LJ (2009) Nine exceptinoal radiations plus high turnover explain species diversity in jawed vertebrates.Proceedings of the National Academy of Sciences, USA, 106, 13410-13414. |
| [2] | Ali SS, Yu Y, Pfosser M, Wetschnig W (2012) Inferences of biogeographical histories within subfamily Hyacinthoideae using S-DIVA and Bayesian binary MCMC analysis implemented in RASP (Reconstruct Ancestral State in Phylogenies).Annals of Botany, 109, 95-107. |
| [3] | APG (1998) An ordinal classification for the families of flowering plants.Annals of the Missouri Botanical Garden, 85, 531-553. |
| [4] | APG II (2003) An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants APG II.Botanical Journal of the Linnean Society, 141, 399-436. |
| [5] | APG III (2009) An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG III.Botanical Journal of the Linnean Society, 161, 105-121. |
| [6] | Baker WJ, Couvreur TL (2013) Global biogeography and diversification of palms sheds light on the evolution of tropical lineages. I. Historical biogeography.Journal of Biogeography, 40, 274-285. |
| [7] | Benton MJ (2010) The origins of modern biodiversity on land.Philosophical Transactions of the Royal Society B, 365, 3667-3679. |
| [8] | Benton MJ (2013) Origins of biodiversity.Palaeontology, 56, 1-7. |
| [9] | Benson DA, Karsch-Mizarchi I, Lipman DJ, Ostell J, Sayers EW (2011) GenBank.Nucleic Acids Research, 39, D32-D37. |
| [10] | Bininda-Emonds ORP (2004) Phylogenetic Supertrees: Combining Information to Reveal the Tree of Life. Kluwer Academic Publishing, Dordrecht. |
| [11] | Bininda-Emonds ORP, Cardillo M, Jones KE, MacPhee RDE, Beck RMD, Grenyer R, Price SA, Vos RA, Gittleman JL, Purvis A (2007) The delayed rise of present-day mammals.Nature, 446, 507-512. |
| [12] | Blackledge TA, Scharff N, Coddington J, Szűts T, Wenzel J, Hayashi C, Agnarsson I (2009) Reconstructing web evolution and spider diversification in the molecular era.Proceedings of the National Academy of Sciences,USA, 106, 5229-5234. |
| [13] | Bouchenak-Khelladi Y, Verboom GA, Hodkinson TR, Salamin N, Francois O, Chonghaile GN, Savolainen V (2009) The origins and diversification of C4 grasses and savanna-adapted ungulates.Global Change Biology, 15, 2397-2417. |
| [14] | Burbrink FT, Ruane S, Pyron RA (2012) When are adaptive radiations replicated in areas? Ecological opportunity and unexceptional diversification in West Indian dipsadine snakes (Colubridae: Alsophiini).Journal of Biogeography, 39, 465-475. |
| [15] | Burleigh JG, Bansal MS, Eulenstein O, Hartmann S, Wehe A, Vision TJ (2011) Genome-scale phylogenetics: inferring the plant tree of life from 18,896 gene trees.Systematic Biology, 60, 117-125. |
| [16] | Cadotte MW, Jonathan Davies T, Regetz J, Kembel SW, Cleland E, Oakley TH (2010) Phylogenetic diversity metrics for ecological communities: integrating species richness, abundance and evolutionary history.Ecology Letters, 13, 96-105. |
| [17] | Cavender-Bares J, Kozak KH, Fine PV, Kembel SW (2009) The merging of community ecology and phylogenetic biology.Ecology Letters, 12, 693-715. |
| [18] | Chen ZD (陈之端), Li DZ (李德铢) (2013) On Barcode of Life and Tree of Life. Plant Diversity and Resources (植物分类与资源学报), 35, 675-681. (in Chinese with English abstract) |
| [19] | Christin PA, Besnard G, Samaritani E, Duvall MR, Hodkinson TR, Savolainen V, Salamin N (2008) Oligocene CO2 decline promoted C4 photosynthesis in grass.Current Biology, 18, 37-43. |
| [20] | Clegg MT (1993) Chloroplast gene sequences and the study of plant evolution.Proceedings of the National Academy of Sciences,USA, 90, 363-367. |
| [21] | Cook LG, Crisp MD (2005) Not so ancient: the extant crown group of Nothofagus represents a post-Gondwanan radiation.Proceedings of the Royal Society B: Biological Sciences, 272, 2535-2544. |
| [22] | Couvreur TLP, Forest F, Baker WJ (2011) Origin and global diversification patterns of tropical rain forests: inferences from a complete genus-level phylogeny of palms.BMC Biology, 9, 44. |
| [23] | Cracraft J, Donoghue MJ (2004) Assembling the Tree of Life. Oxford University Press, Oxford. |
| [24] | Crisp MD, Trewick SA, Cook LG (2011) Hypothesis testing in biogeography.Trends in Ecology and Evolution, 26, 66-72. |
| [25] | Cronquist A (1988) The Evolution and Classification of Flowering Plants, 2nd edn. New York Botanical Garden, New York. |
| [26] | Cruaud A, Rønsted N, Chantarasuwan B, Chou LS, Clement WL, Couloux A, Cousins B, Genson G, Harrison RD, Hanson PE (2012) An extreme case of plant-insect codiversification: figs and fig-pollinating wasps.Systematic Biology, 61, 1029-1047. |
| [27] | Dahlgren G (1989) An updated angiosperm classification.Botanical Journal of the Linnean Society, 100, 197-203. |
| [28] | Darwin C (1859) On the Origin of Species by Means of Natural Selection, or the Preservation of Favoured Races in the Struggle for Life. John Murray, London. |
| [29] | De-Nova JA, Medina R, Montero JC, Weeks A, Rosell JA, Olson ME, Eguiarte LE, Magallón S (2012) Insights into the historical construction of species-rich Mesoamerican seasonally dry tropical forests: the diversification of Bursera (Burseraceae, Sapindales).New Phytologist, 193, 276-287. |
| [30] | Degnan JH, Rosenberg NA (2009) Gene tree discordance, phylogenetic inference and the multispecies coalescent.Trends in Ecology and Evolution, 24, 332-340. |
| [31] | Delsuc F, Brinkmann FH, Philippe H (2005) Phylogenomics and the reconstruction of the tree of life.Nature Reviews Genetics, 6, 361-375. |
| [32] | De QueirozK, Gauthier J (1992) Phylogenetic taxonomy.Annual Review of Ecology and Systematics, 23, 449-480. |
| [33] | De Queiroz A, Donoghue MJ, Kim J (1995) Separate versus combined analysis of phylogenetic evidence.Annual Review of Ecology and Systematics, 26, 657-681. |
| [34] | De Queiroz A, Gatesy J (2007) The supermatrix approach to systematics.Trends in Ecology and Evolution, 22, 34-41. |
| [35] | Dobzhansky TG (1937) Genetics and the Origin of Species. Columbia University Press, New York. |
| [36] | Donoghue MJ (2008) A phylogenetic perspective on the distribution of plant diversity.Proceedings of the National Academy of Sciences,USA, 105, 11549-11555. |
| [37] | Doolittle WF (1999) Phylogenetic classification and the universal tree.Science, 284, 2124-2128. |
| [38] | Doolittle WF, Bapteste E (2007) Pattern pluralism and the Tree of Life hypothesis.Proceedings of the National Academy of Sciences,USA, 104, 2043-2049. |
| [39] | Driskell AC, Ane C, Burleigh JG, McMahon MM, O’Meara BC, Sanderson MJ (2004) Prospects for building the tree of life from large sequence databases.Science, 306, 1172-1174. |
| [40] | Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees.BMC Evolutionary Biology, 7, 214. |
| [41] | Eaton DAR, Ree RH (2013) Inferring phylogeny and introgression using RADseq Data: an example from flowering plants (Pedicularis: Orobanchaceae).Systematic Biology, 62, 689-706. |
| [42] | Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput.Nucleic Acids Research, 32, 1792-1797. |
| [43] | Faith DP (1992) Conservation evaluation and phylogenetic diversity.Biological Conservation, 61, 1-10. |
| [44] | Felsenstein J (1978) Cases in which parsimony or compatibility methods will be positively misleading.Systematic Zoology, 27, 401-410. |
| [45] | Fishbein M, Hibsch-Jetter C, Soltis DE, Hufford L (2001) Phylogeny of Saxifragales (Angiosperms, Eudicots): analysis of a rapid, ancient radiation.Systematic Biology, 50, 817-847. |
| [46] | Fitch WM, Margoliash E (1967) Construction of phylogenetic trees.Science, 155, 279-284. |
| [47] | Fitch WM (1971) Toward defining the course of evolution: minimum change for a specific tree topology.Systematic Zoology, 20, 406-416. |
| [48] | FitzJohn RG, Maddison WP, Otto SP (2009) Estimating trait-dependent speciation and extinction rates from incompletely resolved phylogenies.Systematic Biology, 58, 595-611. |
| [49] | Forest F, Grenyer R, Rouget M, Davies TJ, Cowling RM, Faith DP, Balmford A, Manning JC, Proches S, der Bank M, Reeves G, Hedderson TAJ, Savolainen V (2007) Preserving the evolutionary potential of floras in biodiversity hotspots.Nature, 445, 757-760. |
| [50] | Francis AP, Currie DJ (2003) A globally consistent richness-climate relationship for angiosperms.The American Naturist, 161, 323-336. |
| [51] | Fu J (2000) Toward the phylogeny of the family Lacertidae—why 4708 base pairs of mtDNA sequences cannot draw the picture.Biological Journal of the Linnean Society, 71, 203-217. |
| [52] | Galtier N, Daubin V (2008) Dealing with incongruence in phylogenomic analyses.Philosophical Transactions of the Royal Society B: Biological Sciences, 363, 4023-4029. |
| [53] | Gatesy J, Geisler JH, Chang J, Buell C, Berta A, Meredith RW, Springer MS, McGowen MR (2013) A phylogenetic blueprint for a modern whale.Molecular Phylogenetics and Evolution, 66, 479-506. |
| [54] | Ge XJ (葛学军) (2013) Community phylogeny and DNA barcoding. In: Frontier in Plant DNA Barcoding (植物DNA条形码前沿探讨) (ed. Shen AM (沈爱民)), pp. 11-21. China Science and Technology Press, Beijing. (in Chinese) |
| [55] | Gehrke B, Linder HP (2011) Time, space and ecology: why some clades have more species than others.Journal of Biogeography, 38, 1948-1962. |
| [56] | Goldberg EE, Lancaster LT, Ree RH (2011) Phylogenetic inference of reciprocal effects between geographic range evolution and diversification.Systematic Biology, 60, 451-465. |
| [57] | Graham CH, Fine PV (2008) Phylogenetic beta diversity: linking ecological and evolutionary processes across space in time.Ecology Letters, 11, 1265-1277. |
| [58] | Guschanski K, Krause J, Sawyer S, Valente LM, Bailey S, Finstermeier K, Sabin R, Gilissen E, Sonet G, Nagy ZT, Lenglet G, Mayer F, Savolainen V (2013) Next-generation museomics disentangles one of the largest primate radiations.Systematic Biology, 62, 539-554. |
| [59] | Haffer J (1969) Speciation in Amazonian forest birds.Science, 165, 131-137. |
| [60] | Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT.Nucleic Acids Symposium Series, 41, 95-98. |
| [61] | Hall BG (2005) Comparison of the accuracies of several phylogenetic methods using protein and DNA sequences.Molecular Biology and Evolution, 22, 792-802. |
| [62] | Hall BG (2008) Phylogenetic Trees Made Easy. Sunderland, Mariland. |
| [63] | Harris A, Xiang QYJ (2009) Estimating ancestral distributions of lineages with uncertain sister groups: a statistical approach to Dispersal-Vicariance Analysis and a case using Aesculus L. (Sapindaceae) including fossils.Journal of Systematics and Evolution, 47, 349-368. |
| [64] | Hartmann K, André J (2013) Should evolutionary history guide conservation?Biodiversity and Conservation, 22, 449-458. |
| [65] | Hawksworth DL (1996) Biodiversity: Mesurement and Estimation. Springer, New York. |
| [66] | Hennig W (1950) Grundzügeeiner Theorie der Phylogenetischen Systematik. Deutscher Zentralverlag, Berlin. |
| [67] | Hillis DM, Huelsenbeck JP, Cunningham CW (1994) Application and accuracy of molecular phylogenies.Science-AAAS-Weekly Paper Edition-including Guide to Scientific Information, 264, 671-676. |
| [68] | Hodges SA, Derieg NJ (2009) Adaptive radiations: from field to genomic studies.Proceedings of the National Academy of Sciences,USA, 106, 9947-9954. |
| [69] | Hodkinson TR, Parnell JAN (2006) Reconstructing the Tree of Life: Taxonomy and Systematics of Species Rich Taxa. CRC Press, Boca Raton. |
| [70] | Hoorn C, Wesselingh FP, ter Steege H, Bermudez MA, Mora A, Sevink J, Sanmartín I, Sanchez-Meseguer A, Anderson CL, Figueiredo J, Jaramillo C, Riff D, Negri FR, Hooghiemstra H, Lundberg J, Stadler T, Sarkinen T, Antonelli A (2010) Amazonia through time: Andean uplift, climate change, landscape evolution, and biodiversity.Science, 330, 927-931. |
| [71] | Hou Z, Sket B, Fiser C, Li S (2011) Eocene habitat shift from saline to freshwater promoted Tethyan amphipod diversification.Proceedings of the National Academy of Sciences,USA, 108, 14533-14538. |
| [72] | Huang JH, Chen B, Liu CR, Lai JS, Zhang JL, Ma KP (2012) Identifying hotspots of endemic woody seed plant diversity in China.Diversity and Distributions, 18, 673-688. |
| [73] | Isaac NJ, Turvey ST, Collen B, Waterman C, Baillie JE (2007) Mammals on the EDGE: conservation priorities based on threat and phylogeny.PLoS ONE, 2, e296. |
| [74] | Jetz W, Thomas GH, Joy JB, Hartmann K, Mooers AO (2012) The global diversity of birds in space and time.Nature, 491, 444-448. |
| [75] | Jian S, Soltis PS, Gitzendanner MA, Moore MJ, Li RQ, Hendry TA, Qiu YL, Dhingra A, Bell CD, Soltis DE (2008) Resolving an ancient, rapid radiation in Saxifragales.Systematic Biology, 57, 38-57. |
| [76] | Jordan GJ, Hill RS (1999) The phylogenetic affinities of Nothofagus (Nothofagaceae) leaf fossils based on combined molecular and morphological data.International Journal of Plant Sciences, 160, 1177-1188. |
| [77] | Katoh K, MisawaK, Kuma K, MiyataT (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform.Nucleic Acids Research, 30, 3059-3066. |
| [78] | Kearney M, Clark JM (2003) Problems due to missing data in phylogenetic analyses including fossils: a critical review.Journal of Vertebrate Paleontology, 23, 263-274. |
| [79] | Kembel SW, Hubbell SP (2006) The phylogenetic structure of a neotropical forest tree community.Ecology, 87, S86-S99. |
| [80] | Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD, Blomberg SP, Webb CO (2010) Picante: R tools for integrating phylogenies and ecology.Bioinformatics, 26, 1463-1464. |
| [81] | Knapp M, Stöckler K, Havell D, Delsuc F, Sebastiani F, Lockhart PJ (2005) Relaxed molecular clock provides evidence for long-distance dispersal of Nothofagus (southern beech).PLoS Biology, 3, e14. |
| [82] | Kodandaramaiah U (2010) Use of dispersal-vicariance analysis in biogeography—a critique.Journal of Biogeography, 37, 3-11. |
| [83] | Kramer EM (2009) Aquilegia: a new model for plant development, ecology, and evolution.Annual Review of Plant Biology, 60, 261-277. |
| [84] | Kress WJ, Erickson DL, Jones FA, Swenson NG, Perez R, Sanjur O, Bermingham E (2009) Plant DNA barcodes and a community phylogeny of a tropical forest dynamics plot in Panama.Proceedings of the National Academy of Sciences,USA, 106, 18621-18626. |
| [85] | Laffan SW, Lubarsky E, Rosauer D F (2010) Biodiverse, a tool for the spatial analysis of biological and related diversity.Ecography, 33, 643-647. |
| [86] | Lamm KS, Redelings BD (2009) Reconstructing ancestral ranges in historical biogeography: properties and prospects.Journal of Systematics and Evolution, 47, 369-382. |
| [87] | Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0.Bioinformatics, 23, 2947-2948. |
| [88] | Lartillot N, Lepage T, Blanquart S (2009) PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating.Bioinformatics, 25, 2286-2288. |
| [89] | Linder HP (2003) The radiation of the Cape flora, southern Africa.Biological Reviews, 78, 597-638. |
| [90] | Linnaeus C (1753) Species Plantarum, Vol. 2. Stockholm, Sweden. |
| [91] | Li RQ, Chen ZD, Lu AM, Soltis DE, Soltis PS, Manos PS (2004) Phylogenetic relationships in Fagales based on DNA sequences from three genomes.International Journal of Plant Sciences, 165, 311-324. |
| [92] | Liu L (2008) BEST: Bayesian estimation of species trees under the coalescent model.Bioinformatics, 24, 2542-2543. |
| [93] | Lu LM, Wang W, Chen ZD, Wen J (2013) Phylogeny of the non-monophyletic Cayratia Juss. (Vitaceae) and implica- tions for character evolution and biogeography.Molecular Phylogenetics and Evolution, 68, 502-515. |
| [94] | Ma KP (马克平) (2013) A mini review on the advancement of biodiversity research in China in 2012.Biodiversity Science(生物多样性), 21, 1-2. (in Chinese) |
| [95] | Mace GM, Gittleman JL, Purvis A (2003) Preserving the tree of life.Science, 300, 1707-1709. |
| [96] | Maddison WP (1997) Gene trees in species trees.Systematic Biology, 46, 523-536. |
| [97] | Maddison WP, Maddison D (2009) Mesquite: a modular system for evolutionary analysis, version 2.74. Available at. |
| [98] | Maddison WP, Midford PE, Otto SP (2007) Estimating a binary character’s effect on speciation and extinction.Systematic Biology, 56, 701-710. |
| [99] | Magallón S, Sanderson MJ (2001) Absolute diversification rates in angiosperm clades.Evolution, 55, 1762-1780. |
| [100] | Magallón S, Castillo A (2009) Angiosperm diversification through time.American Journal of Botany, 96, 349-365. |
| [101] | Manos P (1997) Systematics of Nothofagus (Nothofagaceae) based on rDNA spacer sequences (ITS): taxonomic congruence with morphology and plastid sequences.American Journal of Botany, 84, 1137. |
| [102] | Manos P, Steele K (1997) Phylogenetic analyses of “higher” Hamamelididae based on plastid sequence data.American Journal of Botany, 84, 1407. |
| [103] | Mao LF (毛岭峰) (2013) Difference of Relationships Between Spatial Pattern and Environment for Seed Plant Diversity in China (中国种子植物多样性的空间格局-环境关系分异研究). PhD dissertation, Institute of Botany, Chinese Academy of Sciences, Beijing. (in Chinese with English abstract) |
| [104] | Marjoram P, Tavaré S (2006) Modern computational approaches for analyzing molecular genetic variation data.Nature Reviews Genetics, 7, 759-770. |
| [105] | Martin P, Dowd J (1993) Using sequences of rbcL to study phylogeny and biogeography of Nothofagus species.Australian Systematic Botany, 6, 441-447. |
| [106] | Mau B, Newton MA, Larget B (1999) Bayesian phylogenetic inference via Markov Chain Monte Carlo methods.Biometrics, 55, 1-12. |
| [107] | Maureira-Butler IJ, Pfeil BE, Muangprom A, Osborn TC, Doyle JJ (2008) The reticulate history of Medicago (Fabaceae).Systematic Biology, 57, 466-482. |
| [108] | McMahon MM, Sanderson MJ (2006) Phylogenetic supermatrix analysis of GenBank sequences from 2228 papilionoid legumes.Systematic Biology, 55, 818-836. |
| [109] | Mi XC, Swenson NG, Valencia R, Kress WJ, Erickson DL, Pérez AJ, Ren HB, Su SH, Gunatilleke N, Gunatilleke S, Hao ZQ, Ye WH, Gao M, Suresh HS, Dattaraja HS, Sukumar R, Ma KP (2012) The contribution of rare species to community phylogenetic diversity across a global network of forest plots.The American Naturalist, 180, 17-30. |
| [110] | Mi XC (米湘成), Pei NC (裴男才), Ma KP (马克平) (2013) On phylogenetic community ecology. In: The New Biology Yearbook 2013 (新生物年鉴2013) (ed. Pu MM (蒲慕明)), pp. 266-289. Science Press, Beijing. (in Chinese) |
| [111] | Michener CD, Sokal RR (1957) A quantitative approach to a problem in classification.Evolution, 11, 130-162. |
| [112] | Morlon H, Parsons TL, Plotkin JB (2011) Reconciling molecular phylogenies with the fossil record.Proceedings of the National Academy of Sciences,USA, 108, 16327-16332. |
| [113] | Near TJ, Meylan PA, Shaffer HB (2005) Assessing concordance of fossil calibration points in molecular clock studies: an example using turtles.The American Naturalist, 165, 137-146. |
| [114] | Nee S, May RM, Harvey PH (1994) The reconstructed evolutionary process.Philosophical Transactions of the Royal Society B: Biological Sciences, 344, 305-311. |
| [115] | Nei M, Kumar S (2000) Molecular Evolution and Phylogenetics. Oxford University Press, Oxford. |
| [116] | Nylander JA (2004) MrModeltest. Version 2. Program distributed by the author. Evolutionary Biology Center, Uppsala University, Uppsala. |
| [117] | Okuyama Y, Fujii N, Wakabayashi M, Kawakita A, Ito M, Watanabe M, Murakami N, Kato M (2005) Nonuniform concerted evolution and chloroplast capture: heterogeneity of observed introgression patterns in three molecular data partition phylogenies of Asian Mitella (Saxifragaceae).Molecular Biology and Evolution, 22, 285-296. |
| [118] | O’Leary MA, Bloch JI, Flynn JJ, Gaudin TJ, Giallombardo A, Giannini NP, Goldberg SL, Kraatz BP, Luo Z-X, Meng J, Ni X, Novacek MJ, Perini FA, Randall ZS, Rougier GW, Sargis EJ, Silcox MT, Simmons NB, Spaulding M, Velazco PM, Weksler M, Wible JR, Cirranello AL (2013) The placental mammal ancestor and the post-K-Pg radiation of Placentals.Science, 339, 662-667. |
| [119] | Page RDM (1996) TreeView: an application to display phylogenetic trees on personal computers.Computer Applications in the Biosciences, 12, 357-358. |
| [120] | Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language.Bioinformatics, 20, 289-290. |
| [121] | Pei NC (裴男才) (2012) Building a subtropical forest community phylogeny based on plant DNA barcodes from Dinghushan plot.Plant Diversity and Resources(植物分类与资源学报), 34, 263-270. (in Chinese with English abstract) |
| [122] | Pei NC, Lian JY, Erickson DL, Swenson NG, Kress WJ, Ye WH, Ge XJ (2011) Exploring tree-habitat associations in a Chinese subtropical forest plot using a molecular phylogeny generated from DNA barcode loci.PLoS ONE, 6, e21273. |
| [123] | Pei NC (裴男才), Zhang JL (张金龙), Mi XC (米湘成), Ge XJ (葛学军) (2011) Plant DNA barcodes promote the develop- ment of phylogenetic community ecology.Biodiversity Science(生物多样性), 19, 284-294. (in Chinese with English abstract) |
| [124] | Posada D, Crandall K (1998) MODELTEST: testing the model of DNA substitution.Bioinformatics, 14, 817-818. |
| [125] | Posada D (2008) jModelTest: phylogenetic model averaging.Molecular Biology and Evolution, 25, 1253-1256. |
| [126] | Puzey JR, Gerbode SJ, Hodges SA, Kramer EM, Mahadevan L (2012) Evolution of spur-length diversity in Aquilegia petals is achieved solely through cell-shape anisotropy.Proceedings of the Royal Society B: Biological Sciences, 279, 1640-1645. |
| [127] | Qiu YL, Li LB, Wang B, Xue JY, Hendry TA, Li RQ, Brown JW, Liu Y, Hudson YH, Chen ZD (2010) Angiosperm phylogeny inferred from sequences of four mitochondrial genes.Journal of Systematics and Evolution, 48, 391-425. |
| [128] | Rabosky DL (2006a) Likelihood methods for detecting temporal shifts in diversification rates.Evolution, 60, 1152-1164. |
| [129] | Rabosky DL (2006b) LASER: a maximum likelihood toolkit for detecting temporal shifts in diversification rates from molecular phylogenies.Evolutionary Bioinformatics, 2, 247-250. |
| [130] | Rambaut A (2002) Se-Al: Sequence Alignment Editor. Available at. |
| [131] | Rambaut A (2009) FigTree 1.3.1. Available at. |
| [132] | Rannala B, Yang Z (1996) Probability distribution of molecualr evolutionary tree: a new method of phylogenetic inference.Journal of Molecualr Evoluiton, 43, 304-311. |
| [133] | Raven P (1996) Preface. In: The Ecology and Biogeography of Nothofagus Forests (eds Veblen TT, Hill RS, Read J), pp. vii, Yale University Press, New Haven. |
| [134] | Redding DW, Mooers AO (2006) Incorporating evolutionary measures into conservation prioritization.Conservation Biology, 20, 1670-1678. |
| [135] | Ree RH, Smith SA (2008) Maximum likelihood inference of geographic range evolution by dispersal, local extinction, and cladogenesis.Systematic Biology, 57, 4-14. |
| [136] | Ren BQ (任保青), Chen ZD (陈之端) (2010) DNA barcoding of plant life.Chinese Bulletin of Botany(植物学报), 45, 1-12. (in Chinese with English abstract) |
| [137] | Renaud S, Michaux J, Schmidt D, Aguilar J, Mein P, Auffray J (2005) Morphological evolution, ecological diversification and climate change in rodents.Proceedings of the Royal Society B: Biological Sciences, 272, 609-617. |
| [138] | Richter S, Meier R (1994) The development of phylogenetic concepts in Hennig's early theoretical publications (1947-1966).Systematic Biology, 43, 212-221. |
| [139] | Rokas A, Carroll SB (2006) Bushes in the Tree of Life.PLoS Biology, 4, e352. |
| [140] | Ronquist F (1997) Dispersal-Vicariance Analysis: a new approach to the quantification of historical biogeography.Systematic Biology, 46, 195-203. |
| [141] | Ronquist F (2001) DIVA version 1.2. Computer Program for MacOS and Win32. Evolutionary Biology Centre, Uppsala University. Available at. |
| [142] | Ronqust F, Huelsenbeck J (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models.Bioinformatics, 19, 1572-1574. |
| [143] | Ronquist F, Sanmartín I (2011) Phylogenetic methods in biogeography.Annual Review of Ecology, Evolution, and Systematics, 42, 441. |
| [144] | Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Hohna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space.Systematic Biology, 61, 539-542. |
| [145] | Rosauer D, Laffan SW, Crisp MD, Donnellan SC, Cook LG (2009) Phylogenetic endemism: a new approach for identifying geographical concentrations of evolutionary history.Molecular Ecology, 18, 4061-4072. |
| [146] | Rosauer DF, Mooers AO (2013) Nurturing the use of evolutionary diversity in nature conservation.Trends in Ecology and Evolution, 28, 322-323. |
| [147] | Rutschmann F (2006) Molecular dating of phylogenetic trees: a brief review of current methods that estimate divergence times.Diversity and Distributions, 12, 35-48. |
| [148] | Salamin N, Hodkinson TR, Savolainen V (2002) Building supertrees: an empirical assessment using the grass family (Poaceae).Systematic Biology, 51, 136-150. |
| [149] | Sanderson MJ, Purvis A, Henze C (1998) Phylogenetic supertrees: assembling the trees of life.Trends in Ecology and Evolution, 13, 105-109. |
| [150] | Sanderson MJ, Shaffer HB (2002) Troubleshooting molecular phylogenetic analyses.Annual Review of Ecology and Systematics, 33, 49-72. |
| [151] | Sanderson MJ (2003) r8s: inferring absolute rates of molecular evolution and divergence times in the absence of a molecular clock.Bioinformatics, 19, 301-302. |
| [152] | Sanderson MJ, McMahon MM (2007) Inferring angiosperm phylogeny from EST data with widespread gene duplication.BMC Evolutionary Biology, 7, S3. |
| [153] | Santamaria L, Mendez PF (2012) Evolution in biodiversity policy—current gaps and future needs.Evolutionary Applications, 5, 202-218. |
| [154] | Sauquet H (2013) A practical guide to molecular dating.Comptes Rendus Palevol, 6, 355-367. |
| [155] | Schulte P, Alegret L, Arenillas I, Arz JA, Barton PJ, Bown PR, Bralower TJ, Christeson GL, Claeys P, Cockell CS, Collins GS, Deutsch A, Goldin TJ, Goto K, Grajales-Nishimura JM, Grieve RAF, Gulick SPS, Johnson KR, Kiessling W, Koeberl C, Kring DA, MacLeod KG, Matsui T, Melosh J, Montanari A, Morgan JV, Neal CR, Nichols DJ, Norris RD, Pierazzo E, Ravizza G, Rebolledo-Vieyra M, Reimold WU, Robin E, Salge T, Speijer RP, Sweet AR, Urrutia-Fucugauchi J, Vajda V, Whalen MT, Willumsen PS (2010) The Chicxulub asteroid impact and mass extinction at the Cretaceous-Paleogene boundary.Science, 327, 1214-1218. |
| [156] | Schweizer M, Seehausen O, Hertwig ST (2011) Macroevolutionary patterns in the diversification of parrots: effects of climate change, geological events and key innovations.Journal of Biogeography, 38, 2176-2194. |
| [157] | Seelanan T, Schnabel A, Wendel JF (1997) Congruence and consensus in the cotton tribe (Malvaceae).Systematic Botany, 22, 259-290. |
| [158] | Silvertown J, Dodd M, Gowing D, Lawson C, McConway K (2006) Phylogeny and the hierarchical organization of plant diversity.Ecology, 87, S39-S49. |
| [159] | Simpson MG (2012) Plant systematics. In: Introgression, Hybridization and Polyploidy, 2nd edn. pp. 653-655. Science Press, Beijing. |
| [160] | Smith SA, O’Meara BC (2012) treePL: divergence time estimation using penalized likelihood for large phylogenies.Bioinformatics, 28, 2689-2690. |
| [161] | Sneath PHA (1995) Thirty years of numerical taxonomy.Systematic Biology, 44, 281-298. |
| [162] | Sokal RR, Sneath PHA (1963) Principles of Numerical Taxonomy. Freeman, San Francisco. |
| [163] | Soltis DE, Burleigh G, Barbazuk WB, Moore MJ, Soltis PS (2009) Advances in the use of next-generation sequence data in plant systematics and evolution.ISHS Acta Horticulturae, 859, 193-206. |
| [164] | Soltis DE, Smith SA, Cellinese N, Wurdack KJ, Tank DC, Brockington SF, Refulio-Rodriguez NF, Walker JB, Moore MJ, Carlsward BS, Bell CD, Latvis M, Crawley S, Black C, Diouf D, Xi Z, Rushworth CA, Gitzendanner MA, Sytsma KJ, Qiu YL, Hilu KW, Davis CC, Sanderson MJ, Beaman RS, Olmstead RG, Judd WS, Donoghue MJ, Soltis PS (2011) Angiosperm phylogeny: 17 genes, 640 taxa.American Journal of Botany, 98, 704-730. |
| [165] | Soltis PS, Soltis DE (2013) Angiosperm phylogeny: a framework for studies of genome evolution.Plant Genome Diversity, 2, 1-11. |
| [166] | Song S, Liu L, Edwards SV, Wu S (2012) Resolving conflict in eutherian mammal phylogeny using phylogenomics and the multispecies coalescent model.Proceedings of the National Academy of Sciences,USA, 109, 14942-14947. |
| [167] | Springer MS, Meredith RW, Gatesy JG, Emerling CA, Park J, Rabosky DL, Stadler T, Steiner C, Ryder OA, Janecka JE, Fisher CA, Murphy WJ (2012) Macroevolutionary dynamics and historical biogeography of primate diversification inferred from a species supermatrix.PLoS ONE, 7, e49521. |
| [168] | Stadler T (2011) Mammalian phylogeny reveals recent diversification rate shifts.Proceedings of the National Academy of Sciences,USA, 108, 6187-6192. |
| [169] | Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models.Bioinformatics, 22, 2688-2690. |
| [170] | Stebbins GL (1974) Flowering Plants: Evolution Above the Species Level. Harvard University Press, Cambridge. |
| [171] | Steel M, Penny D (2000) Parsimony, likelihood, and the role of models in molecular phylogenetics.Molecular Biology and Evolution, 4, 64-71. |
| [172] | Stevens PF (2001) Angiosperm Phylogeny Website. Version 9, June 2008. Available at. |
| [173] | Su JX (苏俊霞) (2012) The Study on Molecular Phylogeny of Rosids (蔷薇分支的分子系统学研究). PhD dissertation, Institute of Botany, Chinese Academy of Sciences, Beijing. (in Chinese with English abstract) |
| [174] | Su JX, Wang W, Zhang LB, Chen ZD (2012) Phylogentic placement of two enigmatic genera, Borthwickia and Stixis, based on molecular and pollen data, and the description of a new family of Brassicales, Borthwickiaceae.Taxon, 61, 601-611. |
| [175] | Swofford DL (2003)Paup*: Phylogenetic Anlysis Using Parsimony (*and Other Methods), Version 4.0b10. Sinauer Associates, Sunderland. |
| [176] | Takhtajan AL (1997) Diversity and the Classification of Flowering Plants. Columbia University Press, New York. |
| [177] | Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods.Molecular Biology and Evolution, 28, 2731-2739. |
| [178] | Tang ZY, Wang ZH, Zheng CY, Fang JY (2006) Biodiversity in China’s mountains.Frontiers in Ecology and the Environment, 4, 347-352. |
| [179] | Thomas JA, Ware JL (2011) The northeastern symposium on evolutionary divergence time: fossil and molecular dating: molecular and fossil dating: a compatible match?Entomologica Americana, 117, 1-8. |
| [180] | Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools.Nucleic Acids Research, 25, 4876-4882. |
| [181] | Thomson RC, Shaffer HB (2010) Sparse supermatrices for phylogenetic inference: taxonomy alignment, rogue taxa, and the phylogeny of living turtles.Systematic Biology, 59, 42-58. |
| [182] | Thorley JL, Wilkinson M (1999) Testing the phylogenetic stability of early tetrapods.Journal of Theoretical Biology, 200, 343-344. |
| [183] | Thorne JL, Kishino H (2002) Divergence time and evolutionary rate estimation with multilocus data.Systematic Biology, 51, 689-702. |
| [184] | Töpel M, Antonelli A, Yesson C, Eriksen B (2012) Past climate change and plant evolution in western north America: a case study in Rosaceae.PLoS ONE, 7, e50358. |
| [185] | Vane-Wright RI, Humphries CJ, Williams PH (1991) What to protect? Systematics and the agony of choice.Biological Conservation, 55, 235-254. |
| [186] | Wang W, Lu AM, Ren Y, Endress ME, Chen ZD (2009) Phy- logeny and classification of Ranunculales: Evidence from four molecular loci and morphological data.Perspectives in Plant Ecology, Evolution and Systematics, 11, 81-110. |
| [187] | Wang W, Ortiz RDC, Jacques FMB, Xiang XG, Li H-L, Lin L, Li RQ, Liu Y, Soltis PS, Soltis DE, Chen ZD (2012) Menispermaceae and the diversification of tropical rainforests near the Cretaceous-Paleogene boundary.New Phytologist, 195, 470-478. |
| [188] | Wang ZH (王志恒), Tang ZY (唐志尧), Fang JY (方精云) (2009) Metabolic theory of ecology: an explanation for species richness patterns based on the metabolic processes of organisms.Biodiversity Science(生物多样性), 17, 625-634. (in Chinese with English abstract) |
| [189] | Wang ZH, Brown JH, Tang ZY, Fang JY (2009) Temperature dependence, spatial scale, and tree species diversity in eastern Asia and North America.Proceedings of the National Academy of Sciences,USA, 106, 13388-13392. |
| [190] | Webb CO, Ackerly DD, Kembel SW (2008) Phylocom: software for the analysis of phylogenetic community structure and trait evolution.Bioinformatics, 24, 2098-2100. |
| [191] | Webb CO, Ackerly DD, McPeek MA, Donoghue MJ (2002) Phylogenies and community ecology.Annual Review of Ecology and Systematics, 33, 475-505. |
| [192] | Webb CO, Donoghue MJ (2005) Phylomatic: tree assembly for applied phylogenetics.Molecular Ecology Notes, 5, 181-183. |
| [193] | Weir J, Schluter D (2008) Calibrating the avian molecular clock.Molecular Ecology, 17, 2321-2328. |
| [194] | Welch JJ, Bromham L (2005) Molecular dating when rates vary.Trends in Ecology and Evolution, 20, 320-327. |
| [195] | Wen J, Ickert-Bond S, Nie Z-L, Li R (2010) Timing and modes of evolution of eastern Asian-North American biogeo- grap- hic disjunctions in seed plants.Darwin’s Heritage Today: Proceedings of the Darwin 2010 Beijing International Conference, pp. 252-269. |
| [196] | Wen J, Ree RH, Ickert-Bond SM, Nie Z, Funk V (2013) Biogeography: where do we go from here?Taxon, 62, 912-927. |
| [197] | Wendel JF, Doyle JJ (1998) Phylogenetic incongruence: window into genome history and molecular evolution. In: Molecular Systematics of Plants II: DNA Sequencing (eds Soltis DE, Soltis PS, Doyle JJ), pp. 265-296. Kluwer, Boston. |
| [198] | Wiens JJ, Graham CH (2005) Niche conservatism: integrating evolution, ecology, and conservation biology.Annual Review of Ecology, Evolution, and Systematics, 36, 519-539. |
| [199] | Wiley EO, Siegel-Causey D, Brooks DR, Funk V (1991) The Compleat Cladist: A Primer of Phylogenetic Procedures. Museum of Natural History, University of Kansas Lawrence. |
| [200] | Williams TA, Foster PG, Nye TMW, Cox CJ, Embley TM (2012) A congruent phylogenomic signal places eukaryotes within the Archaea.Proceedings of the Royal Society B: Biological Sciences, 1749, 4870-4879. |
| [201] | Winter M, Devicto, V, Schweiger O (2013) Phylogenetic diversity and nature conservation: where are we?Trends in Ecology and Evolution, 28, 199-204. |
| [202] | Woese CR, Fox GE (1977) Phylogenetic structure of the prokaryotic domain: the primary kingdoms.Proceedings of the National Academy of Sciences,USA, 74, 5088-5090. |
| [203] | Wu ZQ, Ge S (2012) The phylogeny of the BEP clade in grasses revisited: evidence from the whole-genome sequences of chloroplasts.Molecular Phylogenetics and Evolution, 62, 573-578. |
| [204] | Yang Z (2004) A heuristic rate smoothing procedure for maximum likelihood estimation of species divergence times.Acta Zoologica Sinica, 50, 656. |
| [205] | Yang Z (2006) Computational Molecular Evolution. Oxford University Press, New York. |
| [206] | Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood.Molecular Biology and Evolution, 24, 1586-1591. |
| [207] | Yang Z, Rannala B (2012) Molecular phylogenetics: principles and practice.Nature Reviews Genetics,13, 303-314. |
| [208] | Yoder AD, Clancey E, Des Roches S, Eastman JM, Gentry L, Godsoe WKW, Hagey T, Jochimsen D, Oswald BP, Robertson J, Sarver BAJ, Schenk JJ, Spear S, Harmon LJ (2010) Ecology opportunity and the origin of adaptive radiations.Journal of Evolutionary Biology, 23, 1581-1596. |
| [209] | Yu Y, Harris A, He X (2010) S-DIVA (Statistical Dispersal- Vicariance Analysis): a tool for inferring biogeographic histories.Molecular Phylogenetics and Evolution, 56, 848-850. |
| [210] | Zhang JB, Li RQ, Xiang XG, Manchester SR, Lin L, Wang W, Wen J, Chen ZD (2013) Integrated fossil and molecular data reveal the biogeographic diversification of the eastern Asian-eastern North American disjunct hickory genus (Carya Nutt.).PLoS ONE, 8, e70449. |
| [211] | Zhang N, Zeng LP, Shan HY, Ma H (2012) Highly conserved low-copy nuclear genes as effective markers for phylogenetic analyses in angiosperms.New Phytologist, 195, 923-937. |
| [212] | Zhang SB, Slik JWF, Zhang JL, Cao KF (2011) Spatial patterns of wood traits in China are controlled by phylogeny and the environment.Global Ecology and Biogeography, 20, 241-250. |
| [213] | Zhu XY, Chase MW, Qiu YL, Kong HZ, Dilcher DL, Li JH, Chen ZD (2007) Mitochondrial matR sequences help to resolve deep phylogenetic relationships in rosids.BMC Evolutionary Biology, 7, 217. |
| [214] | Zou XH (邹新慧), Ge S (葛颂) (2008) Conflicting gene trees and phylogenomics.Journal of Systematics and Evolution(植物分类学报), 46, 795-807. (in Chinese with English abstract) |
| [215] | Zuckerkandl E, Pauling L (1965) Molecules as documents of evolutionary history.Journal of Theoretical Biology, 8, 357-366. |
| [1] | Jing Gan Xiangxu Liu Xueming Lu Xing Yue. China's Large Cities in Global Biodiversity Hotspots: Conservation Policies and Optimization Directions [J]. Biodiv Sci, 2025, 33(5): 24529-. |
| [2] | Zixuan Zeng Rui Yang Yue Huang Luyao Chen. Characteristics of bird diversity and environmental relationships in Tsinghua University campus [J]. Biodiv Sci, 2025, 33(5): 24373-. |
| [3] | Mingyue Zang, Li Liu, Yue Ma, Xu Xu, Feilong Hu, Xiaoqiang Lu, Jiaqi Li, Cigang Yu, Yan Liu. China’s urban biodiversity conservation under the Kunming-Montreal Global Biodiversity Framework [J]. Biodiv Sci, 2025, 33(5): 24482-. |
| [4] | Xiaoyu Zhu, Chenhao Wang, Zhongjun Wang, Yujun Zhang. Research progress and prospect of urban green space biodiversity [J]. Biodiv Sci, 2025, 33(5): 25027-. |
| [5] | Lin Yuan, Siqi Wang, Jingxuan Hou. “Leaving space for wildness” in metropolitan region: Trends and prospects [J]. Biodiv Sci, 2025, 33(5): 24481-. |
| [6] | Min Hu, Binbin Li, Coraline Goron. Green is not enough: A management framework for urban biodiversity-friendly parks [J]. Biodiv Sci, 2025, 33(5): 24483-. |
| [7] | Xin Wang, Femgyu Bao. Analysis of the ecological restoration effect of South Dianchi National Wetland Park based on the enhancement of bird diversity [J]. Biodiv Sci, 2025, 33(5): 24531-. |
| [8] | Yue Ming, Peiyao Hao, Lingqian Tan, Xi Zheng. A study on urban biodiversity conservation and enhancement in china based on the concept of green and high-quality development of cities [J]. Biodiv Sci, 2025, 33(5): 24524-. |
| [9] | Gan Xie, Jing Xuan, Qidi Fu, Ze Wei, Kai Xue, Hairui Luo, Jixi Gao, Min Li. Establishing an intelligent identification model for unmanned aerial vehicle surveys of grassland plant diversity [J]. Biodiv Sci, 2025, 33(4): 24236-. |
| [10] | Xiaolin Chu, Quanguo Zhang. A review of experimental evidence for the evolutionary speed hypothesis [J]. Biodiv Sci, 2025, 33(4): 25019-. |
| [11] | Zhiyu Liu, Xin Ji, Guohui Sui, Ding Yang, Xuankun Li. Invertebrate diversity in buffalo grass and weedy lawns at Beijing Capital International Airport [J]. Biodiv Sci, 2025, 33(4): 24456-. |
| [12] | Xiaoqiang Lu, Shanshan Dong, Yue Ma, Xu Xu, Feng Qiu, Mingyue Zang, Yaqiong Wan, Luanxin Li, Cigang Yu, Yan Liu. Current status, challenges, and prospects of frontier technologies in biodiversity conservation applications [J]. Biodiv Sci, 2025, 33(4): 24440-. |
| [13] | Qiaoyi Nong, Jun Cao, Wenda Cheng, Yanqiong Peng. Comparative study of monitoring methods for Apoidea resources and diversity [J]. Biodiv Sci, 2025, 33(4): 25057-. |
| [14] | Guo Yutong, Li Sucui, Wang Zhi, Xie Yan, Yang Xue, Zhou Guangjin, You Chunhe, Zhu Saning, Gao Jixi. Coverage and distribution of national key protected wild species in China’s nature reserves [J]. Biodiv Sci, 2025, 33(3): 24423-. |
| [15] | Zhao Weiyang, Wang Wei, Ma Bingran. Advances and prospects in research on other effective area-based conservation measures (OECMs) [J]. Biodiv Sci, 2025, 33(3): 24525-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2022 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn ![]()