



Biodiv Sci ›› 2018, Vol. 26 ›› Issue (5): 445-456. DOI: 10.17520/biods.2018058 cstr: 32101.14.biods.2018058
Special Issue: 传粉生物学; 物种形成与系统进化; 昆虫多样性与生态功能
• Reviews • Previous Articles Next Articles
Dandan Lang1,2, Min Tang1,3, Xin Zhou1,3,*( )
)
Received:2018-02-15
Accepted:2018-05-11
Online:2018-05-20
Published:2018-09-11
Contact:
Zhou Xin
About author:# Co-first authors
Dandan Lang,Min Tang,Xin Zhou. Qualitative and quantitative molecular construction of plant-pollinator network: Application and prospective[J]. Biodiv Sci, 2018, 26(5): 445-456.
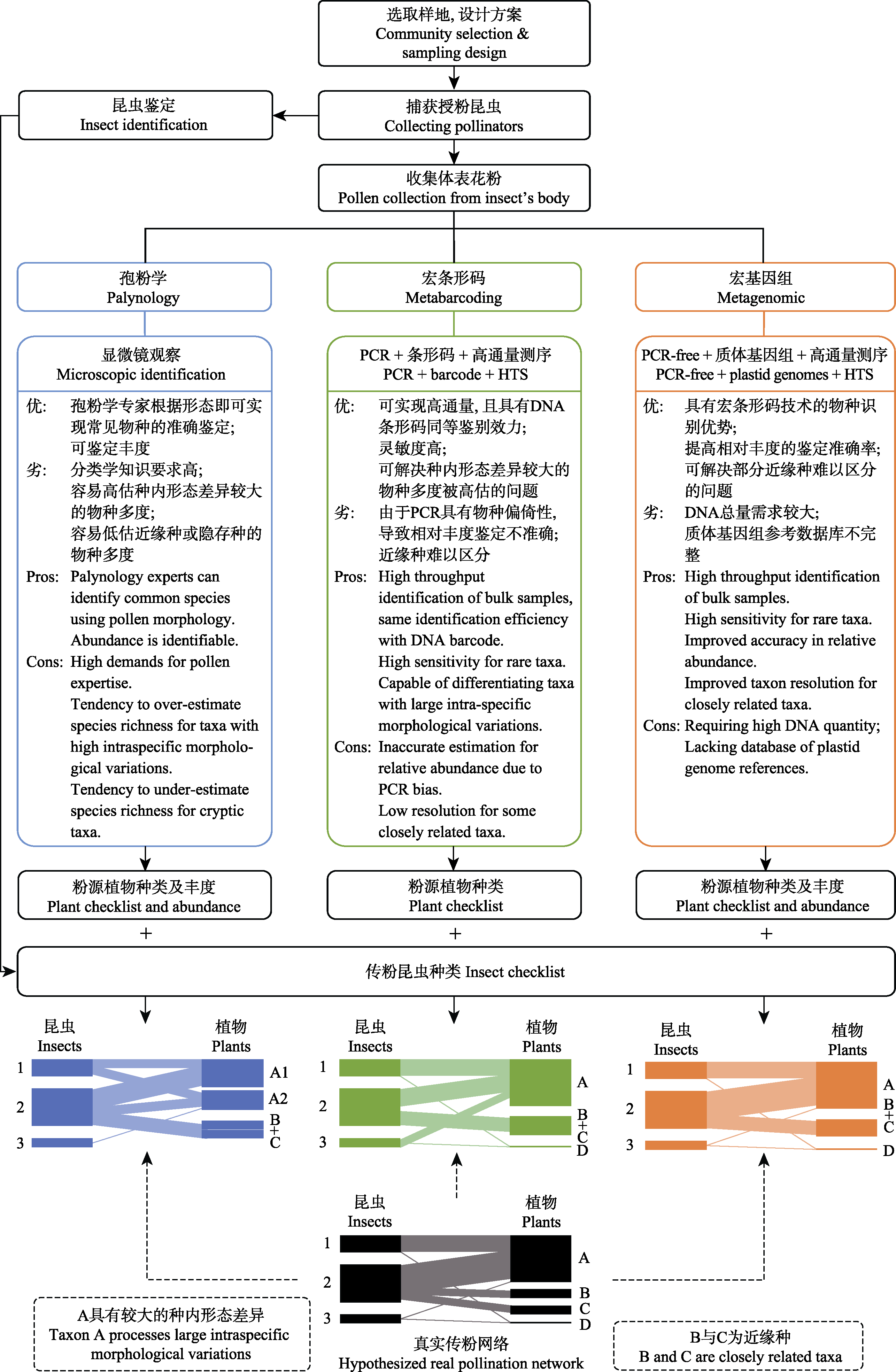
Fig. 1 Construction of pollination network and comparison of analysis methods of mixed pollen composition. The contents of the blue and the green boxes refer to morphological and molecular methods, respectively. The content in the orange box is the PCR-free genome-skimming (metagenomics) approach proposed in this paper. The black network model at the bottom represents the real pollination network, and the blue, green and orange network models represent network structures constructed by the corresponding methods. Due to the limitations of various methods, the constructed networks are potentially deviated from the real network. Metabarcoding and metagenomic techniques can alleviate issues caused by intra-specific morphological variations; but some closely related species remain difficult to differentiate. Compared to metabarcoding, the metagenomic technology can reduce species bias caused by PCR and improve the accuracy in relative abundance.
| 物种 Species | NCBI序列号 NCBI accession number | 物种 Species | NCBI序列号 NCBI accession number |
|---|---|---|---|
| Apis andreniformis | KF736157.1 | Bombus lapidarius | KT164641.1 |
| Apis cerana | NC_014295.1 | Bombus lucorum | KT164681.1 |
| Apis dorsata | KC294229.1 | Bombus pascuorum | KT164630.1 |
| Apis florea | NC_021401.1 | Bombus terrestris | KT368150.1 |
| Apis mellifera sahariensis | NC_035883.1 | Melipona bicolor | NC_004529.1 |
| Apis nigrocincta | KY799147.1 | Melipona scutellaris | NC_026198.1 |
| Bombus breviceps | MF478986.1 | Nomada fabriciana | KT164663.1 |
| Bombus consobrinus | MF995069.1 | Nomada flava | KT164670.1 |
| Bombus hypocrita sapporensis | NC_011923.1 | Nomada flavoguttata | KT164617.1 |
| Bombus ignitus | NC_010967.1 | Nomada goodeniana | KT164660.1 |
Table 1 The list of 20 species of Apidae (including six species of Apis) and their mitochondrial genomes’ accession numbers in NCBI
| 物种 Species | NCBI序列号 NCBI accession number | 物种 Species | NCBI序列号 NCBI accession number |
|---|---|---|---|
| Apis andreniformis | KF736157.1 | Bombus lapidarius | KT164641.1 |
| Apis cerana | NC_014295.1 | Bombus lucorum | KT164681.1 |
| Apis dorsata | KC294229.1 | Bombus pascuorum | KT164630.1 |
| Apis florea | NC_021401.1 | Bombus terrestris | KT368150.1 |
| Apis mellifera sahariensis | NC_035883.1 | Melipona bicolor | NC_004529.1 |
| Apis nigrocincta | KY799147.1 | Melipona scutellaris | NC_026198.1 |
| Bombus breviceps | MF478986.1 | Nomada fabriciana | KT164663.1 |
| Bombus consobrinus | MF995069.1 | Nomada flava | KT164670.1 |
| Bombus hypocrita sapporensis | NC_011923.1 | Nomada flavoguttata | KT164617.1 |
| Bombus ignitus | NC_010967.1 | Nomada goodeniana | KT164660.1 |
| 物种 Species | NCBI序列号 NCBI accession number | 物种 Species | NCBI序列号 NCBI accession number |
|---|---|---|---|
| Anoectochilus emeiensis | NC_033895.1 | Dendrobium parciflorum | NC_035334.1 |
| Apostasia odorata | NC_030722.1 | Dendrobium parishii | NC_035339.1 |
| Bletilla ochracea | NC_029483.1 | Dendrobium pendulum | NC_029705.1 |
| Bletilla striata | NC_028422.1 | Dendrobium primulinum | NC_035321.1 |
| Calanthe triplicata | NC_024544.1 | Dendrobium salaccense | NC_035332.1 |
| Cattleya crispata | NC_026568.1 | Dendrobium spatella | NC_035333.1 |
| Cattleya liliputana | NC_032083.1 | Dendrobium strongylanthum | NC_027691.1 |
| Cephalanthera longifolia | NC_030704.1 | Dendrobium wardianum | NC_035329.1 |
| Cymbidium aloifolium | NC_021429.1 | Dendrobium wilsonii | NC_035330.1 |
| Cymbidium ensifolium | NC_028525.1 | Dendrobium xichouense | NC_035341.1 |
| Cymbidium faberi | NC_027743.1 | Elleanthus sodiroi | NC_027266.1 |
| Cymbidium goeringii | NC_028524.1 | Epipactis mairei | NC_030705.1 |
| Cymbidium kanran | NC_029711.1 | Epipactis veratrifolia | NC_030708.1 |
| Cymbidium lancifolium | NC_029712.1 | Erycina pusilla | NC_018114.1 |
| Cymbidium macrorhizon | NC_029713.1 | Gastrochilus fuscopunctatus | NC_035830.1 |
| Cymbidium mannii | NC_021433.1 | Gastrochilus japonicus | NC_035833.1 |
| Cymbidium sinense | NC_021430.1 | Goodyera fumata | NC_026773.1 |
| Cymbidium tortisepalum | NC_021431.1 | Goodyera procera | NC_029363.1 |
| Cymbidium tracyanum | NC_021432.1 | Goodyera schlechtendaliana | NC_029364.1 |
| Cypripedium formosanum | NC_026772.1 | Goodyera velutina | NC_029365.1 |
| Cypripedium macranthos | NC_024421.1 | Habenaria pantlingiana | NC_026775.1 |
| Dendrobium aphyllum | NC_035322.1 | Habenaria radiata | NC_035834.1 |
| Dendrobium brymerianum | NC_035323.1 | Listera fugongensis | NC_030711.1 |
| Dendrobium catenatum | NC_024019.1 | Ludisia discolor | NC_030540.1 |
| Dendrobium chrysanthum | NC_035336.1 | Masdevallia coccinea | NC_026541.1 |
| Dendrobium chrysotoxum | NC_028549.1 | Masdevallia picturata | NC_026777.1 |
| Dendrobium crepidatum | NC_035331.1 | Neottia ovate | NC_030712.1 |
| Dendrobium denneanum | NC_035324.1 | Neottia pinetorum | NC_030710.1 |
| Dendrobium devonianum | NC_035325.1 | Oberonia japonica | NC_035832.1 |
| Dendrobium ellipsophyllum | NC_035340.1 | Paphiopedilum armeniacum | NC_026779.1 |
| Dendrobium exile | NC_035343.1 | Paphiopedilum niveum | NC_026776.1 |
| Dendrobium falconeri | NC_035326.1 | Pelatantheria scolopendrifolia | NC_035829.1 |
| Dendrobium fanjingshanense | NC_035344.1 | Phalaenopsis equestris | NC_017609.1 |
| Dendrobium fimbriatum | NC_035342.1 | Phragmipedium longifolium | NC_028149.1 |
| Dendrobium gratiosissimum | NC_035327.1 | Sobralia callosa | NC_028147.1 |
| Dendrobium henryi | NC_035335.1 | Thrixspermum japonicum | NC_035831.1 |
| Dendrobium hercoglossum | NC_035328.1 | Vanilla aphylla | NC_035320.1 |
| Dendrobium huoshanense | NC_028430.1 | Vanilla planifolia | NC_026778.1 |
| Dendrobium jenkinsii | NC_035337.1 | Sobralia aff. bouchei | NC_028209.1 |
| Dendrobium lohohense | NC_035338.1 | Phalaenopsis hybrid | NC_025593.1 |
| Dendrobium moniliforme | NC_035154.1 | Phalaenopsis aphrodite formosana | NC_007499.1 |
| Dendrobium nobile | NC_029456.1 | Oncidium hybrid | NC_014056.1 |
Table 2 The list of 84 species of Orchidaceae (including 31 species of Dendrobium) and their chloroplast genomes’ accession numbers in NCBI
| 物种 Species | NCBI序列号 NCBI accession number | 物种 Species | NCBI序列号 NCBI accession number |
|---|---|---|---|
| Anoectochilus emeiensis | NC_033895.1 | Dendrobium parciflorum | NC_035334.1 |
| Apostasia odorata | NC_030722.1 | Dendrobium parishii | NC_035339.1 |
| Bletilla ochracea | NC_029483.1 | Dendrobium pendulum | NC_029705.1 |
| Bletilla striata | NC_028422.1 | Dendrobium primulinum | NC_035321.1 |
| Calanthe triplicata | NC_024544.1 | Dendrobium salaccense | NC_035332.1 |
| Cattleya crispata | NC_026568.1 | Dendrobium spatella | NC_035333.1 |
| Cattleya liliputana | NC_032083.1 | Dendrobium strongylanthum | NC_027691.1 |
| Cephalanthera longifolia | NC_030704.1 | Dendrobium wardianum | NC_035329.1 |
| Cymbidium aloifolium | NC_021429.1 | Dendrobium wilsonii | NC_035330.1 |
| Cymbidium ensifolium | NC_028525.1 | Dendrobium xichouense | NC_035341.1 |
| Cymbidium faberi | NC_027743.1 | Elleanthus sodiroi | NC_027266.1 |
| Cymbidium goeringii | NC_028524.1 | Epipactis mairei | NC_030705.1 |
| Cymbidium kanran | NC_029711.1 | Epipactis veratrifolia | NC_030708.1 |
| Cymbidium lancifolium | NC_029712.1 | Erycina pusilla | NC_018114.1 |
| Cymbidium macrorhizon | NC_029713.1 | Gastrochilus fuscopunctatus | NC_035830.1 |
| Cymbidium mannii | NC_021433.1 | Gastrochilus japonicus | NC_035833.1 |
| Cymbidium sinense | NC_021430.1 | Goodyera fumata | NC_026773.1 |
| Cymbidium tortisepalum | NC_021431.1 | Goodyera procera | NC_029363.1 |
| Cymbidium tracyanum | NC_021432.1 | Goodyera schlechtendaliana | NC_029364.1 |
| Cypripedium formosanum | NC_026772.1 | Goodyera velutina | NC_029365.1 |
| Cypripedium macranthos | NC_024421.1 | Habenaria pantlingiana | NC_026775.1 |
| Dendrobium aphyllum | NC_035322.1 | Habenaria radiata | NC_035834.1 |
| Dendrobium brymerianum | NC_035323.1 | Listera fugongensis | NC_030711.1 |
| Dendrobium catenatum | NC_024019.1 | Ludisia discolor | NC_030540.1 |
| Dendrobium chrysanthum | NC_035336.1 | Masdevallia coccinea | NC_026541.1 |
| Dendrobium chrysotoxum | NC_028549.1 | Masdevallia picturata | NC_026777.1 |
| Dendrobium crepidatum | NC_035331.1 | Neottia ovate | NC_030712.1 |
| Dendrobium denneanum | NC_035324.1 | Neottia pinetorum | NC_030710.1 |
| Dendrobium devonianum | NC_035325.1 | Oberonia japonica | NC_035832.1 |
| Dendrobium ellipsophyllum | NC_035340.1 | Paphiopedilum armeniacum | NC_026779.1 |
| Dendrobium exile | NC_035343.1 | Paphiopedilum niveum | NC_026776.1 |
| Dendrobium falconeri | NC_035326.1 | Pelatantheria scolopendrifolia | NC_035829.1 |
| Dendrobium fanjingshanense | NC_035344.1 | Phalaenopsis equestris | NC_017609.1 |
| Dendrobium fimbriatum | NC_035342.1 | Phragmipedium longifolium | NC_028149.1 |
| Dendrobium gratiosissimum | NC_035327.1 | Sobralia callosa | NC_028147.1 |
| Dendrobium henryi | NC_035335.1 | Thrixspermum japonicum | NC_035831.1 |
| Dendrobium hercoglossum | NC_035328.1 | Vanilla aphylla | NC_035320.1 |
| Dendrobium huoshanense | NC_028430.1 | Vanilla planifolia | NC_026778.1 |
| Dendrobium jenkinsii | NC_035337.1 | Sobralia aff. bouchei | NC_028209.1 |
| Dendrobium lohohense | NC_035338.1 | Phalaenopsis hybrid | NC_025593.1 |
| Dendrobium moniliforme | NC_035154.1 | Phalaenopsis aphrodite formosana | NC_007499.1 |
| Dendrobium nobile | NC_029456.1 | Oncidium hybrid | NC_014056.1 |
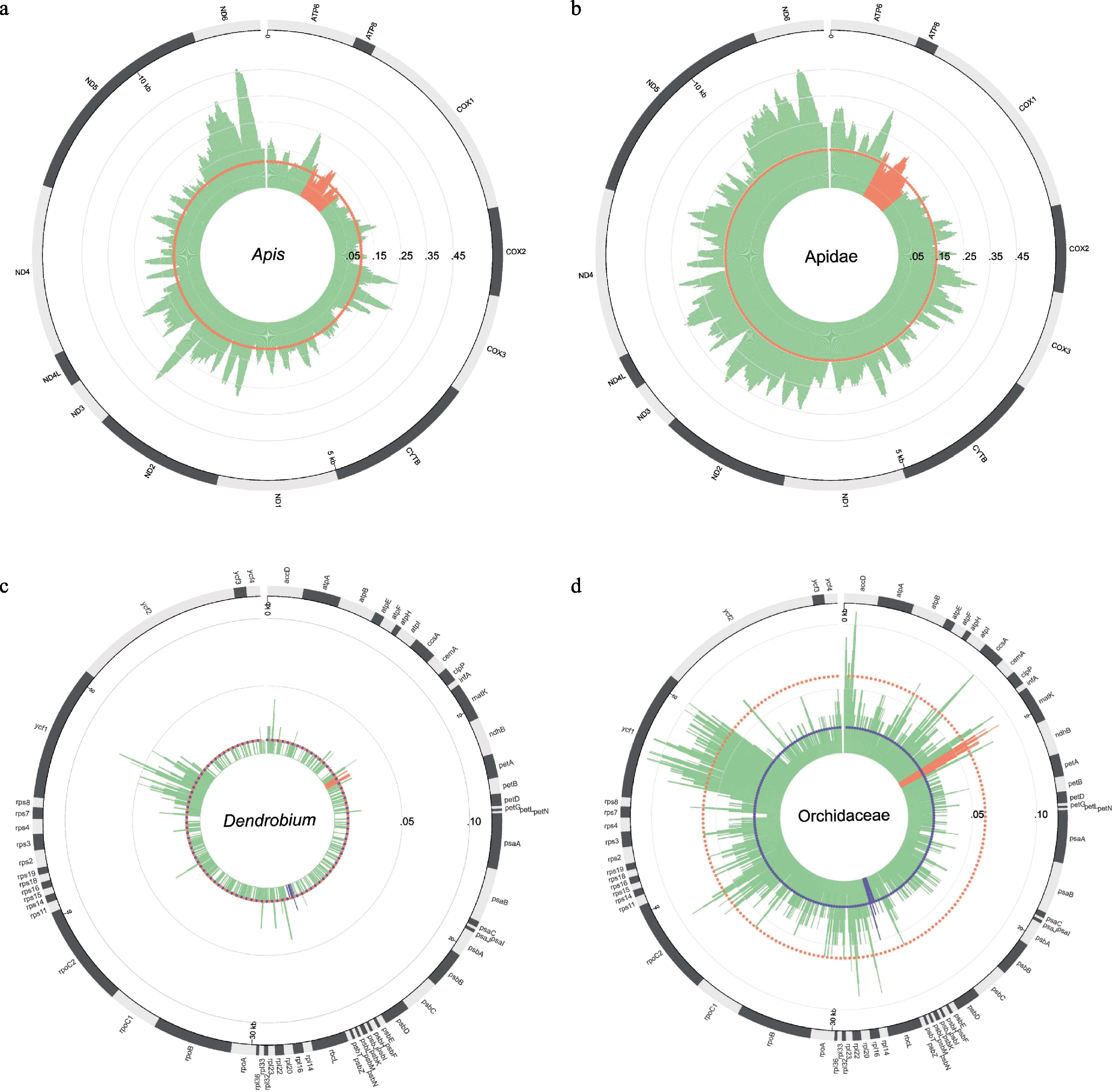
Fig. 3 The P-distance map among species and genera of the mitochondrial genomes and the chloroplast genomes. (a) The P-distance map of 13 protein-coding genes in the mitochondrial genomes among six species in Apis; (b) The P-distance map of 13 protein-coding genes in the mitochondrial genomes among 20 species in Apidae; (c) The P-distance map of 67 protein-coding genes in the chloroplast genomes among 31 species in Dendrobium; (d) The P-distance map of 67 protein-coding genes in the chloroplast genomes among 84 species in Orchidaceae. In panels a & b, the orange area is the location of the COI barcode, and the orange dotted circle represents the median value of the COI barcode’s P-distance. In panels c & d, the orange area is the location of the matK barcode, the purple area is the location of the rbcL barcode, and the orange dotted circle represents the median value of the matK barcode’s P-distance, the purple dotted circle represents the median value of the rbcL barcode’s P-distance.
| 1 | Aguilar R, Ashworth L, Galetto L, Aizen MA (2006) Plant reproductive susceptibility to habitat fragmentation: Review and synthesis through a meta-analysis. Ecology Letters, 9, 968-980. |
| 2 | Arif IA, Khan HA, Al Sadoon M, Shobrak M (2011) Limited efficiency of universal mini-barcode primers for DNA amplification from desert reptiles, birds and mammals. Genetics and Molecular Research, 10, 3559-3564. |
| 3 | Ashman TL, Knight TM, Steets JA, Amarasekare P, Burd M, Campbell DR, Dudash MR, Johnston MO, Mazer SJ, Mitchell RJ, Morgan MT, Wilson WG (2004) Pollen limitation of plant reproduction: Ecological and evolutionary causes and consequences. Ecology, 85, 2408-2421. |
| 4 | Aziz AN, Sauve RJ (2008) Genetic mapping of Echinacea purpurea via individual pollen DNA fingerprinting. Molecular Breeding, 21, 227-232. |
| 5 | Bagella S, Satta A, Floris I, Caria MC, Rossetti I, Podani J (2013) Effects of plant community composition and flowering phenology on honeybee foraging in Mediterranean sylvo-pastoral systems. Applied Vegetation Science, 16, 689-697. |
| 6 | Bambara SB (1991) Using pollen to identify honey. American Bee Journal, 131, 242-243. |
| 7 | Bascompte J, Jordano P (2007) Plant-animal mutualistic networks: The architecture of biodiversity. Annual Review of Ecology, Evolution and Systematics, 38, 567-593. |
| 8 | Bell KL, Julie F, Kevin SB, Emily KD, David G, Brice L, Connor M, Berry JB (2017) Applying pollen DNA metabarcoding to the study of plant-pollinator interactions. Applications in Plant Sciences, 5, 1600124. |
| 9 | Bell KL, Kevin SB, Kazufusa CO, Roman A, Berry JB (2016) Review and future prospects for DNA barcoding methods in forensic palynology. Forensic Science International: Genetics, 21, 110-116. |
| 10 | Bellemain E, Carlsen T, Brochmann C, Coissac E, Taberlet P, Kauserud H (2010) ITS as an environmental DNA barcode for fungi: An in silico approach reveals potential PCR biases. BMC Microbiology, 10, 189. |
| 11 | Binladen J, Gilbert MT, Bollback JP, Panitz F, Bendixen C, Nielsen R, Willerslev E (2007) The use of coded PCR primers enables high-throughput sequencing of multiple homolog amplification products by 454 parallel sequencing. PLoS ONE, 2, e197. |
| 12 | Bosch J, González AM, Rodrigo A, Navarro D (2009) Plant- pollinator networks: Adding the pollinator’s perspective. Ecology Letters, 12, 409-419. |
| 13 | Bruni I, Galimberti A, Caridi L, Scaccabarozzi D, Mattia DF, Casiraghi M, Labra M (2015) A DNA barcoding approach to identify plant species in multiflower honey. Food Chemistry, 170, 308-315. |
| 14 | Cameron SA, Hines HM, Williams PH (2007) A comprehensive phylogeny of the bumble bees (Bombus). Biological Journal of the Linnean Society, 91, 161-188. |
| 15 | CBOL Plant Working Group (2009) A DNA barcode for land plants. Proceedings of the National Academy of Sciences, USA, 106, 12794-12797. |
| 16 | Chen SL, Yao H, Han JP, Liu C, Song JY, Shi LC, Zhu YJ, Ma XY, Gao T, Pang XH, Luo K, Li Y, Li XW, Jia XC, Lin YL, Leon C (2010) Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS ONE, 5, 1-8. |
| 17 | China Plant BOL Group, Li DZ, Gao LM, Li HT, Wang H, Ge XJ, Liu JQ, Chen ZD, Zhou SL, Chen SL, Yang JB, Fu CX, Zeng CX, Yan HF, Zhu YJ, Sun YS, Chen SY, Zhao L, Wang K, Yang T, Duan GW (2011) Comparative analysis of a large dataset indicates that internal transcribed spacer (ITS) should be incorporated into the core barcode for seed plants. Proceedings of the National Academy of Sciences, USA, 108, 19641-19646. |
| 18 | Coissac E, Hollingsworth PM, Lavergne S, Taberlet P (2016) From barcodes to genomes: Extending the concept of DNA barcoding. Molecular Ecology, 25, 1423-1428. |
| 19 | Danforth BN, Cardinal S, Praz C, Almeida EA, Michez D (2013) The impact of molecular data on our understanding of bee phylogeny and evolution. Annual Review of Entomology, 58, 57-78. |
| 20 | Escriche I, Kadar M, Juan-Borrás M, Domenech E (2011) Using flavonoids, phenolic compounds and headspace volatile profile for botanical authentication of lemon and orange honeys. Food Research International, 44, 1504-1513. |
| 21 | Fang Q, Huang SQ (2014) Progress in pollination ecology at the community level. Chinese Science Bulletin, 59, 449-458. (in Chinese with English abstract) |
| [方强, 黄双全 (2014) 群落水平上传粉生态学的研究进展. 科学通报, 59, 449-458.] | |
| 22 | Gómez-Rodríguez C, Crampton-Platt A, Timmermans MJTN, Baselga A, Vogler AP (2015) Validating the power of mitochondrial metagenomics for community ecology and phylogenetics of complex assemblages. Methods in Ecology and Evolution, 6, 883-894. |
| 23 | Hajibabaei M, Shokralla S, Zhou X, Singer GAC, Baird DJ (2011) Environmental Barcoding: A next-generation sequencing approach for biomonitoring Applications Using River Benthos. PLoS ONE, 6, e17497. |
| 24 | Hebert PD, Cywinska A, Ball SL (2003) Biological identifications through DNA barcodes. Proceedings of the Royal Society of London B: Biological Sciences, 270, 313-321. |
| 25 | Hollingsworth PM, Graham SW, Little DP (2011) Choosing and using a plant DNA barcode. PLoS ONE, 6, e19254. |
| 26 | Hollingsworth PM, Li DZ, van der Bank M, Twyford AD (2016) Telling plant species apart with DNA: From barcodes to genomes. Philosophical Transactions of the Royal Society B: Biological Sciences, 371, 20150338. |
| 27 | Holt KA, Bennett KD (2014) Principles and methods for automated palynology. New Phytologist, 203, 735-742. |
| 28 | Janzen DH, Hallwachs W, Burns J, Solis MA, Woodley NE (2009) Integration of DNA barcoding into an ongoing inventory of complex tropical biodiversity. Molecular Ecology Resources, 9, 1-26. |
| 29 | Kaiser-Bunbury CN, James M, Andrew EW, Terence V, Ronny G, Jens MO, Nico B (2017) Ecosystem restoration strengthens pollination network resilience and function. Nature, 542, 223-227. |
| 30 | Keller A, Danner N, Grimmer G, Ankenbrand M, von der Ohe K, von der Ohe W, Rost S, Härtel S, Steffan-Dewenter I (2015) Evaluating multiplexed next-generation sequencing as a method in palynology for mixed pollen samples. Plant Biology, 17, 558-566. |
| 31 | Khansari E, Zarre S, Alizadeh K, Attar F, Aghabeigi F, Salmaki Y (2012) Pollen morphology of Campanula (Campanulaceae) and allied genera in Iran with special focus on its systematic implication. Flora-Morphology, Distribution, Functional Ecology of Plants, 207, 203-211. |
| 32 | King C, Ballantyne G, Willmer PG (2013) Why flower visitation is a poor proxy for pollination: Measuring single-visit pollen deposition, with implications for pollination networks and conservation. Methods in Ecology and Evolution, 4, 811-818. |
| 33 | Klein AM, Vaissière BE, Cane JH, Steffandewenter I, Cunningham SA, Kremen C, Tscharntke T (2007) Importance of pollinators in changing landscapes for world crops. Proceedings of the Royal Society B: Biological Sciences, 274, 303-313. |
| 34 | Lahaye R, Savolainen V, Duthoit S, Maurin O, van der Bank M (2008) A test of psbK-psbI and atpF-atpH as potential plant DNA barcodes using the flora of the Kruger National Park as a model system (South Africa).. |
| 35 | Li XW, Yang Y, Henry RJ, Rossetto M, Wang YT, Chen SL (2015) Plant DNA barcoding: From gene to genome. Biological Reviews, 90, 157-166. |
| 36 | Li YW, Zhou X, Feng G, Hu HY, Niu LM, Hebert PD, Huang DW (2010) COI and ITS2 sequences delimit species, reveal cryptic taxa and host specificity of fig-associated Sycophila (Hymenoptera, Eurytomidae). Molecular Ecology Resources, 10, 31-40. |
| 37 | Liu SL, Li YY, Lu JL, Su X, Tang M, Zhang R, Zhou LL, Zhou CR, Yang Q, Ji YQ, Yu DW, Zhou X (2013) SOAPBarcode: Revealing arthropod biodiversity through assembly of Illumina shotgun sequences of PCR amplicons. Methods in Ecology and Evolution, 4, 1142-1150. |
| 38 | Liu SL, Wang X, Xie L, Tan MH, Li ZY, Su X, Zhang H, Misof B, Kjer KM, Tang M, Niehuis O, Jiang H, Zhou X (2016) Mitochondrial capture enriches mito-DNA 100 fold, enabling PCR-free mitogenomics biodiversity analysis. Molecular Ecology Resources, 16, 470. |
| 39 | Liu SL, Yang CT, Zhou CR, Zhou X (2017) Filling reference gaps via assembling DNA barcodes using high-throughput sequencing—moving toward barcoding the world. GigaScience, 6, 1-8. |
| 40 | Loreau M, Mazancourt CD (2013) Biodiversity and ecosystem stability: A synthesis of underlying mechanisms. Ecology Letters, 16, 106-115. |
| 41 | Macher JN, Zizka VMA, Weigand AM, Leese F (2017) A simple centrifugation protocol for metagenomic studies increases mitochondrial DNA yield by two orders of magnitude. Methods in Ecology and Evolution, 7, 1071-1075. |
| 42 | Matsuki Y, Isagi Y, Suyama Y (2007) The determination of multiple microsatellite genotypes and DNA sequences from a single pollen grain. Molecular Ecology Notes, 7, 194-198. |
| 43 | Murray TE, Úna F, Brown MJF, Paxton RJ (2008) Cryptic species diversity in a widespread bumble bee complex revealed using mitochondrial DNA RFLPs. Conservation Genetics, 9, 653-666. |
| 44 | Parks M, Cronn R, Liston A (2009) Increasing phylogenetic resolution at low taxonomic levels using massively parallel sequencing of chloroplast genomes. BMC Biology, 7, 84. |
| 45 | Pecnikar ZF, Buzan EV (2014) 20 years since the introduction of DNA barcoding: From theory to application. Journal of Applied Genetics, 55, 43-52. |
| 46 | Piñol J, Mir G, Gomez-Polo P, Agustí N (2015) Universal and blocking primer mismatches limit the use of high-throughput DNA sequencing for the quantitative metabarcoding of arthropods. Molecular Ecology Resources, 15, 819-830. |
| 47 | Popic TJ, Wardle GM, Davila YC (2013) Flower-visitor networks only partially predict the function of pollen transport by bees. Austral Ecology, 38, 76-86. |
| 48 | Pornon A, Escaravage N, Burrus M, Holota H, Khimoun A, Mariette J, Pellizzari C, Iribar A, Etienne R, Taberlet P, Vidal M, Winterton P, Zinger L, Andalo C (2016) Using metabarcoding to reveal and quantify plant-pollinator interactions. Scientific Reports, 6, 27282. |
| 49 | Potts SG, Biesmeijer JC, Kremen C, Neumann P, Schweiger O, Kunin WE (2010) Global pollinator declines: Trends, impacts and drivers. Trends in Ecology and Evolution, 25, 345-353. |
| 50 | Potts SG, Imperatriz-Fonseca V, Ngo HT, Aizen MA, Biesmeijer JC, Breeze TD, Dicks LV, Garibaldi LA, Hill R, Settele J, Vanbergen AJ (2016) Safeguarding pollinators and their values to human well-being. Nature, 540, 220-229. |
| 51 | Rahl M (2008) Microscopic identification and purity determination of pollen grains. Humana Press, 138, 263-269. |
| 52 | Raphaël C, Tony D, Alice V, Nicolas R, Jean-Claude R, Aurélie B, Taberlet P, Pont D (2016) Spatial representativeness of environmental DNA metabarcoding signal for fish biodiversity assessment in a natural freshwater system. PLoS ONE, 11, e0157366. |
| 53 | Rasmont P, Coppée A, Michez D, Meulemeester TD (2008) An overview of the Bombus terrestris (L.1758) subspecies (Aymenoptera: Apidae). Annales-Societe Entomologique de France, 44, 243-250. |
| 54 | Richardson RT, Lin CH, Quijia JO, Riusech NS, Goodell K, Johnson RM (2015a) Rank-based characterization of pollen assemblages collected by honey bees using a multi-locus metabarcoding approach. Applications in Plant Sciences, 3, 1500043. |
| 55 | Richardson RT, Lin CH, Sponsler DB, Quijia JO, Goodell K, Johnson RM (2015b) Application of ITS2 metabarcoding to determine the provenance of pollen collected by honey bees in an agroecosystem. Applications in Plant Sciences, 3, 1400066. |
| 56 | Ricketts TH, Regetz J, Steffan-Dewenter I, Cunningham SA, Kremen C, Bogdanski A, Gemmill-Herren B, Greenleaf SS, Klein AM, Mayfield MM, Morandin LA, Ochieng A, Potts SG, Viana BF (2008) Landscape effects on crop pollination services: Are there general patterns? Ecology Letters, 11, 499-515. |
| 57 | Ruhsam M, Rai HS, Mathews S, Ross TG, Graham SW, Raubeson LA, Mei W, Thomas PI, Gardner MF, Ennos RA, Hollingsworth PM (2015) Does complete plastid genome sequencing improve species discrimination and phylogenetic resolution in Araucaria? Molecular Ecology Resources, 15, 1067-1078. |
| 58 | Schmidt S, Schmid-Egger C, Moriniere J, Haszprunar G, Hebert PDN (2015) DNA barcoding largely supports 250 years of classical taxonomy: Identifications for Central European bees (Hymenoptera, Apoidea partim). Molecular Ecology Resources, 15, 985-1000. |
| 59 | Sheffield CS, Hebert PDN, Kevan P, Packer L (2009) DNA barcoding a regional bee (Hymenoptera: Apoidea) fauna and its potential for ecological studies. Molecular Ecology Resources, 9, 196-207. |
| 60 | Shi ZY, Yang CQ, Hao MD, Wang XY, Ward RD, Zhang AB (2018) FuzzyID2: A software package for large dataset species identification via barcoding and metabarcoding using Hidden Markov models and fuzzy set methods. Molecular Ecology Resources, 18, 666-675. |
| 61 | Smart MD, Cornman RS, Iwanowicz DD, McDermott- Kubeczko M, Pettis JS, Spivak MS, Otto CRV (2017) A comparison of honey bee-collected pollen from working agricultural lands using light microscopy and its metabarcoding. Environmental Entomology, 46, 38-49. |
| 62 | Smith MA, Woodley NE, Janzen DH, Hallwachs W, Hebert PDN (2006) DNA barcodes reveal cryptic host-specificity within the presumed polyphagous members of a genus of parasitoid flies (Diptera: Tachinidae). Proceedings of the National Academy of Sciences, USA, 103, 3657-3662. |
| 63 | Taberlet P, Coissac E, Pompanon F, Brochmann C, Willerslev E (2012) Towards next-generation biodiversity assessment using DNA metabarcoding. Molecular Ecology, 21, 2045-2050. |
| 64 | Tang M, Hardman CJ, Ji YQ, Meng GL, Liu SL, Tan MH, Yang SZ, Moss ED, Wang JX, Yang CX, Bruce C, Nevard T, Potts SG, Zhou X, Yu DW (2015) High-throughput monitoring of wild bee diversity and abundance via mitogenomics. Methods in Ecology and Evolution, 6, 1034-1043. |
| 65 | Tang M, Tan MH, Meng GL, Yang SZ, Su X, Liu SL, Song WH, Li YY, Wu Q, Zhang AB, Zhou X (2014) Multiplex sequencing of pooled mitochondrial genomes—a crucial step toward biodiversity analysis using mito-metagenomics. Nucleic Acids Research, 42, e166. |
| 66 | Tang M, Yi TS, Wang X, Tan MH, Zhou X (2013) The application of metabarcoding technology in identification of plant species diversity. Plant Diversity and Resources, 35, 769-773. (in Chinese with English abstract) |
| [唐敏, 伊廷双, 王欣, 谭美华, 周欣 (2013) Metabarcoding技术在植物鉴定和多样性研究中的应用. 植物分类与资源学报, 35, 769-773.] | |
| 67 | Weiner CN, Werner M, Linsenmair KE, Blüthgen N (2014) Land-use impacts on plant-pollinator networks: Interaction strength and specialization predict pollinator declines. Ecology, 95, 466-474. |
| 68 | Whitlock BA, Hale AM, Groff PA (2010) Intraspecific inversions pose a challenge for the trnH-psbA plant DNA barcode. PLoS ONE, 5, e11533. |
| 69 | Widmer A, Cozzolino S, Pellegrino G, Soliva M, Dafni A (2000) Molecular analysis of orchid pollinaria and pollinaria-remains found on insects. Molecular Ecology, 9, 1911-1914. |
| 70 | Wilson EE, Sidhu CS, LeVan KE, Holway DA (2010) Pollen foraging behavior of solitary Hawaiian bees revealed through molecular pollen analysis. Molecular Ecology, 19, 4823-4829. |
| 71 | Yamasaki YK, Nieman CC, Chang AN, Collier TC, Main BJ, Lee Y (2016) Improved tools for genomic DNA library construction of small insects. https://f1000research.com/posters/5-211. |
| 72 | Yu DW, Ji YQ, Emerson BC, Wang XY, Ye CX, Yang CY, Ding ZL (2012) Biodiversity soup: Metabarcoding of arthropods for rapid biodiversity assessment and biomonitoring. Methods in Ecology and Evolution, 3, 613-623. |
| 73 | Zhang AB, Hao MD, Yang CQ, Shi ZY (2017) BarcodingR: An integrated R package for species identification using DNA barcodes. Methods in Ecology and Evolution, 8, 627-634. |
| 74 | Zhou X, Li YY, Liu SL, Yang Q, Su X, Zhou LL, Tang M, Fu RB, Li JG, Huang QF (2013) Ultra-deep sequencing enables high-fidelity recovery of biodiversity for bulk arthropod samples without PCR amplification. GigaScience, 2, 4. |
| [1] | Lin Zhen, Xiang Jiabao, Cai Hejiayi, Gao Bei, Yang Jintao, Li Junyi, Zhou Qingsong, Huang Xiaolei, Deng Jun. Mitochondrial genomic data for seven Hemipteran species [J]. Biodiv Sci, 2025, 33(2): 24434-. |
| [2] | Hong Deng, Zhanyou Zhong, Chunni Kou, Shuli Zhu, Yuefei Li, Yuguo Xia, Zhi Wu, Jie Li, Weitao Chen. Population genetic structure and evolutionary history of Hemibagrus guttatus based on mitochondrial genomes [J]. Biodiv Sci, 2025, 33(1): 24241-. |
| [3] | Jiabei He, Ke Ke, Haiming Sun, Liping Hu, Xiaowei Zhao, Wenhao Wang, Qiang Zhao. Diet analysis of Neptunea cumingii using metabarcoding [J]. Biodiv Sci, 2025, 33(1): 24403-. |
| [4] | Suyan Ba, Chunyan Zhao, Yuan Liu, Qiang Fang. Constructing a pollination network by identifying pollen on insect bodies: Consistency between human recognition and an AI model [J]. Biodiv Sci, 2024, 32(6): 24088-. |
| [5] | Feifei Zhang, Tianfeng Yang, Lirong Chen, Dongmei Liu, Liuyuan Yang, Duyu Yang, Peng Ju, Lu Lu. Review of pollen color diversity in Angiosperms [J]. Biodiv Sci, 2024, 32(1): 23346-. |
| [6] | Xiaoqin Lü, Yang Li, Shunyu Wang, Renxiu Yao, Xiaoyue Wang. No significant differences found in chemical traits of pollen and nectar located in different positions across Aconitum piepunense racemes [J]. Biodiv Sci, 2024, 32(1): 23371-. |
| [7] | Zhiyuan Dong, Linlin Chen, Naipeng Zhang, Li Chen, Debin Sun, Yanmei Ni, Baoquan Li. Response of fish diversity to hydrological connectivity of typical tidal creek system in the Yellow River Delta based on environmental DNA metabarcoding [J]. Biodiv Sci, 2023, 31(7): 23073-. |
| [8] | Zhenjie Zhan, Chao Zhang, Minhao Chen, Jiadong Wang, Aihua Fu, Yuwei Fan, Xiaofeng Luan. DNA metabarcoding-based winter diet analysis of Eurasian otter (Lutra lutra) in the northern Greater Khingan Mountains [J]. Biodiv Sci, 2023, 31(6): 22586-. |
| [9] | Fan Wu, Shenyun Liu, Huqiang Jiang, Qian Wang, Kaiwei Chen, Hongliang Li. Pollination difference between Apis cerana cerana and Apis mellifera ligustica during the late autumn and winter [J]. Biodiv Sci, 2023, 31(5): 22528-. |
| [10] | Buqing Peng, Ling Tao, Jing Li, Ronghui Fan, Shunde Chen, Changkun Fu, Qiong Wang, Keyi Tang. DNA metabarcoding dietary analysis of six sympatric small mammals at the Laojunshan National Nature Reserve, Sichuan Province [J]. Biodiv Sci, 2023, 31(4): 22474-. |
| [11] | Jiajia Pu, Pingjun Yang, Yang Dai, Kexin Tao, Lei Gao, Yuzhou Du, Jun Cao, Xiaoping Yu, Qianqian Yang. Species identification and population genetic structure of non-native apple snails (Ampullariidea: Pomacea) in the lower reaches of the Yangtze River [J]. Biodiv Sci, 2023, 31(3): 22346-. |
| [12] | Jie Tao, Benqiang Li, Jinghua Cheng, Ying Shi, Peihong Liu, Guixia He, Weijie Xu, Huili Liu. Viral metagenome analysis of the viral community composition of the porcine diarrhea feaces [J]. Biodiv Sci, 2023, 31(11): 23170-. |
| [13] | Congcong Lu, Qian Liu, Xiaolei Huang. Mitochondrial genome data of three aphid species [J]. Biodiv Sci, 2022, 30(7): 22204-. |
| [14] | Mei Shen, Ningning Guo, Zunlan Luo, Xiaochen Guo, Guang Sun, Nengwen Xiao. Explore the distribution and influencing factors of fish in major rivers in Beijing with eDNA metabarcoding technology [J]. Biodiv Sci, 2022, 30(7): 22240-. |
| [15] | Junjie Zhai, Huifeng Zhao, Guangshen Shang, Zhenjun Sun, Yufeng Zhang, Xing Wang. Advances in earthworm genomics: Based on whole genome and mitochondrial genome [J]. Biodiv Sci, 2022, 30(12): 22257-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn