



Biodiv Sci ›› 2021, Vol. 29 ›› Issue (10): 1308-1320. DOI: 10.17520/biods.2021057 cstr: 32101.14.biods.2021057
Special Issue: 物种形成与系统进化
• Original Papers: Plant Diversity • Previous Articles Next Articles
Xingke Fan1,2, Xia Yan3, Yuanyuan Feng4, Jinhua Ran4, Chaoju Qian1, Xiaoyue Yin1,2, Shanshan Zhou1,2, Tingzhou Fang1,2, Xiaofei Ma1,3,*( )
)
Received:2021-02-09
Accepted:2021-06-10
Online:2021-10-20
Published:2021-10-20
Contact:
Xiaofei Ma
Xingke Fan, Xia Yan, Yuanyuan Feng, Jinhua Ran, Chaoju Qian, Xiaoyue Yin, Shanshan Zhou, Tingzhou Fang, Xiaofei Ma. Genome size variations and species differentiation of Reaumuria soongarica[J]. Biodiv Sci, 2021, 29(10): 1308-1320.
| 简称 Code | 地理位置 Location | 经纬度 Locality | 海拔 Altitude (m) | 年均降水量 Mean annual precipitation (mm) | 遗传支系 Lineage |
|---|---|---|---|---|---|
| HG | 甘肃红古 Honggu, Gansu | 103.03° E, 36.26° N | 1,775 | 369 | 东部支系 Eastern |
| SMY | 宁夏四马营 Simaying, Ningxia | 106.01° E, 38.50° N | 1,175 | 193 | 东部支系 Eastern |
| AKS | 新疆阿克苏 Akesu, Xinjiang | 79.70° E, 41.57° N | 1,491 | 164 | 西部支系 Western |
| CDY | 新疆策达雅 Cedaya, Xinjiang | 84.86° E, 42.01° N | 1,078 | 80 | 西部支系 Western |
| SW | 新疆沙湾 Shawan, Xinjiang | 85.33° E, 44.32° N | 519 | 150 | 北疆支系 Northern |
| FK | 新疆阜康 Fukang, Xinjiang | 88.13° E, 44.32° N | 491 | 170 | 北疆支系 Northern |
| HSS | 新疆火烧山 Huoshaoshan, Xinjiang | 88.99° E, 44.86° N | 478 | 155 | 北疆支系 Northern |
| WCC | 新疆五彩城 Wucaicheng, Xinjiang | 88.99° E, 45.16° N | 773 | 176 | 北疆支系 Northern |
Table 1 Sample information of Reaumuria soongarica populations
| 简称 Code | 地理位置 Location | 经纬度 Locality | 海拔 Altitude (m) | 年均降水量 Mean annual precipitation (mm) | 遗传支系 Lineage |
|---|---|---|---|---|---|
| HG | 甘肃红古 Honggu, Gansu | 103.03° E, 36.26° N | 1,775 | 369 | 东部支系 Eastern |
| SMY | 宁夏四马营 Simaying, Ningxia | 106.01° E, 38.50° N | 1,175 | 193 | 东部支系 Eastern |
| AKS | 新疆阿克苏 Akesu, Xinjiang | 79.70° E, 41.57° N | 1,491 | 164 | 西部支系 Western |
| CDY | 新疆策达雅 Cedaya, Xinjiang | 84.86° E, 42.01° N | 1,078 | 80 | 西部支系 Western |
| SW | 新疆沙湾 Shawan, Xinjiang | 85.33° E, 44.32° N | 519 | 150 | 北疆支系 Northern |
| FK | 新疆阜康 Fukang, Xinjiang | 88.13° E, 44.32° N | 491 | 170 | 北疆支系 Northern |
| HSS | 新疆火烧山 Huoshaoshan, Xinjiang | 88.99° E, 44.86° N | 478 | 155 | 北疆支系 Northern |
| WCC | 新疆五彩城 Wucaicheng, Xinjiang | 88.99° E, 45.16° N | 773 | 176 | 北疆支系 Northern |
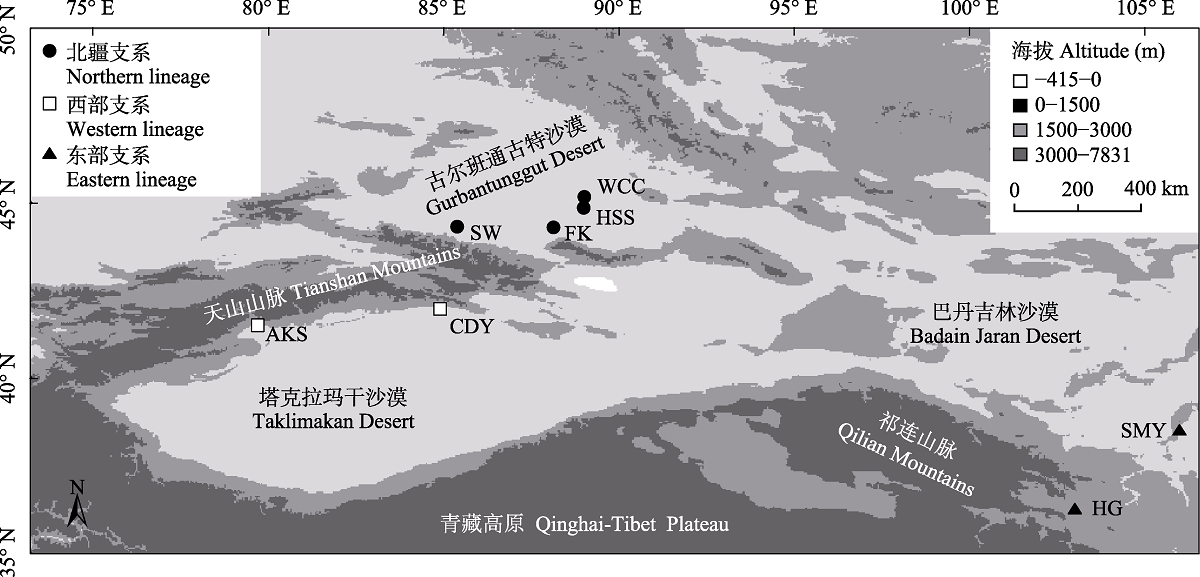
Fig. 1 Geographic locations of the eight Reaumuria soongarica populations measured in this study. The specific information of the populations is shown in Table 1.
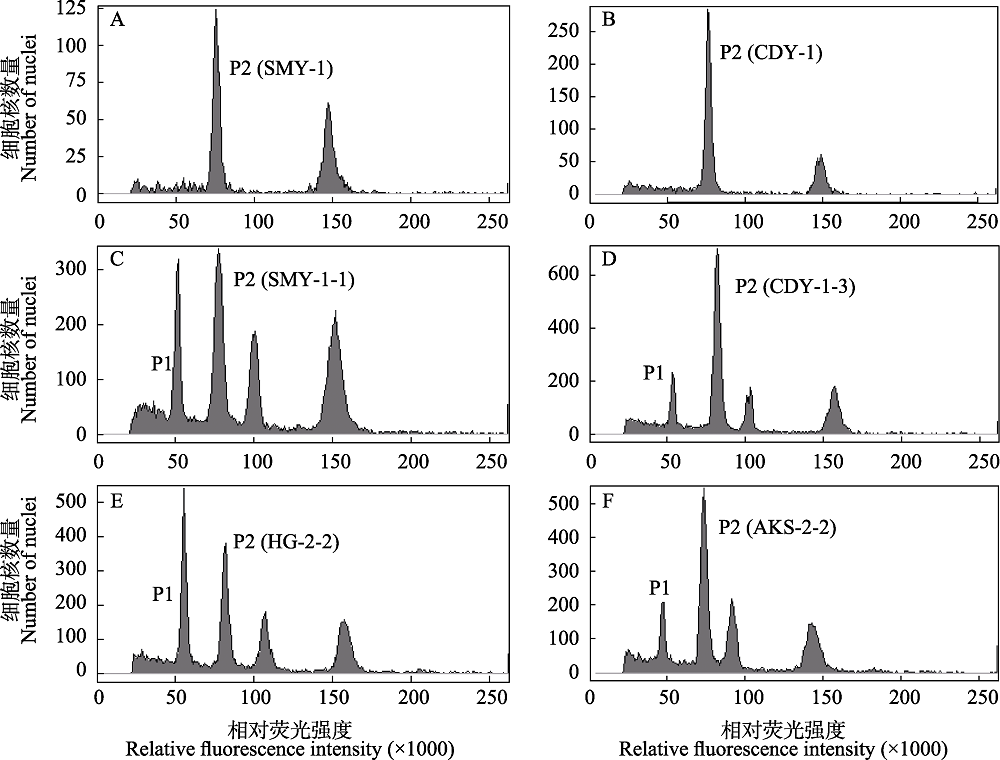
Fig. 2 Test results of different populations of the eastern and western lineages of Reaumuria soongarica using flow cytometry. A, B, The samples of SMY and CDY populations were measured without the internal standard; C-F, the mixed samples of Solanum pimpinellifolium and each Reaumuria soongarica population were tested, including SMY, CDY, HG, and AKS populations. P1 represents the nuclear DNA content of S. pimpinellifolium in G0/G1 phase, and P2 represents that of R. soongarica samples in G0/G1 phase. The population codes see Table 1.
| 支系 Lineage | 群体 Population | 红砂属样品G0/G1峰值 Peak values of Reaumuria (G0/G1) | 内标番茄G0/G1峰值 Peak values of Solanum pimpinellifolium (G0/G1) | DNA 1C-值 DNA 1C-values (pg) | 倍性 Ploidy | 一倍体基因组大小 Monoploid genome size (Mb) |
|---|---|---|---|---|---|---|
| 西部支系 Western lineage of R. soongarica | AKS | 74,192.667 ± 1,381.685 | 45,988.667 ± 1,264.577 | 1.219 ± 0.025 | 2n = 2x | 1,192.639 ± 24.701 |
| CDY | 76,255.167 ± 2,643.503 | 49,239.500 ± 1,725.806 | 1.170 ± 0.005 | 2n = 2x | 1,144.481 ± 4.816 | |
| 平均值 Average | - | - | 1.195 ± 0.031 | 2n = 2x | 1,168.560 ± 30.338 | |
| 东部支系 Eastern lineage of R. soongarica | HG | 75,210.667 ± 7,241.285 | 49,762.333 ± 5,100.729 | 1.143 ± 0.014 | 2n = 2x | 1,117.586 ± 13.338 |
| SMY | 74,570.500 ± 2,960.292 | 48,737.333 ± 1,845.872 | 1.156 ± 0.006 | 2n = 2x | 1,130.655 ± 5.787 | |
| 平均值 Average | - | - | 1.149 ± 0.012 | 2n = 2x | 1,124.120 ± 11.944 | |
| 北疆支系 Northern lineage of R. soongarica | SW | 77,156.167 ± 3,492.550 | 49,210.000 ± 1,972.505 | 1.185 ± 0.009 | 2n = 2x | 1,158.498 ± 9.009 |
| FK | 68,354.000 ± 4,150.933 | 44,074.167 ± 2,919.187 | 1.172 ± 0.013 | 2n = 2x | 1,146.490 ± 12.237 | |
| HSS | 141,698.000 ± 8,912.053 | 46,786.333 ± 2,805.389 | 2.288 ± 0.015 | 2n = 4x | 1,118.938 ± 7.437 | |
| WCC | 144,188.167 ± 5,140.113 | 47,052.500 ± 1,598.277 | 2.316 ± 0.023 | 2n = 4x | 1,132.291 ± 11.403 | |
| 五柱红砂 R. kaschgarica | DLH | 200,628.333 ± 1,704.432 | 38,574.000 ± 351.629 | 3.930 ± 0.006 | - | - |
| 红砂干燥叶片 Dehydrated leaves of R. soongarica | AKS | 72,426.000 ± 2,382.788 | 43,371.000 ± 1,417.229 | 1.262 ± 0.014 | 2n = 2x | 1,234.122 ± 14.060 |
Table 2 Measurement results of Reaumuria samples using flow cytometry (Mean ± SD)
| 支系 Lineage | 群体 Population | 红砂属样品G0/G1峰值 Peak values of Reaumuria (G0/G1) | 内标番茄G0/G1峰值 Peak values of Solanum pimpinellifolium (G0/G1) | DNA 1C-值 DNA 1C-values (pg) | 倍性 Ploidy | 一倍体基因组大小 Monoploid genome size (Mb) |
|---|---|---|---|---|---|---|
| 西部支系 Western lineage of R. soongarica | AKS | 74,192.667 ± 1,381.685 | 45,988.667 ± 1,264.577 | 1.219 ± 0.025 | 2n = 2x | 1,192.639 ± 24.701 |
| CDY | 76,255.167 ± 2,643.503 | 49,239.500 ± 1,725.806 | 1.170 ± 0.005 | 2n = 2x | 1,144.481 ± 4.816 | |
| 平均值 Average | - | - | 1.195 ± 0.031 | 2n = 2x | 1,168.560 ± 30.338 | |
| 东部支系 Eastern lineage of R. soongarica | HG | 75,210.667 ± 7,241.285 | 49,762.333 ± 5,100.729 | 1.143 ± 0.014 | 2n = 2x | 1,117.586 ± 13.338 |
| SMY | 74,570.500 ± 2,960.292 | 48,737.333 ± 1,845.872 | 1.156 ± 0.006 | 2n = 2x | 1,130.655 ± 5.787 | |
| 平均值 Average | - | - | 1.149 ± 0.012 | 2n = 2x | 1,124.120 ± 11.944 | |
| 北疆支系 Northern lineage of R. soongarica | SW | 77,156.167 ± 3,492.550 | 49,210.000 ± 1,972.505 | 1.185 ± 0.009 | 2n = 2x | 1,158.498 ± 9.009 |
| FK | 68,354.000 ± 4,150.933 | 44,074.167 ± 2,919.187 | 1.172 ± 0.013 | 2n = 2x | 1,146.490 ± 12.237 | |
| HSS | 141,698.000 ± 8,912.053 | 46,786.333 ± 2,805.389 | 2.288 ± 0.015 | 2n = 4x | 1,118.938 ± 7.437 | |
| WCC | 144,188.167 ± 5,140.113 | 47,052.500 ± 1,598.277 | 2.316 ± 0.023 | 2n = 4x | 1,132.291 ± 11.403 | |
| 五柱红砂 R. kaschgarica | DLH | 200,628.333 ± 1,704.432 | 38,574.000 ± 351.629 | 3.930 ± 0.006 | - | - |
| 红砂干燥叶片 Dehydrated leaves of R. soongarica | AKS | 72,426.000 ± 2,382.788 | 43,371.000 ± 1,417.229 | 1.262 ± 0.014 | 2n = 2x | 1,234.122 ± 14.060 |
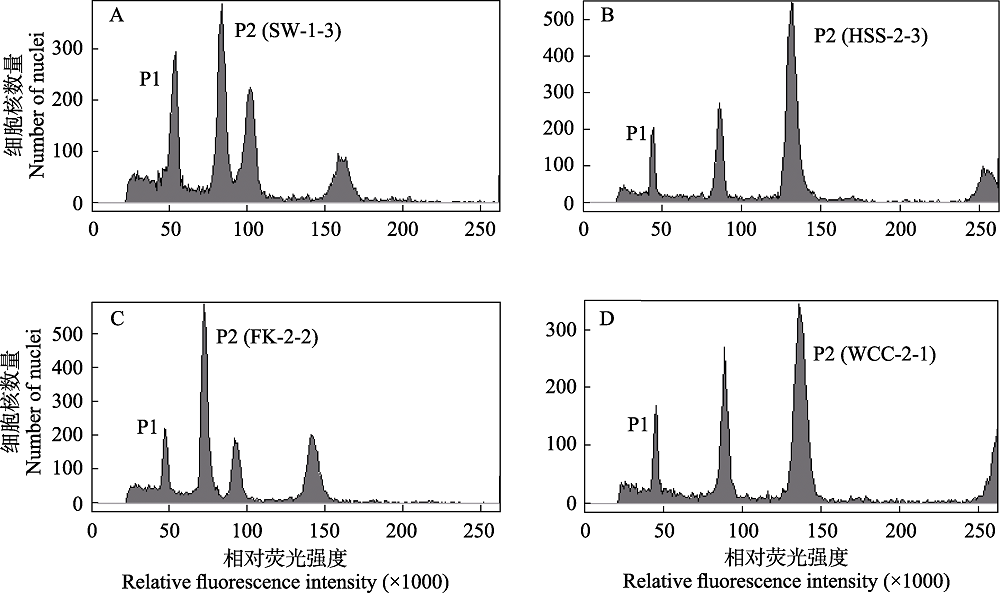
Fig. 3 Test results of the mixed sample of Solanum pimpinellifolium and different populations of the northern lineage of Reaumuria soongarica using flow cytometry. The populations of the northern lineage include SW (A), HSS (B), FK (C), and WCC (D). P1 represents the nuclear DNA content of S. pimpinellifolium in G0/G1 phase, and P2 represents that of R. soongarica samples in G0/G1 phase.
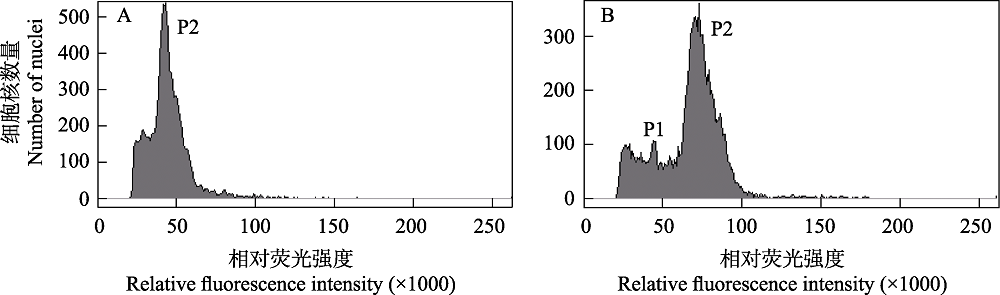
Fig. 4 Histograms of the nuclear DNA content of Reaumuria soongarica dry leaves (the AKS population). (A) R. soongarica sample alone; (B) the mixed sample of R. soongarica (P2, G0/G1) and Solanum pimpinellifolium (P1, G0/G1).
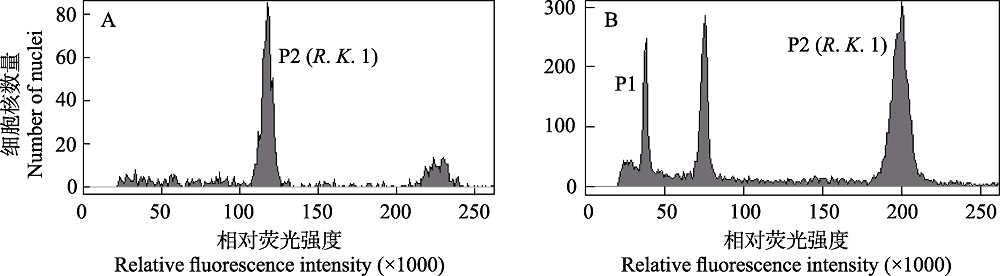
Fig. 5 Histograms of the nuclear DNA content of Reaumuria kaschgarica. (A) The pure sample of R. kaschgarica; (B) the mixed sample of R. kaschgarica (P2, G0/G1) and Solanum pimpinellifolium (P1, G0/G1).
| [1] |
Abbott R, Albach D, Ansell S, Arntzen JW, Baird SJE, Bierne N, Boughman J, Brelsford A, Buerkle CA, Buggs R, Butlin RK, Dieckmann U, Eroukhmanoff F, Grill A, Cahan SH, Hermansen JS, Hewitt G, Hudson AG, Jiggins C, Jones J, Keller B, Marczewski T, Mallet J, Martinez-Rodriguez P, Möst M, Mullen S, Nichols R, Nolte AW, Parisod C, Pfennig K, Rice AM, Ritchie MG, Seifert B, Smadja CM, Stelkens R, Szymura JM, Väinölä R, Wolf JBW, Zinner D (2013) Hybridization and speciation. Journal of Evolutionary Biology, 26, 229-246.
DOI PMID |
| [2] |
Barton NH (2001) The role of hybridization in evolution. Molecular Ecology, 10, 551-568.
PMID |
| [3] |
Buerkle CA, Morris RJ, Asmussen MA, Rieseberg LH (2000) The likelihood of homoploid hybrid speciation. Heredity, 84, 441-451.
DOI URL |
| [4] |
Čertnerová D, Škaloud P (2020) Substantial intraspecific genome size variation in golden-brown algae and its phenotypic consequences. Annals of Botany, 126, 1077-1087.
DOI URL |
| [5] | Cui DF, Liao WB, Zhang B (2000) Determination of flavonoid compounds of Reaumuria L. (Tamaricaceae) and their taxonomical significance. Acta Botanica Boreali-Occidentalia Sinica, 20, 283-287. (in Chinese with English abstract) |
| [崔大方, 廖文波, 张冰 (2000) 琵琶柴属植物黄酮类化合物的测定及分类学意义. 西北植物学报, 20, 283-287.] | |
| [6] | Cui DF, Zhong MJ (1999) A new species of Reaumuria L. from Xinjiang. Acta Botanica Boreali-Occidentalia Sinica, 19, 552-554. (in Chinese) |
| [崔大方, 仲铭锦 (1999) 新疆琵琶柴属一新种. 西北植物学报, 19, 552-554.] | |
| [7] |
Doležel J, Bartoš J (2005) Plant DNA flow cytometry and estimation of nuclear genome size. Annals of Botany, 95, 99-110.
DOI URL |
| [8] | Doležel J, Bartoš J, Voglmayr H, Greilhuber J (2003) Nuclear DNA content and genome size of trout and human. Cytometry Part A, 51A, 127-128. |
| [9] |
Doležel J, Greilhuber J, Suda J (2007) Estimation of nuclear DNA content in plants using flow cytometry. Nature Protocols, 2, 2233-2244.
DOI URL |
| [10] |
Fan XK, Yan X, Qian CJ, Bachir DG, Yin XY, Sun PP, Ma XF (2020) Leaf size variations in a dominant desert shrub, Reaumuria soongarica, adapted to heterogeneous environments. Ecology and Evolution, 10, 10076-10094.
DOI URL |
| [11] |
Garcia S, Garnatje T, Hidalgo O de Xaxars GM, Pellicer J, Sánchez-Jiménez I, Vitales D, Vallès J (2010) First genome size estimations for some eudicot families and genera. Collectanea Botanica, 29, 7-16.
DOI URL |
| [12] | Gaskin JF, Ghahremani-nejad F, Zhang DY, Londo JP (2004) A systematic overview of Frankeniaceae and Tamaricaceae from nuclear rDNA and plastid sequence data. Annals of the Missouri Botanical Garden, 91, 401-409. |
| [13] |
Greilhuber J, Doležel J, Lysák MA, Bennett MD (2005) The origin, evolution and proposed stabilization of the terms ‘genome size' and ‘C-value' to describe nuclear DNA contents. Annals of Botany, 95, 255-260.
PMID |
| [14] |
Gross BL, Rieseberg LH (2005) The ecological genetics of homoploid hybrid speciation. Journal of Heredity, 96, 241-252.
PMID |
| [15] | Guignard MS, Crawley MJ, Kovalenko D, Nichols RA, Trimmer M, Leitch AR, Leitch IJ (2019) Interactions between plant genome size, nutrients and herbivory by rabbits, molluscs and insects on a temperate grassland. Proceedings of the Royal Society B: Biological Sciences, 286, 20182619. |
| [16] |
Guignard MS, Nichols RA, Knell RJ, Macdonald A, Romila CA, Trimmer M, Leitch IJ, Leitch AR (2016) Genome size and ploidy influence angiosperm species' biomass under nitrogen and phosphorus limitation. New Phytologist, 210, 1195-1206.
DOI PMID |
| [17] | Guo SL, Zhou P, Yin LP, Lou YX (2011) Interspecific and intraspecific variations of plant DNA C-values and its biological significance. Journal of Shanghai Normal University (Natural Sciences), 40, 102-110. (in Chinese with English abstract) |
| [郭水良, 周平, 印丽萍, 娄玉霞 (2011) 植物DNA C-值在种间和种内的变异及其生物学意义. 上海师范大学学报(自然科学版), 40, 102-110.] | |
| [18] | Hao XL, Zhang ML, Wang SM, Zhang X (2014) Classification and distribution of the genus Reaumuria L. in Tamaricaceae. Arid Zone Research, 31, 838-843. (in Chinese with English abstract) |
| [郝晓莉, 张明理, 王绍明, 张霞 (2014) 柽柳科红砂属(Reaumuria L.)的分类与分布. 干旱区研究, 31, 838-843.] | |
| [19] |
Hidalgo O, Vallès J, Romo A, Canela MÁ, Garnatje T (2015) Genome size variation in gymnosperms under different growth conditions. Caryologia, 68, 92-96.
DOI URL |
| [20] | Hou YJ (2015) Study on the Transport Processes and Ecological Effect of Heavy Metals in Dust-soil-plant in the East Junggar Region. PhD dissertation, Xinjiang University, Urumqi. (in Chinese with English abstract) |
| [侯艳军 (2015) 准东地区降尘-土壤-植物重金属迁移过程及生态效应研究. 博士学位论文, 新疆大学, 乌鲁木齐.] | |
| [21] | Husband BC (2000) Constraints on polyploid evolution:A test of the minority cytotype exclusion principle. Proceedings of the Royal Society B: Biological Sciences, 267, 217-223. |
| [22] |
Li ZH, Chen J, Zhao GF, Guo YP, Kou YX, Ma YZ, Wang G, Ma XF (2012) Response of a desert shrub to past geological and climatic change: A phylogeographic study of Reaumuria soongarica (Tamaricaceae) in western China. Journal of Systematics and Evolution, 50, 351-361.
DOI URL |
| [23] | Lin F, Xiao YE, Zhou XY, Tang Y, Gao BH (2018) Estimation of genomic C value of 25 samples of Iris plants by flow cytometry. Acta Agrestia Sinica, 26, 985-990. (in Chinese with English abstract) |
| [林峰, 肖月娥, 周翔宇, 唐颖, 高步红 (2018) 25份鸢尾属植物基因组DNA C值的流式测定. 草地学报, 26, 985-990.] | |
| [24] |
Liu JQ (2016) “The integrative species concept” and “species on the speciation way”. Biodiversity Science, 24, 1004-1008. (in Chinese with English abstract)
DOI URL |
| [刘建全 (2016) “整合物种概念”和“分化路上的物种”. 生物多样性, 24, 1004-1008.] | |
| [25] | Liu JQ, Qiu MX, Pu JC, Lu ZM (1982) The typical extreme xerophyte Reaumuria soongorica in the desert of China. Acta Botanica Sinica, 24, 485-488. (in Chinese) |
| [刘家琼, 邱明新, 蒲锦春, 鲁作民 (1982) 我国荒漠典型超旱生植物-红砂. 植物学报, 24, 485-488.] | |
| [26] | Liu YX (1995) A study on origin and formation of the Chinese desert floras. Acta Phytotaxonomica Sinica, 33, 131-143. (in Chinese with English abstract) |
| [刘媖心 (1995) 试论我国沙漠地区植物区系的发生与形成. 植物分类学报, 33, 131-143.] | |
| [27] |
Loureiro J, Rodriguez E, Doležel J, Santos C (2007) Two new nuclear isolation buffers for plant DNA flow cytometry: A test with 37 species. Annals of Botany, 100, 875-888.
PMID |
| [28] | Loureiro J, Trávníček P, Rauchová J, Urfus T, Vít P, Štech M, Castro S, Suda J (2010) The use of flow cytometry in the biosystematics, ecology and population biology of homoploid plants. Preslia, 82, 3-21. |
| [29] |
Ma JY, Chen T, Qiang WY, Wang G (2005) Correlations between foliar stable carbon isotope composition and environmental factors in desert plant Reaumuria soongorica (Pall.) Maxim. Journal of Integrative Plant Biology, 47, 1065-1073.
DOI URL |
| [30] |
Ma XF, Szmidt AE, Wang XR (2006) Genetic structure and evolutionary history of a diploid hybrid pine Pinus densata inferred from the nucleotide variation at seven gene loci. Molecular Biology and Evolution, 23, 807-816.
DOI URL |
| [31] |
Mahelka V, Suda J, Jarolímová V, Trávníček P, Krahulec F (2005) Genome size discriminates between closely related taxa Elytrigia repens and E. intermedia (Poaceae: Triticeae) and their hybrid. Folia Geobotanica, 40, 367-384.
DOI URL |
| [32] |
Mallet J (2005) Hybridization as an invasion of the genome. Trends in Ecology & Evolution, 20, 229-237.
DOI URL |
| [33] |
Mallet J (2007) Hybrid speciation. Nature, 446, 279-283.
DOI URL |
| [34] |
Morigengaowa, Shang H, Liu BD, Kang M, Yan YH (2019) One or more species? GBS sequencing and morphological traits evidence reveal species diversification of Sphaeropteris brunoniana in China. Biodiversity Science, 27, 1196-1204. (in Chinese with English abstract)
DOI |
|
[莫日根高娃, 商辉, 刘保东, 康明, 严岳鸿 (2019) 一个种还是多个种? 简化基因组及其形态学证据揭示中国白桫椤植物的物种多样性分化. 生物多样性, 27, 1196-1204.]
DOI |
|
| [35] |
Pellicer J, Hidalgo O, Dodsworth S, Leitch IJ (2018) Genome size diversity and its impact on the evolution of land plants. Genes, 9, 88.
DOI URL |
| [36] | Pellicer J, Leitch IJ (2020) The Plant DNA C-values database (release 7.1): An updated online repository of plant genome size data for comparative studies. New Phytologist, 226, 301-305. |
| [37] |
Qian ZQ, Xu L, Wang YL, Yang J, Zhao GF (2008) Ecological genetics of Reaumuria soongorica (Pall.) Maxim. population in the oasis-desert ecotone in Fukang, Xinjiang, and its implications for molecular evolution. Biochemical Systematics and Ecology, 36, 593-601.
DOI URL |
| [38] |
Qiu F, Baack EJ, Whitney KD, Bock DG, Tetreault HM, Rieseberg LH, Ungerer MC (2019) Phylogenetic trends and environmental correlates of nuclear genome size variation in Helianthus sunflowers. New Phytologist, 221, 1609-1618.
DOI URL |
| [39] |
Rieseberg LH (1991) Homoploid reticulate evolution in Helianthus (Asteraceae): Evidence from ribosomal genes. American Journal of Botany, 78, 1218-1237.
DOI URL |
| [40] |
Rieseberg LH, Raymond O, Rosenthal DM, Lai Z, Livingstone K, Nakazato T, Durphy JL, Schwarzbach AE, Donovan LA, Lexer C (2003) Major ecological transitions in wild sunflowers facilitated by hybridization. Science, 301, 1211-1216.
PMID |
| [41] |
Samadi N, Ghaffari SM, Akhani H (2013) Meiotic behaviour, karyotype analyses and pollen viability in species of Tamarix (Tamaricaceae). Willdenowia, 43, 195-203.
DOI URL |
| [42] |
Schwarzbach AE, Rieseberg LH (2002) Likely multiple origins of a diploid hybrid sunflower species. Molecular Ecology, 11, 1703-1715.
PMID |
| [43] | Shen J, Xu J, Liu GX, Shi JS (2015) Effects of different isolation buffers on DNA resolution in nucleus suspension of root tip cell of Cunninghamia lanceolata (Lamb.) Hook. Molecular Plant Breeding, 13, 190-196. (in Chinese with English abstract) |
| [沈捷, 徐进, 刘光欣, 施季森 (2015) 不同分离缓冲液对杉木根尖细胞核悬液DNA分辨率的影响. 分子植物育种, 13, 190-196.] | |
| [44] | Shi Y, Liu Y, Yin HX, Yan X, Ma XF (2016) Seed germination characteristics and local adaptation of Reaumuria soongarica. Journal of Desert Research, 36, 644-650. (in Chinese with English abstract) |
| [石勇, 刘源, 殷恒霞, 燕霞, 马小飞 (2016) 红砂(Reaumuria soongarica)种子萌发特性及其局部适应性. 中国沙漠, 36, 644-650.] | |
| [45] |
Shi Y, Yan X, Yin HX, Qian CJ, Fan XK, Yin XY, Chang YX, Zhang CJ, Ma XF (2020) Divergence and hybridization in the desert plant Reaumuria soongarica. Journal of Systematics and Evolution, 58, 159-173.
DOI URL |
| [46] |
Shi Y, Yan X, Zhao PS, Yin HX, Zhao X, Xiao HL, Li XR, Chen GX, Ma XF (2013) Transcriptomic analysis of a tertiary relict plant, extreme xerophyte Reaumuria soongorica to identify genes related to drought adaptation. PLoS ONE, 8, e63993.
DOI URL |
| [47] | Smarda P, Bures P (2010) Understanding intraspecific variation in genome size in plants. Preslia, 82, 41-61. |
| [48] |
Soltis DE, Soltis PS, Tate JA (2004) Advances in the study of polyploidy since plant speciation. New Phytologist, 161, 173-191.
DOI URL |
| [49] |
Suda J, Trávníček P (2006) Reliable DNA ploidy determination in dehydrated tissues of vascular plants by DAPI flow cytometry-New prospects for plant research. Cytometry Part A, 69A, 273-280.
DOI URL |
| [50] |
Sun YS, Lu ZQ, Zhu XF, Ma H (2020) Genomic basis of homoploid hybrid speciation within chestnut trees. Nature Communications, 11, 3375.
DOI URL |
| [51] | Taylor SA, Larson EL (2019) Insights from genomes into the evolutionary importance and prevalence of hybridization in nature. Nature Ecology & Evolution, 3, 170-177. |
| [52] |
The Tomato Genome Consortium (2012) The tomato genome sequence provides insights into fleshy fruit evolution. Nature, 485, 635-641.
DOI URL |
| [53] |
Urfusová R, Mahelka V, Krahulec F, Urfus T (2021) Evidence of widespread hybridization among couch grasses (Elymus, Poaceae). Journal of Systematics and Evolution, 59, 113-124.
DOI URL |
| [54] |
Van de Peer Y, Ashman TL, Soltis PS, Soltis DE (2021) Polyploidy: An evolutionary and ecological force in stressful times. The Plant Cell, 33, 11-26.
DOI URL |
| [55] |
Wan JZ, Chen LX, Gao S, Song YB, Tang SL, Yu FH, Li JM, Dong M (2019) Ecological niche shift between diploid and tetraploid plants of Fragaria (Rosaceae) in China. South African Journal of Botany, 121, 68-75.
DOI |
| [56] |
Wang XH, Zhang T, Wen ZN, Xiao HL, Yang ZJ, Chen GX, Zhao X (2011) The chromosome number, karyotype and genome size of the desert plant diploid Reaumuria soongorica (Pall.) Maxim. Plant Cell Reports, 30, 955-964.
DOI URL |
| [57] | Wang Y, Xiao Y, Liu W, Li TT, Hu R, Qiao ZX (2015) Operation skills of flow cytometer for detecting nuclear DNA contents in higher plant cells. Plant Science Journal, 33, 126-131. (in Chinese with English abstract) |
| [汪艳, 肖媛, 刘伟, 李婷婷, 胡锐, 乔志仙 (2015) 流式细胞仪检测高等植物细胞核DNA含量的方法. 植物科学学报, 33, 126-131.] | |
| [58] |
Wang YG (2017) Natural hybridization and speciation. Biodiversity Science, 25, 565-576. (in Chinese with English abstract)
DOI URL |
|
[王玉国 (2017) 自然杂交与物种形成. 生物多样性, 25, 565-576.]
DOI |
|
| [59] |
Wang ZF, Jiang YZ, Bi H, Lu ZQ, Ma YZ, Yang XY, Chen NN, Tian B, Liu BB, Mao XX, Ma T, DiFazio SP, Hu QJ, Abbott RJ, Liu JQ (2021) Hybrid speciation via inheritance of alternate alleles of parental isolating genes. Molecular Plant, 14, 208-222.
DOI URL |
| [60] |
Yakimowski SB, Rieseberg LH (2014) The role of homoploid hybridization in evolution: A century of studies synthesizing genetics and ecology. American Journal of Botany, 101, 1247-1258.
DOI PMID |
| [61] | Yang HW, Chen WL (2012) The effects of different growth regulators on the Reaumuria soongorica (Pall.) Maxim seedling emergence and growth. Journal of Fujian Forestry Science and Technology, 39, 71-73. (in Chinese with English abstract) |
| [杨海文, 陈文莲 (2012) 植物生长调节剂对红砂出苗及苗期生长的影响. 福建林业科技, 39, 71-73.] | |
| [62] |
Yang JY, Cushman SA, Song XM, Yang J, Zhang PJ (2015) Genetic diversity and drivers of genetic differentiation of Reaumuria soongorica of the Inner Mongolia Plateau in China. Plant Ecology, 216, 925-937.
DOI URL |
| [63] |
Yin HX, Yan X, Shi Y, Qian CJ, Li ZH, Zhang W, Wang LR, Li Y, Li XZ, Chen GX, Li XR, Nevo E, Ma XF (2015) The role of East Asian monsoon system in shaping population divergence and dynamics of a constructive desert shrub Reaumuria soongarica. Scientific Reports, 5, 15823.
DOI URL |
| [64] | Yu HM, Wang J, Zhao MZ, Meng XF (2012) Optimization of strawberry ploidy identification method using flow cytometry. Journal of Southern Agriculture, 43, 1530-1533. (in Chinese with English abstract) |
| [于红梅, 王静, 赵密珍, 孟宪凤 (2012) 利用流式细胞仪检测草莓倍性方法的优化. 南方农业学报, 43, 1530-1533.] | |
| [65] | Zhang DY, Pan BR, Yin LK (2003) The photogeographical studies of Tamarix (Tamaricaceae). Acta Botanica Yunnanica, 25, 415-427. (in Chinese with English abstract) |
| [张道远, 潘伯荣, 尹林克 (2003) 柽柳科柽柳属的植物地理研究. 云南植物研究, 25, 415-427.] | |
| [66] | Zhang LY, Chen CD (2002) On the general characteristics of plant diversity of Gurbantunggut sandy desert. Acta Ecologica Sinica, 22, 1923-1932. (in Chinese with English abstract) |
| [张立运, 陈昌笃 (2002) 论古尔班通古特沙漠植物多样性的一般特点. 生态学报, 22, 1923-1932.] | |
| [67] |
Zhang ML, Hao XL, Sanderson SC, Vyacheslav BV, Sukhorukov AP, Zhang X (2014a) Spatiotemporal evolution of Reaumuria (Tamaricaceae) in Central Asia: Insights from molecular biogeography. Phytotaxa, 167, 89-103.
DOI URL |
| [68] |
Zhang ML, Meng HH, Zhang HX, Vyacheslav BV, Sanderson SC (2014b) Himalayan origin and evolution of Myricaria (Tamaricaeae) in the Neogene. PLoS ONE, 9, e97582.
DOI URL |
| [69] | Zhang YM, Pan BR, Yin LK (1998) Seed morphology of Tamaricaceae in China arid areas and its systematic evolution. Journal of Plant Resources and Environment, 7, 22-27. (in Chinese with English abstract) |
| [张元明, 潘伯荣, 尹林克 (1998) 中国干旱区柽柳科植物种子形态特征及其系统学意义. 植物资源与环境, 7, 22-27.] | |
| [70] |
Zhao W, Meng JX, Wang BS, Zhang LS, Xu YL, Zeng QY, Li Y, Mao JF, Wang XR (2014) Weak crossability barrier but strong juvenile selection supports ecological speciation of the hybrid pine Pinus densata on the Tibetan Plateau. Evolution, 68, 3120-3133.
DOI PMID |
| [71] |
Zonneveld BJM, Duncan GD (2010) Genome sizes of Eucomis L'Hér. (Hyacinthaceae) and a description of the new species Eucomis grimshawii G. D. Duncan & Zonneveld. Plant Systematics and Evolution, 284, 99-109.
DOI URL |
| [72] |
Zonneveld BJM, Leitch IJ, Bennett MD (2005) First nuclear DNA amounts in more than 300 angiosperms. Annals of Botany, 96, 229-244.
PMID |
| [73] | Zou X, Sun LY, Wan XX, Chen Y, Lin F, Yin ZF (2020) Establishment of flow cytometry system for DNA C-value determination of polyploid species in subgenus Yulania. Plant Science Journal, 38, 369-377. (in Chinese with English abstract) |
| [邹璇, 孙李勇, 万小霞, 陈瑶, 林峰, 尹增芳 (2020) 玉兰亚属多倍体植物DNA C-值流式细胞术测定体系的建立. 植物科学学报, 38, 369-377.] |
| [1] | Xiao-Qing Wu Meihui Zhang Suting Ge Manshu Li Kun Song Guochun Shen Jian Zhang. Spatiotemporal Dynamics of Woody Plant Species Diversity and Aboveground Biomass during Near-Natural Forest Reconstruction in Shanghai: A Case Study from the Eco-Island in Minhang District [J]. Biodiv Sci, 2025, 33(5): 24444-. |
| [2] | Tai Wang, Fujun Song, Yongsheng Zhang, Zhongyu Lou, Yanping Zhang, Yanyan Du. Fish diversity and resource status in interior drainage systems of Hexi Corridor [J]. Biodiv Sci, 2025, 33(4): 24387-. |
| [3] | Jingjing Zhang, Wenbin Huang, Yiting Chen, Zepeng Yang, Weiye Ke, Zhaojie Peng, Shichao Wei, Zhiwei Zhang, Yisi Hu, Wenhua Yu, Wenliang Zhou. Reef-building coral diversity and distribution characteristics in the National Nature Reserve for Marine Ecology of Guangdong Nanpeng Islands [J]. Biodiv Sci, 2025, 33(4): 24424-. |
| [4] | Shang Huadan, Zhang Chuqing, Wang Mei, Pei Wenya, Li Guohong, Wang Hongbin. Species diversity and geographic distribution of poplar pests in China [J]. Biodiv Sci, 2025, 33(2): 24370-. |
| [5] | Wu Yuxuan, Wang Ping, Hu Xiaosheng, Ding Yi, Peng Tiantian, Zhi Qiuying, Bademu Qiqige, Li Wenjie, Guan Xiao, Li Junsheng. Evaluation of grassland degradation status and vegetation characteristics changes in Hulunbuir [J]. Biodiv Sci, 2025, 33(2): 24118-. |
| [6] | Zihong Chen, Yifei Zhang, Kai Chen, Jianying Chen, Ling Xu. Species diversity of entomopathogenic fungi and the influencing factors in the Southern Gaoligong Mountains [J]. Biodiv Sci, 2025, 33(1): 24228-. |
| [7] | Ke Tan, Yao Ning, Renfen Wang, Qing Wang, Danping Liang, Zibing Xin, Fang Wen. A dataset on the checklist and geographical distribution of Gesneriaceae in China [J]. Biodiv Sci, 2025, 33(1): 23275-. |
| [8] | Jianan Han, Yang Su, Fei Li, Junyan Liu, Yilin Zhao, Lin Li, Jiancheng Zhao, Hongzhu Liang, Min Li. Bryophytes diversity of Hebei Province, China [J]. Biodiv Sci, 2024, 32(9): 24096-. |
| [9] | Hongyu Niu, Lu Chen, Hengyue Zhao, Gulzar Abdukirim, Hongmao Zhang. Effects of urbanization on animals: From community to individual level [J]. Biodiv Sci, 2024, 32(8): 23489-. |
| [10] | Xue Bai, Zhengfei Li, Yang Liu, Junqian Zhang, Duopeng Zhang, Xin Luo, Jiali Yang, Lina Du, Xuankong Jiang, Ruiwen Wu, Zhicai Xie. Species diversity and maintenance mechanisms of benthic macroinvertebrate assemblages in the Xijiang River [J]. Biodiv Sci, 2024, 32(7): 23499-. |
| [11] | Jia Xu, Xiaojuan Cui, Yifei Zhang, Chang Wu, Yuandong Sun. Fish diversity and distribution in the Nanling region [J]. Biodiv Sci, 2024, 32(7): 23482-. |
| [12] | Qiyu Kuang, Liang Hu. Species diversity and geographical distribution of marine benthic shell-bearing mollusks around Donghai Island and Naozhou Island, Guangdong Province [J]. Biodiv Sci, 2024, 32(5): 24065-. |
| [13] | Yongqiang Zhao, Xiyu Yan, Jiaqi Xie, Mengting Hou, Danmei Chen, Lipeng Zang, Qingfu Liu, Mingzhen Sui, Guangqi Zhang. Species diversity and community assembly of woody plants at different life history stages during the natural restoration of degraded karst forests [J]. Biodiv Sci, 2024, 32(5): 23462-. |
| [14] | Weiqiang Xu, Qiang Su. Exploring the interplay of fractal model and species abundance distribution: A case study of shellfish and insect [J]. Biodiv Sci, 2024, 32(4): 23410-. |
| [15] | Hui Ran, Tianyou Yang, Xiaoqi Mi. The updated checklist of reptiles in Guizhou Province, China [J]. Biodiv Sci, 2024, 32(4): 23348-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn