



Biodiv Sci ›› 2026, Vol. 34 ›› Issue (1): 25364. DOI: 10.17520/biods.2025364 cstr: 32101.14.biods.2025364
• Special Feature: Methods for Ecological Data Analysis • Previous Articles Next Articles
Jiqi Gu1, Jiangshan Lai2,3, Ying Wang4, Haoran Wu5, Xue Zhang6, Xiaotong Song7, Xiaoming Shao8,*( ), Anru Lou1,*(
), Anru Lou1,*( )
)
Received:2025-09-09
Accepted:2026-01-08
Online:2026-01-20
Published:2026-02-06
Contact:
Xiaoming Shao, Anru Lou
Supported by:Jiqi Gu, Jiangshan Lai, Ying Wang, Haoran Wu, Xue Zhang, Xiaotong Song, Xiaoming Shao, Anru Lou. Theoretical foundations, methodological advances, and applications of joint species distribution models with a focus on the HMSC framework in ecology[J]. Biodiv Sci, 2026, 34(1): 25364.
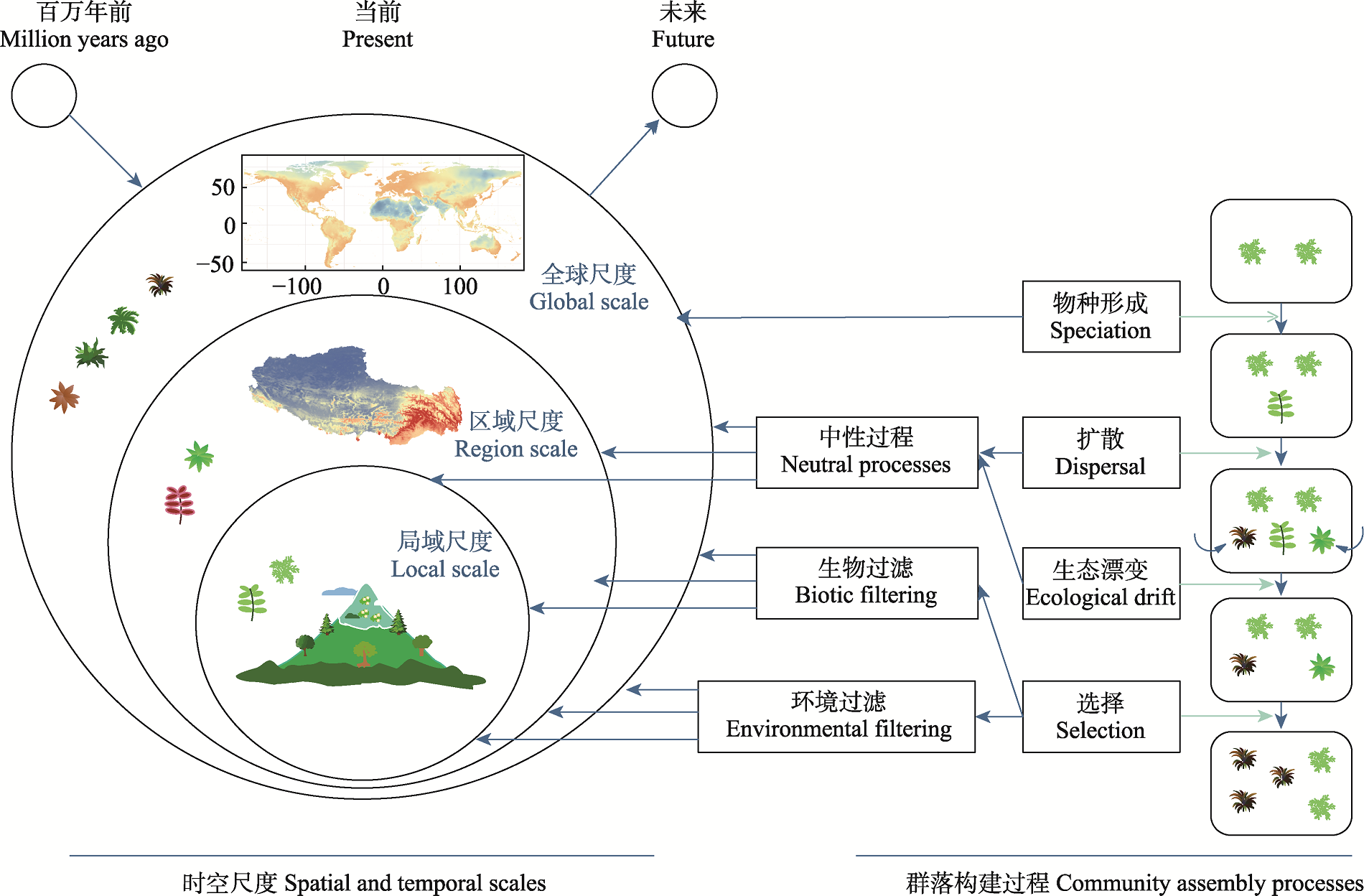
Fig. 1 Schematic diagram of the multi-scale framework of community assembly processes and corresponding types of observational data. The figure illustrates the different temporal and spatial scales at which community assembly processes occur, including global, regional, and local scales. Community assembly processes can be divided into speciation, neutral processes (dispersal, ecological drift, etc.), biotic filtering (species interactions), and environmental filtering (selection by abiotic factors). Blue arrows indicate the direction of mechanisms, and green arrows denote the pathways of different community assembly processes.
| 索引及其范围 Index and range | 含义说明 Description |
|---|---|
| i = 1,..., n | 样方(采样单元) Sampling plots (Sampling units) |
| j = 1,..., ns | 物种 Species |
| k = 1,..., nc | 环境协变量 Environmental covariates |
| l = 1,..., nt | 物种性状 Species traits |
| h = 1,..., nf | 潜在因子 Latent factors |
| u = 1,..., nᵤ | 层级单元 Hierarchical units |
| q = 1,..., d | 空间坐标维度 Spatial coordinate dimensions |
| r = 1,..., nᵣ | 随机效应 Random effects |
Table 1 Indices and their ranges in the HMSC framwork
| 索引及其范围 Index and range | 含义说明 Description |
|---|---|
| i = 1,..., n | 样方(采样单元) Sampling plots (Sampling units) |
| j = 1,..., ns | 物种 Species |
| k = 1,..., nc | 环境协变量 Environmental covariates |
| l = 1,..., nt | 物种性状 Species traits |
| h = 1,..., nf | 潜在因子 Latent factors |
| u = 1,..., nᵤ | 层级单元 Hierarchical units |
| q = 1,..., d | 空间坐标维度 Spatial coordinate dimensions |
| r = 1,..., nᵣ | 随机效应 Random effects |
| 数据矩阵 Data matrix | 数据维度 Data dimension | 含义说明 Description |
|---|---|---|
| 𝐘, 元素 𝐘, elements | 群落数据 Community data | |
| 𝐗, 元素 𝐗, elements | 环境数据 Environmental data | |
| 𝐓, 元素 𝐓, elements | 物种性状数据 Species trait data | |
| 𝐂, 元素 𝐂, elements | 系统发育数据 Phylogenetic data | |
| 𝚷, 元素 𝚷, elements | 研究设计 Study design | |
| 𝐒, 元素 𝐒, elements | 空间坐标 Spatial coordinates |
Table 2 Data matrices and their dimensions in the core model of HMSC framework
| 数据矩阵 Data matrix | 数据维度 Data dimension | 含义说明 Description |
|---|---|---|
| 𝐘, 元素 𝐘, elements | 群落数据 Community data | |
| 𝐗, 元素 𝐗, elements | 环境数据 Environmental data | |
| 𝐓, 元素 𝐓, elements | 物种性状数据 Species trait data | |
| 𝐂, 元素 𝐂, elements | 系统发育数据 Phylogenetic data | |
| 𝚷, 元素 𝚷, elements | 研究设计 Study design | |
| 𝐒, 元素 𝐒, elements | 空间坐标 Spatial coordinates |
| 类别 Category | 参数 Parameter | 类型 Type | 含义 Description |
|---|---|---|---|
| 固定效应 Fixed effect | LF, 元素 LF, elements | 固定效应的线性预测量 Linear predictor of fixed effects | |
| 固定效应 Fixed effect | B, 元素 B, elements | 物种生态位 Species ecological niches | |
| 固定效应 Fixed effect | M, 元素 M, elements | 基于性状的物种生态位期望值 Trait‐based expected values of species niches | |
| 固定效应 Fixed effect | ρ | 标量 Scalar | 物种生态位的系统发育信号 Phylogenetic signal in species niches |
| 固定效应 Fixed effect | Γ, 元素 Γ, elements | 性状对生态位的影响 Effects of traits on species niches | |
| 固定效应 Fixed effect | V, 元素 V, elements | 物种生态位的残差协方差 Residual covariance of species niches | |
| 随机效应 Random effect | Lᴿ, 元素 Lᴿ, elements | 随机效应的线性预测量 Linear predictor of random effects | |
| 随机效应 Random effect | H, 元素 H, elements | 样地载荷 Site loadings | |
| 随机效应 Random effect | α, 元素 α, elements | 长度为 Vector of length | 样地载荷的空间尺度 Spatial scale of site loadings |
| 随机效应 Random effect | Λ, 元素 Λ, elements | 物种载荷 Species loadings | |
| 随机效应 Random effect | Ω, 元素 Ω, elements | 物种间的关联关系 Interspecific association matrix | |
| 随机效应 Random effect | Φ, 元素 Φ, elements | 物种载荷的局部收缩项 Local shrinkage parameters of species loadings | |
| 随机效应 Random effect | δ, 元素 δ, elements | 长度为 vector of length | 物种载荷的全局收缩项 Global shrinkage parameters of species loadings |
| 数据模型 Data model | L, 元素 L, elements | 线性预测量 Linear predictor | |
| 数据模型 Data model | Σ, 元素 Σ, elements | 残差方差 Residual variances |
Table 3 Parameters in the core model of the HMSC framework and their interpretations
| 类别 Category | 参数 Parameter | 类型 Type | 含义 Description |
|---|---|---|---|
| 固定效应 Fixed effect | LF, 元素 LF, elements | 固定效应的线性预测量 Linear predictor of fixed effects | |
| 固定效应 Fixed effect | B, 元素 B, elements | 物种生态位 Species ecological niches | |
| 固定效应 Fixed effect | M, 元素 M, elements | 基于性状的物种生态位期望值 Trait‐based expected values of species niches | |
| 固定效应 Fixed effect | ρ | 标量 Scalar | 物种生态位的系统发育信号 Phylogenetic signal in species niches |
| 固定效应 Fixed effect | Γ, 元素 Γ, elements | 性状对生态位的影响 Effects of traits on species niches | |
| 固定效应 Fixed effect | V, 元素 V, elements | 物种生态位的残差协方差 Residual covariance of species niches | |
| 随机效应 Random effect | Lᴿ, 元素 Lᴿ, elements | 随机效应的线性预测量 Linear predictor of random effects | |
| 随机效应 Random effect | H, 元素 H, elements | 样地载荷 Site loadings | |
| 随机效应 Random effect | α, 元素 α, elements | 长度为 Vector of length | 样地载荷的空间尺度 Spatial scale of site loadings |
| 随机效应 Random effect | Λ, 元素 Λ, elements | 物种载荷 Species loadings | |
| 随机效应 Random effect | Ω, 元素 Ω, elements | 物种间的关联关系 Interspecific association matrix | |
| 随机效应 Random effect | Φ, 元素 Φ, elements | 物种载荷的局部收缩项 Local shrinkage parameters of species loadings | |
| 随机效应 Random effect | δ, 元素 δ, elements | 长度为 vector of length | 物种载荷的全局收缩项 Global shrinkage parameters of species loadings |
| 数据模型 Data model | L, 元素 L, elements | 线性预测量 Linear predictor | |
| 数据模型 Data model | Σ, 元素 Σ, elements | 残差方差 Residual variances |
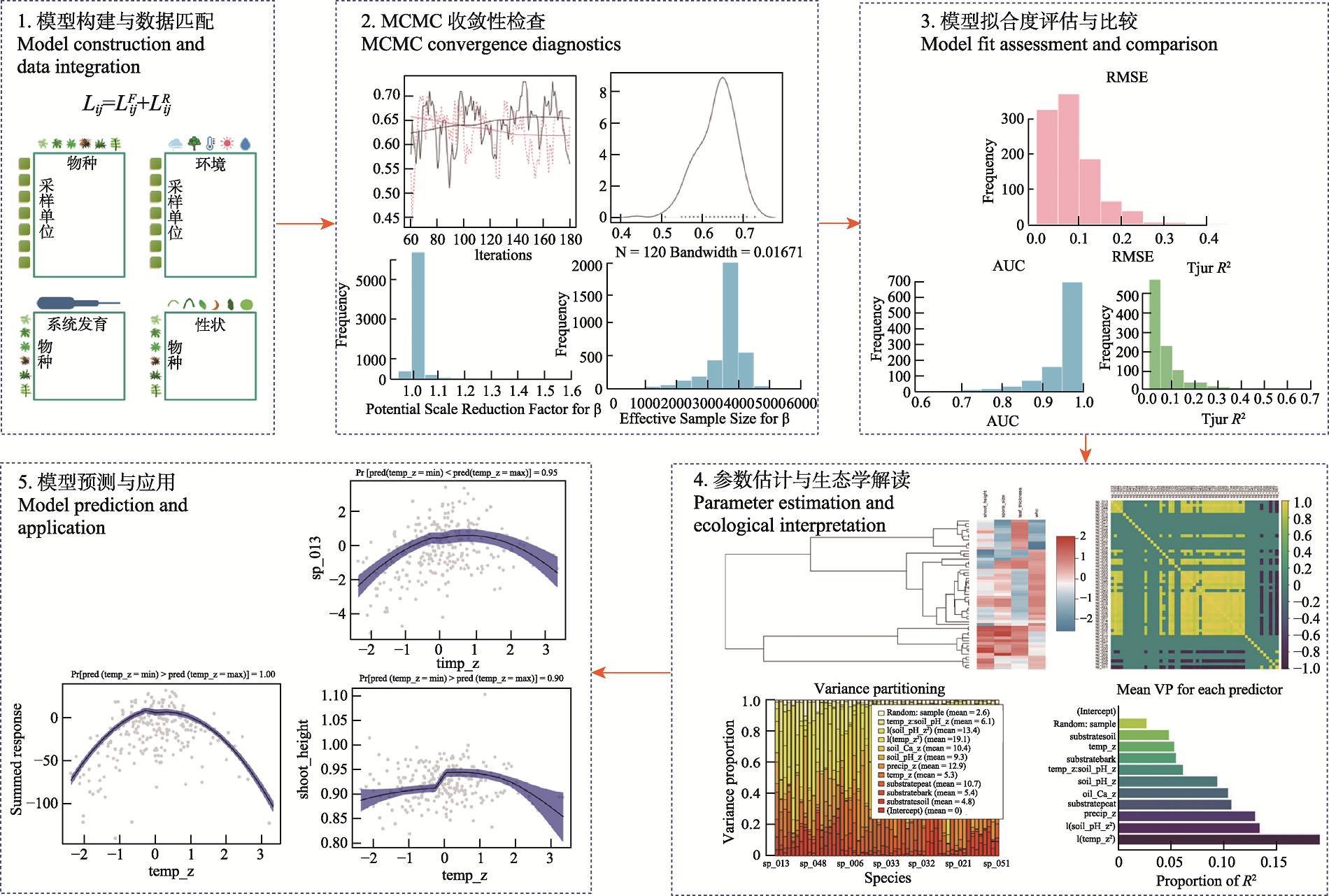
Fig. 2 Schematic overview of the complete analytical workflow and outputs of hierarchical modelling of species communities (HMSC) framework. The figure illustrates five key steps of HMSC-based modelling and inference: (1) model construction and data integration, in which species distribution data are jointly modelled with environmental variables, functional traits, phylogenetic relationships, and spatial random effects within a unified hierarchical Bayesian framework; (2) MCMC convergence diagnostics, where trace plots, posterior distributions, effective sample size (ESS), and potential scale reduction factors (PSRF) are used to assess convergence and fitness parameter mixing; (3) model fit evaluation and comparison, in which predictive performance is quantified using root mean square error (RMSE), area under curve (AUC), and R²; (4) parameter estimation and ecological interpretation, including environmental response parameters, trait and phylogenetic effects, residual correlation structures, and variance partitioning; and (5) model prediction and application, illustrating species responses to key environmental gradients with associated uncertainty intervals for community-level prediction and scenario-based analyses.
| [1] | Abrego N, Niittynen P, Kemppinen J, Ovaskainen O (2025) Joint species-trait distribution modeling: The role of intraspecific trait variation in community assembly. Ecology, 106, e70174. |
| [2] | Abrego N, Norberg A, Ovaskainen O (2017) Measuring and predicting the influence of traits on the assembly processes of wood-inhabiting fungi. Journal of Ecology, 105, 1070-1081. |
| [3] | Agrawal AA, Ackerly DD, Adler F, Arnold AE, Cáceres C, Doak DF, Post E, Hudson PJ, Maron J, Mooney KA, Power M, Schemske D, Stachowicz J, Strauss S, Turner MG, Werner E (2007) Filling key gaps in population and community ecology. Frontiers in Ecology and the Environment, 5, 145-152. |
| [4] | Anderson MJ, Walsh DCI (2013) PERMANOVA, ANOSIM, and the Mantel test in the face of heterogeneous dispersions: What null hypothesis are you testing? Ecological Monographs, 83, 557-574. |
| [5] | Begon M, Townsend CR (2020) Ecology: From Individuals to Ecosystems, 5th edn. John Wiley & Sons, Hoboken, USA. |
| [6] | Blanchet FG, Cazelles K, Gravel D (2020) Co-occurrence is not evidence of ecological interactions. Ecology Letters, 23, 1050-1063. |
| [7] | Bohmann K, Evans A, Gilbert MTP, Carvalho GR, Creer S, Knapp M, Yu DW, de Bruyn M (2014) Environmental DNA for wildlife biology and biodiversity monitoring. Trends in Ecology & Evolution, 29, 358-367. |
| [8] | Breiner FT, Guisan A, Bergamini A, Nobis MP (2015) Overcoming limitations of modelling rare species by using ensembles of small models. Methods in Ecology and Evolution, 6, 1210-1218. |
| [9] | Calabrese JM, Certain G, Kraan C, Dormann CF (2014) Stacking species distribution models and adjusting bias by linking them to macroecological models. Global Ecology and Biogeography, 23, 99-112. |
| [10] | Chase JM, Myers JA (2011) Disentangling the importance of ecological niches from stochastic processes across scales. Philosophical Transactions of the Royal Society of London Series B, Biological Sciences, 366, 2351-2363. |
| [11] | Clark JS, Nemergut D, Seyednasrollah B, Turner PJ, Zhang S (2017) Generalized joint attribute modeling for biodiversity analysis: Median-zero, multivariate, multifarious data. Ecological Monographs, 87, 34-56. |
| [12] | Cranston A, Cooper N, Bro-Jørgensen J (2024) Using joint species distribution modelling to identify climatic and non-climatic drivers of Afrotropical ungulate distributions. Ecography, 2024, e07209. |
| [13] | Dormann CF, Bobrowski M, Dehling DM, Harris DJ, Hartig F, Lischke H, Moretti MD, Pagel J, Pinkert S, Schleuning M, Schmidt SI, Storch I, Weiss IS, Mueller J (2018) Biotic interactions in species distribution modelling: 15 challenges and a way forward. Ecography, 41, 1-15. |
| [14] | Doser JW, Finley AO, Kéry M, Zipkin EF (2022) spOccupancy: An R package for single-species, multi-species, and integrated spatial occupancy models. Methods in Ecology and Evolution, 13, 1670-1678. |
| [15] | Dray S, Legendre P (2008) Testing the species traits-environment relationships: The fourth-corner problem revisited. Ecology, 89, 3400-3412. |
| [16] | Elith J, Leathwick JR (2009) Species distribution models: Ecological explanation and prediction across space and time. Annual Review of Ecology, Evolution, and Systematics, 40, 677-697. |
| [17] | Epele LB, Williams-Subiza EA, Bird MS, Boissezon A, Boix D, Demierre E, Fair CG, García PE, Gascón S, Grech MG, Greig HS, Jeffries M, Kneitel JM, Loskutova O, Maltchik L, Manzo LM, Mataloni G, McLean K, Mlambo MC, Oertli B, Pires MM, Sala J, Scheibler EE, Stenert C, Wu HT, Wissinger SA, Batzer DP (2024) A global assessment of environmental and climate influences on wetland macroinvertebrate community structure and function. Global Change Biology, 30, e17173. |
| [18] | Escamilla Molgora JM, Sedda L, Diggle PJ, Atkinson PM (2022) A taxonomic-based joint species distribution model for presence-only data. Journal of the Royal Society, Interface 19, 20210681. |
| [19] | Fang XN, Zhang P, Xing QW, Chen XJ, Cao J, Zhang H, Yu W (2025) The impact of varying spatiotemporal scales on different joint species distribution models: A case study of pelagic fish species in the northwest Pacific Ocean. Journal of Biogeography, 52, e15154. |
| [20] | Fick SE, Hijmans RJ (2017) WorldClim 2: New 1-km spatial resolution climate surfaces for global land areas. International Journal of Climatology, 37, 4302-4315. |
| [21] | Galante PJ, Alade B, Muscarella R, Jansa SA, Goodman SM, Anderson RP (2018) The challenge of modeling niches and distributions for data-poor species: A comprehensive approach to model complexity. Ecography, 41, 726-736. |
| [22] | Gilarranz LJ, Rayfield B, Liñán-Cembrano G, Bascompte J, Gonzalez A (2017) Effects of network modularity on the spread of perturbation impact in experimental metapopulations. Science, 357, 199-201. |
| [23] | Gotelli NJ (2000) Null model analysis of species co-occurrence patterns. Ecology, 81, 2606-2621. |
| [24] | Gotelli N, Hart E, Ellison A, Hart ME (2015) Package ‘EcoSimR’. R Package. Available at: https://CRAN.R-project.org/package=EcoSimR. (accessed on 2026-01-10) |
| [25] | Götzenberger L, de Bello F, Bråthen KA, Davison J, Dubuis A, Guisan A, Lepš J, Lindborg R, Moora M, Pärtel M, Pellissier L, Pottier J, Vittoz P, Zobel K, Zobel M (2012) Ecological assembly rules in plant communities— Approaches, patterns and prospects. Biological Reviews, 87, 111-127. |
| [26] | Guisan A, Thuiller W (2005) Predicting species distribution: Offering more than simple habitat models. Ecology Letters, 8, 993-1009. |
| [27] | Huang EH, Ji CJ, Liang MX, Zhu JL, Tang ZY, Fang JY (2025) Climatic and non-climatic effects on species occurrence and abundance shift in different trends along elevational gradients. Journal of Plant Ecology, 18, rtaf122. |
| [28] | Hui FKC (2016) Boral-Bayesian ordination and regression analysis of multivariate abundance data in R. Methods in Ecology and Evolution, 7, 744-750. |
| [29] | Hui FKC, Taskinen S, Pledger S, Foster SD, Warton DI (2015) Model-based approaches to unconstrained ordination. Methods in Ecology and Evolution, 6, 399-411. |
| [30] | Itter MS, Kaarlejärvi E, Laine AL, Hamberg L, Tonteri T, Vanhatalo J (2024) Bayesian joint species distribution model selection for community-level prediction. Global Ecology and Biogeography, 33, e13827. |
| [31] | Jeliazkov A, Gavish Y, Marsh CJ, Geschke J, Brummitt N, Rocchini D, Haase P, Kunin WE, Henle K (2022) Sampling and modelling rare species: Conceptual guidelines for the neglected majority. Global Change Biology, 28, 3754-3777. |
| [32] | Kattge J, Diaz S, Lavorel S, Prentice IC, Leadley P, Bönisch G, Garnier E, Westoby M, Reich PB, Wright IJ (2011) TRY—A global database of plant traits. Global Change Biology, 17, 2905-2935. |
| [33] | Korhonen P, Hui FKC, Niku J, Taskinen S, van der Veen B (2024) A comparison of joint species distribution models for percent cover data. Methods in Ecology and Evolution, 15, 2359-2372. |
| [34] | Kraft NJB, Adler PB, Godoy O, James EC, Fuller S, Levine JM (2015) Community assembly, coexistence and the environmental filtering metaphor. Functional Ecology, 29, 592-599. |
| [35] | Legendre P, Galzin R, Harmelin-Vivien ML (1997) Relating behavior to habitat: Solutions to the fourth-corner problem. Ecology, 78, 547-562. |
| [36] | Legendre P, Legendre L (2012) Numerical Ecology. Elsevier, Amsterdam. |
| [37] | Leibold MA, Holyoak M, Mouquet N, Amarasekare P, Chase JM, Hoopes MF, Holt RD, Shurin JB, Law R, Tilman D, Loreau M, Gonzalez A (2004) The metacommunity concept: A framework for multi-scale community ecology. Ecology Letters, 7, 601-613. |
| [38] | Liu C, Van Meerbeek K (2024) Predicting the responses of European grassland communities to climate and land cover change. Philosophical Transactions of the Royal Society of London Series B, Biological Sciences, 379, 20230335. |
| [39] | Logue JB, Mouquet N, Peter H, Hillebrand H, (2011) Empirical approaches to metacommunities: A review and comparison with theory. Trends in Ecology & Evolution, 26, 482–491. |
| [40] | Magurran AE, Mcgill BJ (2010) Biological Diversity:Frontiers in Measurement and Assessment. OUP Oxford. Oxford University Press, Oxford. |
| [41] | McArdle BH, Anderson MJ (2001) Fitting multivariate models to community data: A comment on distance-based redundancy analysis. Ecology, 82, 290-297. |
| [42] | McLaughlin P, Krause K, Maloney K, Woods T, Wagner T (2024) Evaluating the effectiveness of joint species distribution modeling for freshwater fish communities within large watersheds. Canadian Journal of Fisheries and Aquatic Sciences, 81, 1248-1263. |
| [43] | Niku J, Hui FKC, Taskinen S, Warton DI (2019) Gllvm: Fast analysis of multivariate abundance data with generalized linear latent variable models in R. Methods in Ecology and Evolution, 10, 2173-2182. |
| [44] | Norberg A, Abrego N, Blanchet FG, Adler FR, Anderson BJ, Anttila J, Araújo MB, Dallas T, Dunson D, Elith J, Foster SD, Fox R, Franklin J, Godsoe W, Guisan A, O’Hara B, Hill NA, Holt RD, Hui FKC, Husby M, Kålås JA, Lehikoinen A, Luoto M, Mod HK, Newell G, Renner I, Roslin T, Soininen J, Thuiller W, Vanhatalo J, Warton D, White M, Zimmermann NE, Gravel D, Ovaskainen O (2019) A comprehensive evaluation of predictive performance of 33 species distribution models at species and community levels. Ecological Monographs, 89, e01370. |
| [45] | Odriozola I, Abrego N, Tláskal V, Zrůstová P, Morais D, Větrovský T, Ovaskainen O, Baldrian P (2021) Fungal communities are important determinants of bacterial community composition in deadwood. mSystems, 6, e01017-e01020. |
| [46] | Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O Hara RB, Simpson GL, Solymos P, Stevens MHH, Wagner H (2013) Package ‘vegan’. Community Ecology Package, Version 2, 1-295. https://cran.r-project.org/. (accessed on 2026-01-10) |
| [47] | Ovaskainen O, Abrego N (2020) Joint Species Distribution Modelling. Cambridge University Press, Cambridge. |
| [48] | Ovaskainen O, Hottola J, Siitonen J (2010) Modeling species co-occurrence by multivariate logistic regression generates new hypotheses on fungal interactions. Ecology, 91, 2514-2521. |
| [49] | Ovaskainen O, Rybicki J, Abrego N (2019) What can observational data reveal about metacommunity processes? Ecography, 42, 1877-1886. |
| [50] | Ovaskainen O, Soininen J (2011) Making more out of sparse data: Hierarchical modeling of species communities. Ecology, 92, 289-295. |
| [51] | Ovaskainen O, Tikhonov G, Norberg A, Guillaume Blanchet F, Duan L, Dunson D, Roslin T, Abrego N (2017) How to make more out of community data? A conceptual framework and its implementation as models and software. Ecology Letters, 20, 561-576. |
| [52] | Ovaskainen O, Winter S, Tikhonov G, Abrego N, Anslan S, de Waard JR, de Waard SL, Fisher BL, Furneaux B, Hardwick B, Kerdraon D, Pentinsaari M, Raharinjanahary D, Rajoelison ET, Ratnasingham S, Somervuo P, Sones JE, Zakharov EV, Hebert PDN, Roslin T, Dunson D (2025) Common to rare transfer learning (CORAL) enables inference and prediction for a quarter million rare Malagasy arthropods. Nature Methods, 22, 2074-2082. |
| [53] | Pichler M, Hartig F (2021) A new joint species distribution model for faster and more accurate inference of species associations from big community data. Methods in Ecology and Evolution, 12, 2159-2173. |
| [54] | Poggiato G, Münkemüller T, Bystrova D, Arbel J, Clark JS, Thuiller W (2021) On the interpretations of joint modeling in community ecology. Trends in Ecology & Evolution, 36, 391-401. |
| [55] | Pollock LJ, Tingley R, Morris WK, Golding N, O’Hara RB, Parris KM, Vesk PA, McCarthy MA (2014) Understanding co-occurrence by modelling species simultaneously with a Joint Species Distribution Model (JSDM). Methods in Ecology and Evolution, 5, 397-406. |
| [56] | Powell-Romero F, Fountain-Jones NM, Norberg A, Clark NJ (2023) Improving the predictability and interpretability of co-occurrence modelling through feature-based joint species distribution ensembles. Methods in Ecology and Evolution, 14, 146-161. |
| [57] | Rahman AU, Tikhonov G, Oksanen J, Rossi T, Ovaskainen O (2024) Accelerating joint species distribution modelling with Hmsc-HPC by GPU porting. PLoS Computational Biology, 20, e1011914. |
| [58] | Runquist RDB, Lake TA, Moeller DA (2021) Improving predictions of range expansion for invasive species using joint species distribution models and surrogate co-occurring species. Journal of Biogeography, 48, 1693-1705. |
| [59] | Sallinen S, Norberg A, Susi H, Laine AL (2020) Intraspecific host variation plays a key role in virus community assembly. Nature Communications, 11, 5610. |
| [60] | Schliep EM, Lany NK, Zarnetske PL, Schaeffer RN, Orians CM, Orwig DA, Preisser EL (2018) Joint species distribution modelling for spatio-temporal occurrence and ordinal abundance data. Global Ecology and Biogeography, 27, 142-155. |
| [61] | Schmitt S, Pouteau R, Justeau D, de Boissieu F, Birnbaum P (2017) ssdm: An R package to predict distribution of species richness and composition based on stacked species distribution models. Methods in Ecology and Evolution, 8, 1795-1803. |
| [62] | Smith JA, Johnson DD (2024) Evaluating drivers and predictability of catch composition in a highly mixed trawl fishery using stacked and joint species distribution models. Fisheries Research, 279, 107151. |
| [63] | Stephenson F, Bowden DA, Rowden AA, Anderson OF, Clark MR, Bennion M, Finucci B, Pinkerton MH, Goode S, Chin C, Davey N, Hart A, Stewart R (2024) Using joint species distribution modelling to predict distributions of seafloor taxa and identify vulnerable marine ecosystems in New Zealand waters. Biodiversity and Conservation, 33, 3103-3127. |
| [64] | Thorson JT, Ianelli JN, Larsen EA, Ries L, Scheuerell MD, Szuwalski C, Zipkin EF (2016) Joint dynamic species distribution models: A tool for community ordination and spatio-temporal monitoring. Global Ecology and Biogeography, 25, 1144-1158. |
| [65] | Tikhonov G, Duan L, Abrego N, Newell G, White M, Dunson D, Ovaskainen O (2020) Computationally efficient joint species distribution modeling of big spatial data. Ecology, 101, e02929. |
| [66] | Tikhonov G, Opedal ØH, Abrego N, Lehikoinen A, de Jonge MMJ, Oksanen J, Ovaskainen O (2020) Joint species distribution modelling with the R-package HMSC. Methods in Ecology and Evolution, 11, 442-447. |
| [67] | Traylor CR, Ulyshen MD, McHugh JV, Burner RC (2024) Forest age is a primary trait filter for saproxylic beetles in the southeastern United States. Forest Ecology and Management, 553, 121545. |
| [68] | Vellend M (2010) Conceptual synthesis in community ecology. The Quarterly Review of Biology, 85, 183-206. |
| [69] | Violet C, Boyé A, Chevalier M, Gauthier O, Grall J, Marzloff MP (2022) A multifaceted framework to assess tradeoffs in interpretability, Explanatory and Predictive Performances of Alternative Joint Species Distribution Models. bioRxiv, https://doi.org/10.1101/2022.12.19.519605. (accessed on 2026-01-10) |
| [70] | Wang HY, Chen XL, Chen SC (2025) Chinese Seed Trait Database: A curated resource for diaspore traits in the Chinese flora. New Phytologist, 248, 11-16. |
| [71] | Warton DI, Blanchet FG, O’Hara RB, Ovaskainen O, Taskinen S, Walker SC, Hui FKC (2015) So many variables: Joint modeling in community ecology. Trends in Ecology & Evolution, 30, 766-779. |
| [72] | Weigel B, Graco-Roza C, Hultman J, Pajunen V, Teittinen A, Kuzmina M, Zakharov EV, Soininen J, Ovaskainen O (2023) Local eukaryotic and bacterial stream community assembly is shaped by regional land use effects. ISME Communications, 3, 65. |
| [73] | Weiss S, Van Treuren W, Lozupone C, Faust K, Friedman J, Deng Y, Xia LC, Xu ZZ, Ursell L, Alm EJ, Birmingham A, Cram JA, Fuhrman JA, Raes J, Sun FZ, Zhou JZ, Knight R (2016) Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. The ISME Journal, 10, 1669-1681. |
| [74] | Wilkinson DP, Golding N, Guillera-Arroita G, Tingley R, McCarthy MA (2019) A comparison of joint species distribution models for presence-absence data. Methods in Ecology and Evolution, 10, 198-211. |
| [75] | Wilkinson DP, Golding N, Guillera-Arroita G, Tingley R, McCarthy MA (2021) Defining and evaluating predictions of joint species distribution models. Methods in Ecology and Evolution, 12, 394-404. |
| [76] | Wisz MS, Pottier J, Kissling WD, Pellissier L, Lenoir J, Damgaard CF, Dormann CF, Forchhammer MC, Grytnes JA, Guisan A, Heikkinen RK, Høye TT, Kühn I, Luoto M, Maiorano L, Nilsson MC, Normand S, Öckinger E, Schmidt NM, Termansen M, Timmermann A, Wardle DA, Aastrup P, Svenning JC (2013) The role of biotic interactions in shaping distributions and realised assemblages of species: Implications for species distribution modelling. Biological Reviews, 88, 15-30. |
| [77] | Zobel M (1997) The relative of species pools in determining plant species richness: An alternative explanation of species coexistence? Trends in Ecology & Evolution, 12, 266-269. |
| [78] | Zhang JT (2011) Quantitative Ecology, 2nd edn. Science Press, Beijing. (in Chinese) |
| [张金屯 (2011) 数量生态学(第二版). 科学出版社, 北京.] | |
| [79] | Zhu YJ, Shan D, Zhang X, Liu YS, Shi ZJ, Yang XH (2018) Advances in joint species distribution models to reveal community structure and its environmental response. Chinese Journal of Applied Ecology, 29, 4217-4225. (in Chinese with English abstract) |
| [朱媛君, 山丹, 张晓, 刘艳书, 时忠杰, 杨晓晖 (2018) 揭示群落结构及其环境响应的联合物种分布模型的研究进展. 应用生态学报, 29, 4217-4225.] |
| [1] | Yi Zou. Alpha-diversity index selection: Simulation comparison under unequal sampling [J]. Biodiv Sci, 2026, 34(1): 25278-. |
| [2] | Changqiao Chen, Yanfei Feng, Liqi Lu, Huaijiang He, Chunyu Zhang, Xiuhai Zhao, Minhui Hao. β-diversity pattern and mechanism in a conifer-broadleaf mixed forest in Jiaohe, Jilin Province [J]. Biodiv Sci, 2025, 33(12): 25169-. |
| [3] | Guoshan Shi, Feng Liu, Guanghong Cao, Dian Chen, Shangwen Xia, Yun Deng, Bin Wang, Xiaodong Yang, Luxiang Lin. Beta diversity of woody plants in a tropical seasonal rainforest at Xishuangbanna: Roles of space, environment, and forest stand structure [J]. Biodiv Sci, 2024, 32(12): 24285-. |
| [4] | Yanping Wang, Minchu Zhang, Chengxiu Zhan. A review on the nested distribution pattern (nestedness): Analysis methods, mechanisms and conservation implications [J]. Biodiv Sci, 2023, 31(12): 23314-. |
| [5] | Mengjun Qu, Nueryila·Ababaike , Xuge Zou, Hang Zhao, Weilin Zhu, Jianming Wang, Jingwen Li. Influence of geographic distance and environmental factors on beta diversity of plants in the Alxa gobi region in northern China [J]. Biodiv Sci, 2022, 30(11): 22029-. |
| [6] | Jian Zhang, Hongzhi Kong, Xiaolei Huang, Shenglei Fu, Liangdong Guo, Qinghua Guo, Fumin Lei, Zhi Lü, Yurong Zhou, Keping Ma. Thirty key questions for biodiversity science in China [J]. Biodiv Sci, 2022, 30(10): 22609-. |
| [7] | Yi Zou. The calculation of β-diversity for different sample sizes [J]. Biodiv Sci, 2021, 29(6): 790-797. |
| [8] | Mingjia Li, Kaiyuan Wu, Fanfan Meng, Ji Shen, Yongqin Liu, Nengwen Xiao, Jianjun Wang. Beta diversity of stream bacteria in Hengduan Mountains: The effects of climatic and environmental variables [J]. Biodiv Sci, 2020, 28(12): 1570-1580. |
| [9] | Meixiang Gao, Lin Lin, Liang Chang, Xin Sun, Dong Liu, Donghui Wu. Spatial patterns and assembly rules in soil fauna communities: A review [J]. Biodiv Sci, 2018, 26(10): 1034-1050. |
| [10] | Yang Meng, Yue Qiu, Liang Zhang, Cuiling Wang, Zhenhua Zang, Xueyao Zhang, Guozhen Shen, Caifeng Yan, Quansheng Chen. Effects of geographical distance and differences in climate and altitude on species dissimilarity of vascular plant communities in the Dulongjiang River Watershed Area [J]. Biodiv Sci, 2017, 25(12): 1313-1320. |
| [11] | Jianming Wang, Wenjuan Wang, Jingwen Li, Yiming Feng, Bo Wu, Qi Lu. Biogeographic patterns and environmental interpretation of plant species richness in desert regions of Northwest China [J]. Biodiv Sci, 2017, 25(11): 1192-1201. |
| [12] | Nancai Pei, Jinlong Zhang, Xiangcheng Mi, Xuejun Ge. Plant DNA barcodes promote the development of phylogenetic commu- nity ecology [J]. Biodiv Sci, 2011, 19(3): 284-294. |
| [13] | Hongyu Niu, Zhengfeng Wang, Juyu Lian, Wanhui Ye, Hao Shen. New progress in community assembly: community phylogenetic structure combining evolution and ecology [J]. Biodiv Sci, 2011, 19(3): 275-283. |
| [14] | Huixian Wu, Jianliang Yao, Yan Liu, Junzeng Xue, Qinghua Cai, Jiankang Liu. Seasonal variation and longitudinal distribution characters of cladocerans in the Three Gorges Reservoir [J]. Biodiv Sci, 2009, 17(5): 512-517. |
| [15] | Jianliang Yao, Junzeng Xue, Dengyuan Wang, Qinghua Cai, Xiangfei Huang, Jiankang Liu . Seasonal variation and longitudinal distribution of copepods in the main river area of the Three Gorges Reservoir [J]. Biodiv Sci, 2007, 15(3): 300-305. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn