



生物多样性 ›› 2026, Vol. 34 ›› Issue (2): 25256. DOI: 10.17520/biods.2025256 cstr: 32101.14.biods.2025256
纪林1,2,3, 邓宸迅1,2,3, 王丽凤1,2,3( ), 王德港1,2,3, 王建涛4(
), 王德港1,2,3, 王建涛4( ), 于永永4, 张军国1,2,3,*(
), 于永永4, 张军国1,2,3,*( )
)
收稿日期:2025-07-02
接受日期:2026-01-14
出版日期:2026-02-20
发布日期:2026-03-23
通讯作者:
张军国
基金资助:
Lin Ji1,2,3, Chenxun Deng1,2,3, Lifeng Wang1,2,3( ), Degang Wang1,2,3, Jiantao Wang4(
), Degang Wang1,2,3, Jiantao Wang4( ), Yongyong Yu4, Junguo Zhang1,2,3,*(
), Yongyong Yu4, Junguo Zhang1,2,3,*( )
)
Received:2025-07-02
Accepted:2026-01-14
Online:2026-02-20
Published:2026-03-23
Contact:
Junguo Zhang
Supported by:摘要:
随着人工智能技术的快速发展, 利用深度学习方法对野生动物图像进行自动识别, 已成为野生动物调查保护的关键手段。实际采集的野生动物图像数据通常呈现一种偏态分布特征, 即少数高频类别物种样本充足, 而大多数低频类别物种样本稀缺, 影响模型的整体识别性能。针对这一问题, 本文提出一种基于Diff-SCC模型的偏态分布野生动物识别方法。首先, 该方法利用大语言模型生成类别的丰富语义描述, 引导扩散模型生成额外样本, 同时引入多尺度负样本筛选策略, 从像素空间、特征空间及语义空间3个维度进行图像质量评估和筛选, 提升低频类别的特征多样性并平衡数据分布。其次, 本文在主干网络ResNet50中引入SCConv模块以减少空间与通道建模过程中的冗余特征, 并增强模型对前景区域的感知能力, 从而提高模型对低频类别的识别性能。最后, 本文在自建数据集ULB-12和公开野生动物数据集NACTI上开展对比实验以验证模型的性能。实验结果显示, Diff-SCC模型在上述两个数据集上的整体识别准确率分别达到78.71%和80.84%, 低频类别的识别准确率相较基线模型分别提升9.96%和9.99%。上述结果验证了Diff-SCC在处理偏态分布数据集上的有效性, 能够为野生动物智能监测与保护提供可靠的技术支撑。
纪林, 邓宸迅, 王丽凤, 王德港, 王建涛, 于永永, 张军国 (2026) 基于Diff-SCC模型的偏态分布野生动物识别方法. 生物多样性, 34, 25256. DOI: 10.17520/biods.2025256.
Lin Ji, Chenxun Deng, Lifeng Wang, Degang Wang, Jiantao Wang, Yongyong Yu, Junguo Zhang (2026) A wildlife recognition method for skewed distributions based on the Diff-SCC model. Biodiversity Science, 34, 25256. DOI: 10.17520/biods.2025256.
| 物种 Species | 数量 Number | 物种 Species | 数量 Number |
|---|---|---|---|
| 美洲黑熊 Ursus americanus | 2,765 | 条纹臭鼬 Mephitis mephitis | 1,123 |
| 美洲狮 Puma concolor | 2,707 | 驼鹿 Alces alces | 994 |
| 短尾猫 Lynx rufus | 2,310 | 灰松鼠 Sciurus carolinensis | 798 |
| 骡鹿 Odocoileus hemionus | 1,958 | 火鸡 Meleagris gallopavo | 615 |
| 马鹿 Cervus canadensis | 1,956 | 黑尾长耳大野兔 Lepus californicus | 592 |
| 欧洲马鹿 Cervus elaphus | 1,928 | 九带犰狳 Dasypus novemcinctus | 434 |
| 野猪 Sus scrofa | 1,693 | 北美红松鼠 Tamiasciurus hudsonicus | 188 |
| 郊狼 Canis latrans | 1,614 | 加州翎鹑 Callipepla californica | 137 |
| 灰狐 Urocyon cinereoargenteus | 1,339 | 赤狐 Vulpes vulpes | 127 |
| 白靴兔 Lepus americanus | 1,246 | 弗吉尼亚负鼠 Didelphis virginiana | 110 |
| 浣熊 Procyon lotor | 1,176 | 美洲貂 Martes americana | 88 |
表1 NACTI野生动物数据集训练集构成
Table 1 Composition of the training set of the NACTI wildlife dataset
| 物种 Species | 数量 Number | 物种 Species | 数量 Number |
|---|---|---|---|
| 美洲黑熊 Ursus americanus | 2,765 | 条纹臭鼬 Mephitis mephitis | 1,123 |
| 美洲狮 Puma concolor | 2,707 | 驼鹿 Alces alces | 994 |
| 短尾猫 Lynx rufus | 2,310 | 灰松鼠 Sciurus carolinensis | 798 |
| 骡鹿 Odocoileus hemionus | 1,958 | 火鸡 Meleagris gallopavo | 615 |
| 马鹿 Cervus canadensis | 1,956 | 黑尾长耳大野兔 Lepus californicus | 592 |
| 欧洲马鹿 Cervus elaphus | 1,928 | 九带犰狳 Dasypus novemcinctus | 434 |
| 野猪 Sus scrofa | 1,693 | 北美红松鼠 Tamiasciurus hudsonicus | 188 |
| 郊狼 Canis latrans | 1,614 | 加州翎鹑 Callipepla californica | 137 |
| 灰狐 Urocyon cinereoargenteus | 1,339 | 赤狐 Vulpes vulpes | 127 |
| 白靴兔 Lepus americanus | 1,246 | 弗吉尼亚负鼠 Didelphis virginiana | 110 |
| 浣熊 Procyon lotor | 1,176 | 美洲貂 Martes americana | 88 |
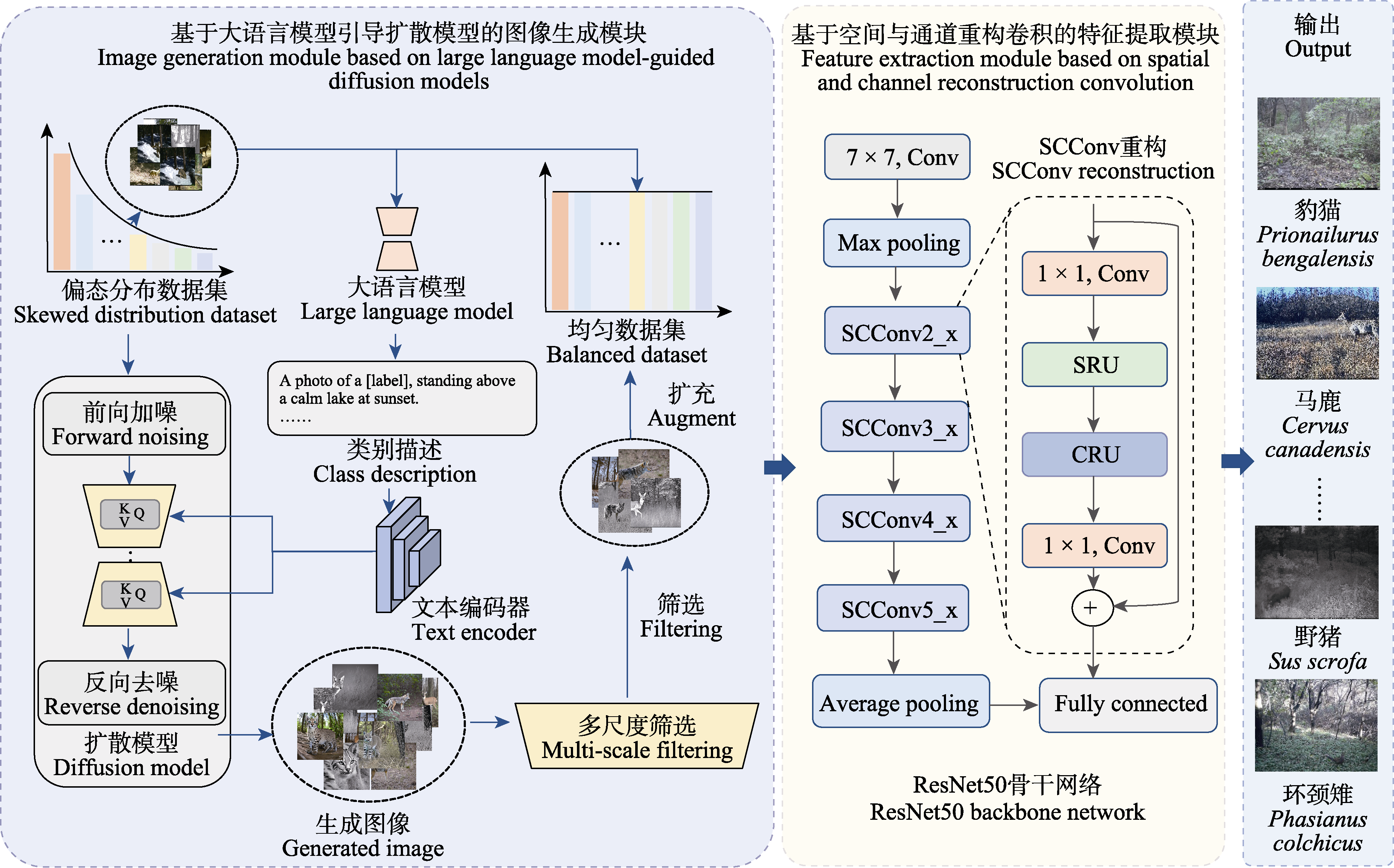
图2 Diff-SCC模型结构。Conv: 卷积; SCConv2_x-SCConv5_x: SCConv残差阶段; Max pooling: 最大池化; Average pooling: 平均池化; Fully connected: 全连接层; SRU: 空间重构单元; CRU: 通道重构单元。
Fig. 2 The architecture of Diff-SCC model. Conv, Convolution; SCConv2_x-SCConv5_x, Residual stages of SCConv; SRU, Spatial reconstruction unit; CRU, Channel reconstruction unit.
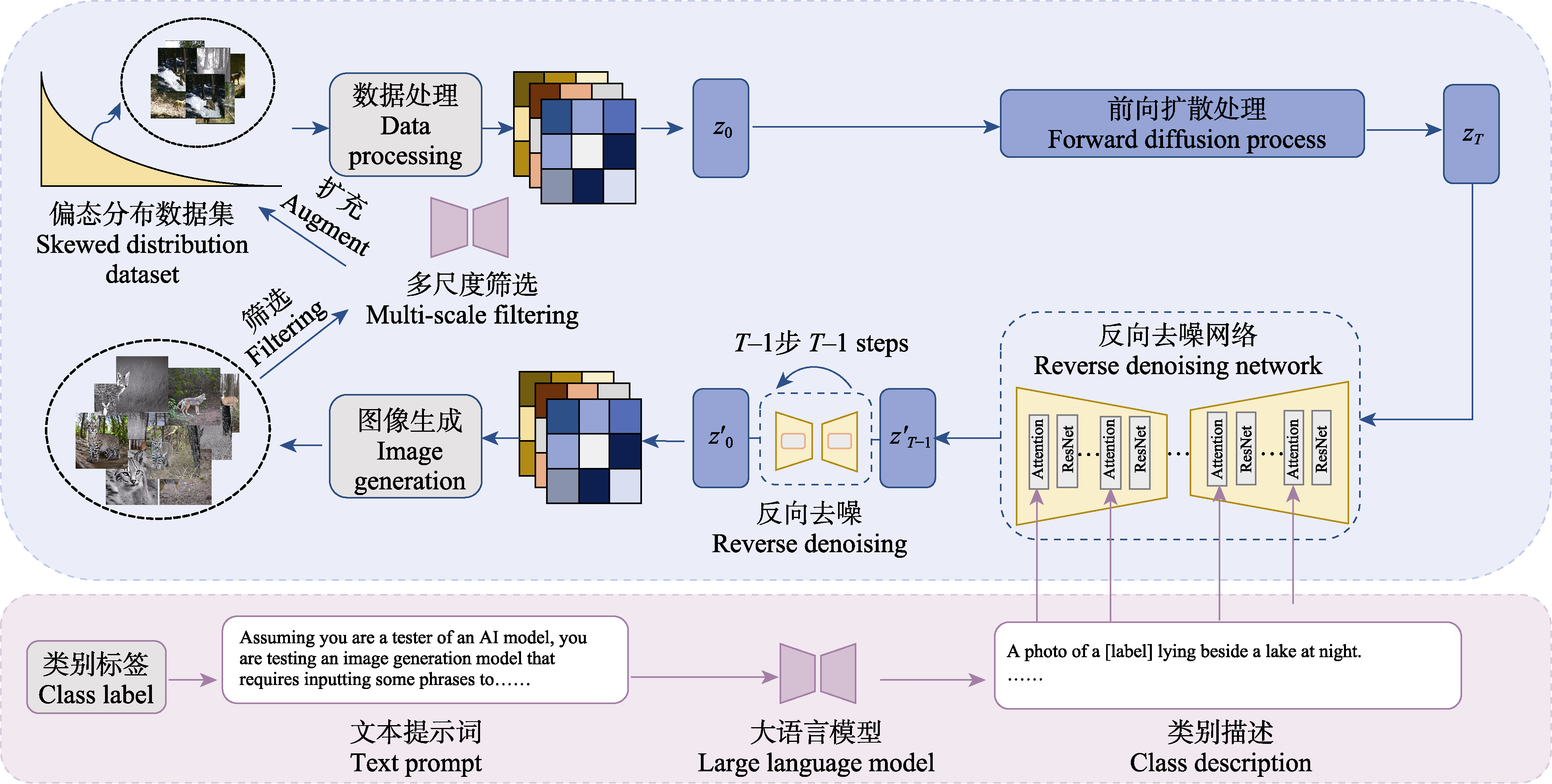
图3 基于扩散模型的偏态分布数据集扩充示意图。z0: 原始图像的初始潜在表示; zT: 经过T步高斯噪声添加后的潜在表示;${{{z}'}_{T1}}$: 在反向扩散过程中的中间潜在表示; ${{{z}'}_{0}}$: 扩散模型生成的图像潜在表示。
Fig. 3 Expansion of skewed dataset based on diffusion model. z0, Initial latent representation of the original image; zT, Latent representation after T steps of Gaussian noise addition; ${{{z}'}_{T1}}$, Intermediate latent representation in the reverse diffusion process;${{{z}'}_{0}}$, Latent representation of the image generated by the diffusion model.
| 编号 Number | 提示模板 Prompt template | 备注 Remarks |
|---|---|---|
| 1 | A photo of a [label] | 基础模板 Basic template |
| 2 | A photo of a [label] at sunset | 添加时间描述 Adding temporal description |
| 3 | A photo of a [label] in winter | 添加时间描述 Adding temporal description |
| 4 | A photo of a [label] at the edge of a canyon | 添加地点描述 Adding location description |
| 5 | A photo of a [label] in a misty forest clearing | 添加地点描述 Adding location description |
| 6 | A photo of a [label] nesting | 添加动作描述 Adding action description |
| 7 | A photo of a [label] landing gracefully | 添加动作描述 Adding action description |
| 8 | A photo of a [label] lying beside a lake at night | 添加3个描述 Adding three descriptions |
表2 文本提示模板
Table 2 Text prompt template
| 编号 Number | 提示模板 Prompt template | 备注 Remarks |
|---|---|---|
| 1 | A photo of a [label] | 基础模板 Basic template |
| 2 | A photo of a [label] at sunset | 添加时间描述 Adding temporal description |
| 3 | A photo of a [label] in winter | 添加时间描述 Adding temporal description |
| 4 | A photo of a [label] at the edge of a canyon | 添加地点描述 Adding location description |
| 5 | A photo of a [label] in a misty forest clearing | 添加地点描述 Adding location description |
| 6 | A photo of a [label] nesting | 添加动作描述 Adding action description |
| 7 | A photo of a [label] landing gracefully | 添加动作描述 Adding action description |
| 8 | A photo of a [label] lying beside a lake at night | 添加3个描述 Adding three descriptions |
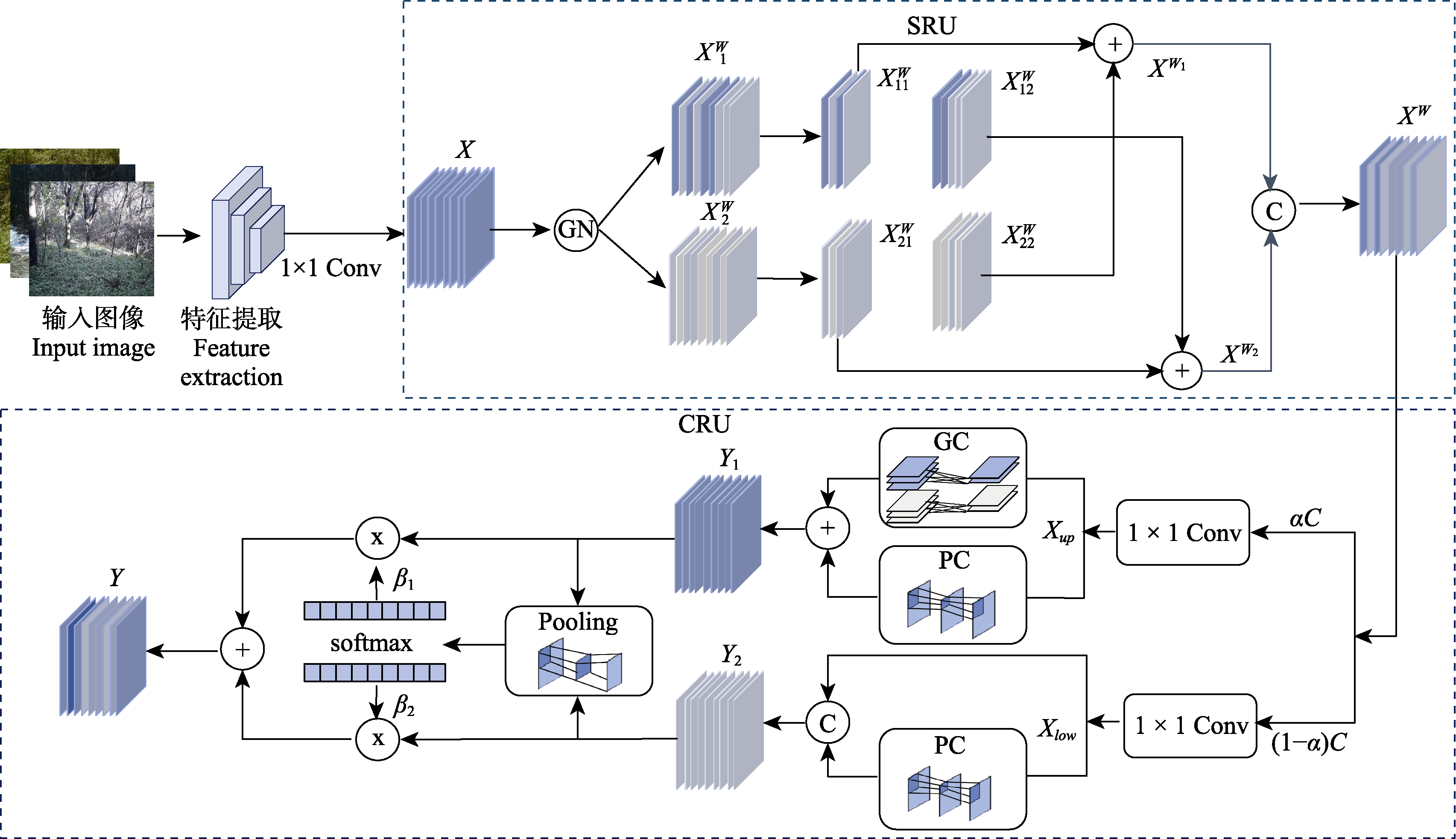
图4 SCConv卷积重建模块流程图。SRU: 空间重构单元; CRU: 通道重构单元; Conv: 卷积单元; X: 输入特征图; GN: 组归一化层; XW 1: 高信息量的特征图; XW 2: 低信息量的特征图; XW 11: 高信息特征的自重构部分; XW 12: 向低信息部分注入的交叉特征; XW 22: 低信息特征的自重构部分; XW 21: 向高信息部分的辅助补充; XW1, XW2 : 增强后的特征图; XW: 空间卷积重构后特征图; αC, (1-α)C: 主通道与辅助通道; Xup: 主通道部分的特征图; Xlow: 辅助通道部分的特征图; GC: 组卷积; PC: 点卷积; Y1: 上分支输出的主特征; Y2: 下分支输出的辅助特征; Pooling: 池化; β1 , β2: 通道注意力权重; Y: 最终的通道重构特征图。
Fig. 4 The workflow of SCConv convolution reconstruction. SRU, Spatial reconstruction unit; CRU, Channel reconstruction unit; Conv, Convolution unit; X, Input feature map; GN, Group normalization; XW 1, Feature map with high information content; XW 2, Feature map with low information content; XW 11, Self-reconstructed part of the high-information feature; XW 12, Cross features injected into the low-information regions; XW 22, Self-reconstructed part of the low-information feature; XW 21, Auxiliary supplement to the high-information regions; XW1, XW2, Enhanced feature maps after reconstruction; XW, Feature map after spatial convolutional reconstruction; αC, (1-α)C, Main and auxiliary channel splits; Xup, Feature map from the main channel group; Xlow, Feature map from the auxiliary channel group; GC, Group convolution; PC, Point convolution; Y1, Main features output by the upper branch; Y2, Auxiliary features output by the lower branch; Pooling, Pooling layer; β1, β2, Channel attention weights; Y, Final channel-reconstructed feature map.
| 方法 Method | 高频类别准确率 High-frequency accuracy (%) | 中频类别准确率 Middle-frequency accuracy (%) | 低频类别准确率 Low-frequency accuracy (%) | 总体准确率 Overall accuracy (%) | Macro F1 |
|---|---|---|---|---|---|
| ResNet50 | 91.37 | 80.38 | 53.82 | 70.76 | 0.29 |
| LDAM | 92.71 | 79.34 | 53.79 | 71.02 | 0.31 |
| Balanced Softmax | 90.38 | 81.49 | 55.37 | 71.39 | 0.32 |
| AREA | 91.09 | 80.25 | 55.65 | 71.69 | 0.32 |
| PaCo | 89.16 | 84.34 | 56.79 | 72.17 | 0.33 |
| ResLT | 93.22 | 85.62 | 61.88 | 76.28 | 0.38 |
| SADE | 92.53 | 87.43 | 62.34 | 76.58 | 0.38 |
| BalPoE | 92.74 | 88.15 | 62.86 | 77.64 | 0.39 |
| Mixup | 92.75 | 85.11 | 57.28 | 73.74 | 0.35 |
| CycleGAN | 92.48 | 86.49 | 58.59 | 74.53 | 0.36 |
| Diff-SCC | 95.32 | 89.75 | 63.78 | 78.71 | 0.40 |
表3 不同模型在ULB-12数据集的实验结果对比
Table 3 Comparison of experimental results on the ULB-12 dataset among different models
| 方法 Method | 高频类别准确率 High-frequency accuracy (%) | 中频类别准确率 Middle-frequency accuracy (%) | 低频类别准确率 Low-frequency accuracy (%) | 总体准确率 Overall accuracy (%) | Macro F1 |
|---|---|---|---|---|---|
| ResNet50 | 91.37 | 80.38 | 53.82 | 70.76 | 0.29 |
| LDAM | 92.71 | 79.34 | 53.79 | 71.02 | 0.31 |
| Balanced Softmax | 90.38 | 81.49 | 55.37 | 71.39 | 0.32 |
| AREA | 91.09 | 80.25 | 55.65 | 71.69 | 0.32 |
| PaCo | 89.16 | 84.34 | 56.79 | 72.17 | 0.33 |
| ResLT | 93.22 | 85.62 | 61.88 | 76.28 | 0.38 |
| SADE | 92.53 | 87.43 | 62.34 | 76.58 | 0.38 |
| BalPoE | 92.74 | 88.15 | 62.86 | 77.64 | 0.39 |
| Mixup | 92.75 | 85.11 | 57.28 | 73.74 | 0.35 |
| CycleGAN | 92.48 | 86.49 | 58.59 | 74.53 | 0.36 |
| Diff-SCC | 95.32 | 89.75 | 63.78 | 78.71 | 0.40 |
| 方法 Method | 高频类别准确率 High-frequency accuracy (%) | 中频类别准确率 Middle-frequency accuracy (%) | 低频类别准确率 Low-frequency accuracy (%) | 总体准确率 Overall accuracy (%) | Macro F1 | |
|---|---|---|---|---|---|---|
| ResNet50 | 89.66 | 87.11 | 56.56 | 73.57 | 0.29 | |
| LDAM | 87.24 | 87.25 | 59.59 | 74.67 | 0.31 | |
| Balanced Softmax | 87.24 | 87.39 | 60.04 | 74.93 | 0.30 | |
| AREA | 88.34 | 88.68 | 57.70 | 74.18 | 0.31 | |
| PaCo | 90.16 | 88.54 | 62.15 | 76.76 | 0.32 | |
| ResLT | 92.76 | 88.57 | 64.36 | 78.13 | 0.34 | |
| SADE | 91.35 | 90.17 | 64.63 | 78.72 | 0.34 | |
| BalPoE | 92.39 | 89.55 | 64.84 | 78.67 | 0.34 | |
| Mixup | 90.88 | 89.32 | 60.96 | 76.64 | 0.32 | |
| CycleGAN | 92.16 | 89.16 | 62.21 | 77.31 | 0.35 | |
| Diff-SCC | 94.75 | 90.84 | 66.55 | 80.84 | 0.36 | |
表4 不同模型在NACTI数据集的实验结果对比
Table 4 Comparison of experimental results on the NACTI dataset among different models
| 方法 Method | 高频类别准确率 High-frequency accuracy (%) | 中频类别准确率 Middle-frequency accuracy (%) | 低频类别准确率 Low-frequency accuracy (%) | 总体准确率 Overall accuracy (%) | Macro F1 | |
|---|---|---|---|---|---|---|
| ResNet50 | 89.66 | 87.11 | 56.56 | 73.57 | 0.29 | |
| LDAM | 87.24 | 87.25 | 59.59 | 74.67 | 0.31 | |
| Balanced Softmax | 87.24 | 87.39 | 60.04 | 74.93 | 0.30 | |
| AREA | 88.34 | 88.68 | 57.70 | 74.18 | 0.31 | |
| PaCo | 90.16 | 88.54 | 62.15 | 76.76 | 0.32 | |
| ResLT | 92.76 | 88.57 | 64.36 | 78.13 | 0.34 | |
| SADE | 91.35 | 90.17 | 64.63 | 78.72 | 0.34 | |
| BalPoE | 92.39 | 89.55 | 64.84 | 78.67 | 0.34 | |
| Mixup | 90.88 | 89.32 | 60.96 | 76.64 | 0.32 | |
| CycleGAN | 92.16 | 89.16 | 62.21 | 77.31 | 0.35 | |
| Diff-SCC | 94.75 | 90.84 | 66.55 | 80.84 | 0.36 | |
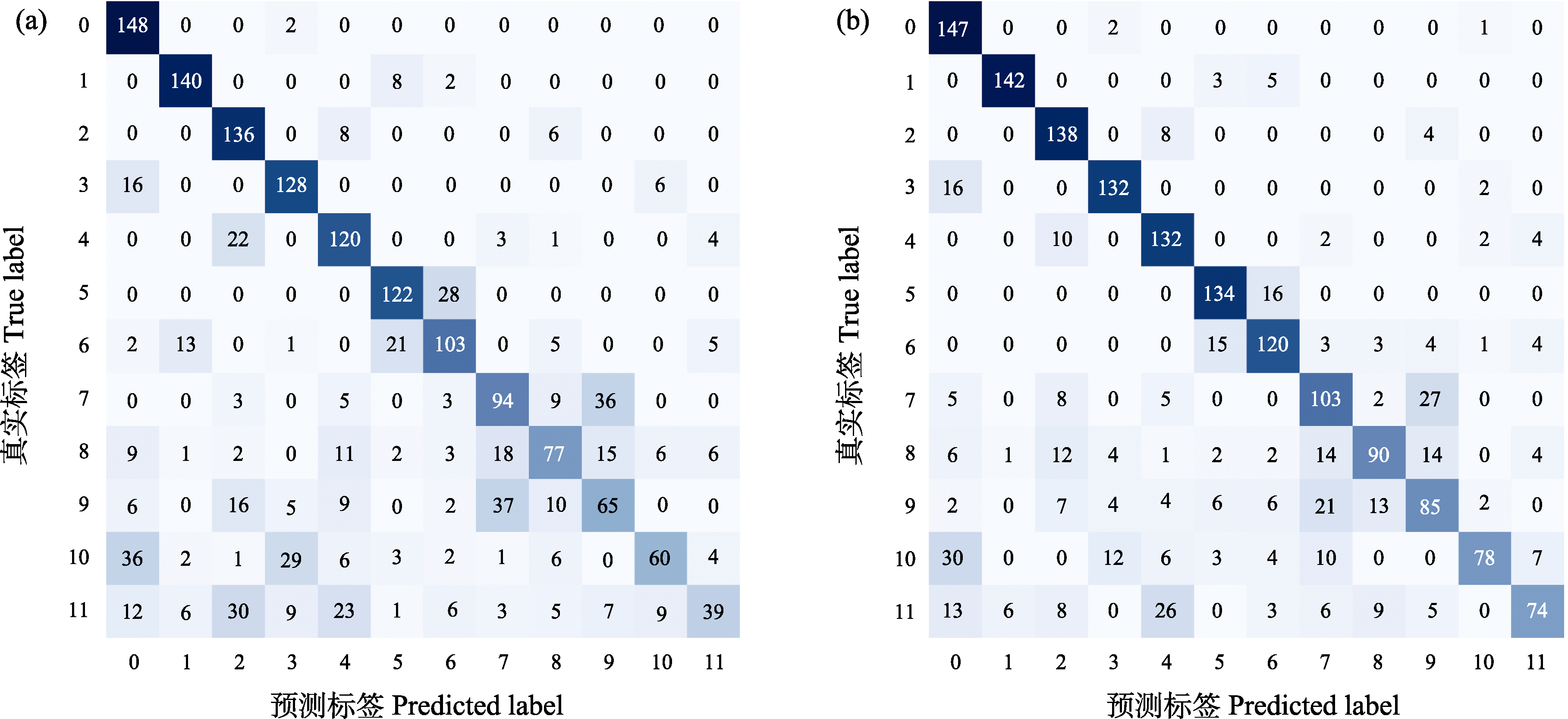

图5 ULB-12数据集上ResNet50 (a)和Diff-SCC (b)模型的分类预测与实际情况对比。矩阵颜色深浅反映对应位置的样本数量, 颜色越深表示样本数越多, 颜色越浅表示样本数越少。对角线区域颜色越集中、越深, 说明模型对该类别的识别能力越强; 非对角线区域颜色越明显, 表明类别间混淆越严重。
Fig. 5 Comparison of predicted vs. actual classification for ResNet50 (a) and Diff-SCC (b) models on the ULB-12 Dataset. The color intensity in the confusion matrix reflects the number of samples at each position: Darker shades indicate a larger number of samples, whereas lighter shades indicate fewer samples. Therefore, a darker and more concentrated diagonal region suggests stronger recognition performance for the corresponding class, while more pronounced off-diagonal regions indicate more severe inter-class confusion.
| 方法 Method | 弗雷歇距离 FID | 方法学习感知图像 块相似度 LPIPS | 准确率 Accuracy (%) |
|---|---|---|---|
| Mixup | 39.5 | 0.42 | 73.74 |
| CycleGAN | 17.6 | 0.28 | 74.53 |
| Diff-SCC | 12.4 | 0.21 | 78.71 |
表5 不同数据增强方法生成图像质量对比
Table 5 Comparison of image quality generated by different data augmentation methods
| 方法 Method | 弗雷歇距离 FID | 方法学习感知图像 块相似度 LPIPS | 准确率 Accuracy (%) |
|---|---|---|---|
| Mixup | 39.5 | 0.42 | 73.74 |
| CycleGAN | 17.6 | 0.28 | 74.53 |
| Diff-SCC | 12.4 | 0.21 | 78.71 |
| 低频类别图像扩充策略 Low-frequency class image expansion strategy | 空间和通道卷积重构单元 Spatial and channel convolution reconstruction unit | 准确率 Accuracy (%) | 模型规模 Model size (MB) | 浮点运算次数 FLOPs (× 109) | 单张图像推理时间 Inference time per image (ms) |
|---|---|---|---|---|---|
| - | - | 70.76 | 90.14 | 4.13 | 3.8 |
| - | + | 73.86 | 56.44 | 2.66 | 2.5 |
| + | - | 77.65 | 90.14 | 4.13 | 3.8 |
| + | + | 78.71 | 56.44 | 3.13 | 2.6 |
表6 消融实验结果
Table 6 Ablation experiment results
| 低频类别图像扩充策略 Low-frequency class image expansion strategy | 空间和通道卷积重构单元 Spatial and channel convolution reconstruction unit | 准确率 Accuracy (%) | 模型规模 Model size (MB) | 浮点运算次数 FLOPs (× 109) | 单张图像推理时间 Inference time per image (ms) |
|---|---|---|---|---|---|
| - | - | 70.76 | 90.14 | 4.13 | 3.8 |
| - | + | 73.86 | 56.44 | 2.66 | 2.5 |
| + | - | 77.65 | 90.14 | 4.13 | 3.8 |
| + | + | 78.71 | 56.44 | 3.13 | 2.6 |
| [1] | Aimar ES, Jonnarth A, Felsberg M, Kuhlmann M (2023) Balanced product of calibrated experts for long-tailed recognition. In: 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 19967-19977. Vancouver, Canada. |
| [2] | Cao KD, Wei C, Gaidon A, Arechiga N, Ma TY (2019) Learning imbalanced datasets with label-distribution-aware margin loss. In:2019 Annual Conference on Neural Information Processing Systems (NeurIPS), pp. 1565-1576. Vancouver, Canada. |
| [3] | Chen JH, Su B (2023) Transfer knowledge from head to tail: Uncertainty calibration under long-tailed distribution. In: 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 19978-19987. Vancouver, Canada. |
| [4] | Chen XH, Zhou YC, Wu DY, Yang CL, Li B, Hu QH, Wang WP (2023) AREA: Adaptive reweighting via effective area for long-tailed classification. In: 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 19277-19287. Vancouver, Canada. |
| [5] | Cui JQ, Liu S, Tian ZT, Zhong ZS, Jia JY (2023) ResLT: Residual learning for long-tailed recognition. IEEE Transactions on Pattern Analysis and Machine Intelligence, 45, 3695-3706. |
| [6] | Cui JQ, Zhong ZS, Liu S, Yu B, Jia JY (2021) Parametric contrastive learning. In: 2021 IEEE/CVF International Conference on Computer Vision (ICCV), pp. 695-704. Montreal, Canada. |
| [7] | Cunha F, dos Santos EM, Colonna JG (2023) Bag of tricks for long-tail visual recognition of animal species in camera-trap images. Ecological Informatics, 76, 102060. |
| [8] | Deng CX, Li DF, Ji L, Zhang CY, Li BC, Yan HY, Zheng JY, Wang LF, Zhang JG (2025) ChatDiff: A ChatGPT-based diffusion model for long-tailed classification. Neural Networks, 181, 106794. |
| [9] | Fabian Z, Miao ZQ, Li CY, Zhang YH, Liu ZW, Hernández A, Montes-Rojas A, Escucha R, Siabatto L, Link A, Arbeláez P, Dodhia R, Ferres LF (2023) Multimodal foundation models for zero-shot animal species recognition in camera trap images. arXiv, doi:10.48550/arXiv.2311.01064. |
| [10] | Gu JQ, Chen JP, Lai JS (2024) Application of large language models in biodiversity research. Biodiversity Science, 32, 24258.(in Chinese with English abstract) |
| [谷际岐, 陈建平, 赖江山 (2024) 大语言模型在生物多样性研究中的应用. 生物多样性, 32, 24258.] | |
| [11] | Guo YT, Li SC, Wang Z, Xie Y, Yang X, Zhou GJ, You CH, Zhu SN, Gao JX (2025) Coverage and distribution of national key protected wild species in China’s nature reserves. Biodiversity Science, 33, 24423.(in Chinese with English abstract) |
| [郭雨桐, 李素萃, 王智, 解焱, 杨雪, 周广金, 尤春赫, 朱萨宁, 高吉喜 (2025) 全国自然保护地对国家重点保护野生物种的覆盖度及其分布状况. 生物多样性, 33, 24423.] | |
| [12] | He KM, Zhang XY, Ren SQ, Sun J (2016) Deep residual learning for image recognition. In: 2016 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 770-778. Las Vegas, USA. |
| [13] | Hernandez A, Miao ZQ, Vargas L, Beery S, Dodhia R, Arbelaez P (2024) Pytorch-Wildlife: A collaborative deep learning framework for conservation. arXiv, doi:10.48550/arXiv.2405.12930. |
| [14] | Ho J, Jain A, Abbeel P (2020) Denoising diffusion probabilistic models. Advances in Neural Information Processing Systems, 33, 6840-6851. |
| [15] | Jin Y, Li MK, Lu Y, Cheung YM, Wang HZ (2023) Long-tailed visual recognition via self-heterogeneous integration with knowledge excavation. In: 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 23695-23704. Vancouver, Canada. |
| [16] | Kim J, Woo S, Park B, Kim C (2022) Temporal flow mask attention for open-set long-tailed recognition of wild animals in camera-trap images. In: 2022 IEEE International Conference on Image Processing (ICIP), pp. 2152-2156. Bordeaux, France. |
| [17] | Li JF, Wen Y, He LH (2023) SCConv: Spatial and channel reconstruction convolution for feature redundancy. In: 2023 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 6153-6162. Vancouver, Canada. |
| [18] | Li MK, Cheung YM, Lu Y (2022) Long-tailed visual recognition via Gaussian clouded logit adjustment. In: 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 6929-6938. New Orleans, USA. |
| [19] | Li X, Wang WH, Hu XL, Yang J (2019) Selective kernel networks. In: 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 510-519. Long Beach, USA. |
| [20] | Mangla P, Singh M, Sinha A, Kumari N, Balasubramanian VN, Krishnamurthy B (2020) Charting the right manifold: Manifold Mixup for few-shot learning. In: 2020 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV), pp. 2207-2216. Snowmass Village, USA. |
| [21] | Miao ZQ, Gaynor KM, Wang JY, Liu ZW, Muellerklein O, Norouzzadeh MS, McInturff A, Bowie RCK, Nathan R, Yu SX, Getz WM (2019) Insights and approaches using deep learning to classify wildlife. Scientific Reports, 9, 8137. |
| [22] | Miao ZQ, Liu ZW, Gaynor KM, Palmer MS, Yu SX, Getz WM (2021) Iterative human and automated identification of wildlife images. Nature Machine Intelligence, 3, 885-895. |
| [23] | Mugerwa B, Niedballa J, Planillo A, Sheil D, Kramer-Schadt S, Wilting A (2024) Global disparity of camera trap research allocation and defaunation risk of terrestrial mammals. Remote Sensing in Ecology and Conservation, 10, 121-136. |
| [24] | Norman DL, Bischoff PH, Wearn OR, Ewers RM, Rowcliffe JM, Evans B, Sethi S, Chapman PM, Freeman R (2023) Can CNN-based species classification generalise across variation in habitat within a camera trap survey? Methods in Ecology and Evolution, 14, 242-251. |
| [25] | Qi JD, Zheng SZ, Chen ZY, Ma ZT (2024) Wildlife image recognition of infrared cameras in Beijing area based on an improvement ConvNeXt model. Scientia Silvae Sinicae, 60(8), 33-45.(in Chinese with English abstract) |
| [齐建东, 郑尚姿, 陈子仪, 马鐘添 (2024) 基于ConvNeXt的北京地区红外相机野生动物图像识别改进模型构建. 林业科学, 60(8), 33-45.] | |
| [26] | Radford A, Kim JW, Hallacy C, Ramesh A, Goh G, Agarwal S, Sastry G, Askell A, Mishkin P, Clark J, Krueger G, Sutskever I (2021) Learning transferable visual models from natural language supervision. In:2021 Proceedings of Machine Learning Research (PMLR), pp. 8748-8763. Virtual meeting. |
| [27] | Rangwani H, Jaswani N, Karmali T, Jampani V, Babu RV (2022) Improving GANs for long-tailed data through group spectral regularization. In:2022 European Conference on Computer Vision (ECCV), pp. 426-442. Tel Aviv, Israel. |
| [28] | Ren JW, Yu CJ, Sheng SN, Ma X (2020) Balanced meta-softmax for long-tailed visual recognition. In:2020 Annual Conference on Neural Information Processing Systems (NeurIPS), pp. 4175-4186. Virtual meeting. |
| [29] | Rombach R, Blattmann A, Lorenz D, Esser P, Ommer B (2022) High-resolution image synthesis with latent diffusion models. In: 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 10684-10695. New Orleans, USA. |
| [30] | Senior RA, Oliveira BF, Dale J, Scheffers BR (2022) Wildlife trade targets colorful birds and threatens the aesthetic value of nature. Current Biology, 32, 4299-4305. |
| [31] | Shi JX, Wei T, Xiang Y, Li YF (2023) How re-sampling helps for long-tail learning? Advances in Neural Information Processing Systems, 36, 75669-75687. |
| [32] | Tabak MA, Norouzzadeh MS, Wolfson DW, Sweeney SJ, Vercauteren KC, Snow NP, Halseth JM, Di Salvo PA, Lewis JS, White MD, Teton B, Beasley JC, Schlichting PE, Bougthon RK, Wight B, Newkirk ES, Ivan JS, Odell EA, Brook RK, Lukcas PM, Moeller AK, Mandeville EG, Clune J, Miller RS (2019) Machine learning to classify animal species in camera trap images: Applications in ecology.Methods in Ecology and Evolution, 10, 585-590. |
| [33] | Tuia D, Kellenberger B, Beery S, Costelloe BR, Zuffi S, Risse B, Mathis A, Mathis MW, van Langevelde F, Burghardt T, Kays R, Klinck H, Wikelski M, Couzin ID, van Horn G, Crofoot MC, Stewart CV, Berger-Wolf T (2022) Perspectives in machine learning for wildlife conservation. Nature Communications, 13, 792. |
| [34] | Wang LF, Wang S, Deng CX, Zhu HW, Tian Y, Zhang JG (2025) DeLoCo: Decoupled location context-guided framework for wildlife species classification using camera trap images. Ecological Informatics, 85, 102949. |
| [35] | Wang XD, Yu SX (2021) Tied Block convolution: Leaner and better CNNs with shared thinner filters. In:2021 Proceedings of the AAAI Conference on Artificial Intelligence (AAAI), pp. 10227-10235. Virtual meeting. |
| [36] | Wu YX, He KM (2018) Group normalization. In:2018 European Conference on Computer Vision (ECCV), pp. 3-19. Munich, Germany. |
| [37] | Xiao ZS, Xiao WH, Wang TM, Li S, Lian XM, Song DZ, Deng XQ, Zhou QH (2022) Wildlife monitoring and research using camera-trapping technology across China: The current status and future issues. Biodiversity Science, 30, 22451.(in Chinese with English abstract) |
| [肖治术, 肖文宏, 王天明, 李晟, 连新明, 宋大昭, 邓雪琴, 周岐海 (2022) 中国野生动物红外相机监测与研究: 现状及未来. 生物多样性, 30, 22451.] | |
| [38] | Xu Q, Xie YH (2024) Automatic individual tracking method of Amur tiger based on attention mechanism fusion of multiple features. Biodiversity Science, 32, 23409.(in Chinese with English abstract) |
| [许群, 谢永华 (2024) 基于注意力机制融合多特征的东北虎个体自动跟踪方法. 生物多样性, 32, 23409.] | |
| [39] | Yang L, Jiang H, Song Q, Guo J (2022) A survey on long-tailed visual recognition. International Journal of Computer Vision, 130, 1837-1872. |
| [40] | Zhang CC, Li DF, Zhang JG (2024) Wildlife images recognition method based on Wasserstein distance and correlation alignment transfer learning. Scientia Silvae Sinicae, 60(8), 25-32.(in Chinese with English abstract) |
| [张长春, 李大方, 张军国 (2024) 基于Wasserstein距离和相关对齐迁移学习的野生动物图像识别方法. 林业科学, 60(8), 25-32.] | |
| [41] | Zhang CS, Almpanidis G, Fan GJ, Deng BQ, Zhang YB, Liu J, Kamel A, Soda P, Gama J (2025) A systematic review on long-tailed learning. IEEE Transactions on Neural Networks and Learning Systems, 36, 13670-13690. |
| [42] | Zhang QY, Yi XL, Guo JL, Tang YD, Feng T, Liu R (2023) A few-shot rare wildlife image classification method based on style migration data augmentation. Ecological Informatics, 77, 102237. |
| [43] | Zhang XY, Zhang HL, Han YY, Weng Q, Yuan ZR, Yao Y (2022) Research progress of the wildlife monitoring and identification based on deep learning. Journal of Wildlife, 43, 251-258.(in Chinese with English abstract) |
| [张雪莹, 张浩林, 韩莹莹, 翁强, 袁峥嵘, 姚远 (2022) 基于深度学习的野生动物监测与识别研究进展. 野生动物学报, 43, 251-258.] | |
| [44] | Zhang YF, Hooi B, Hong LQ, Feng JS (2022) Self-supervised aggregation of diverse experts for test-agnostic long-tailed recognition. In: 2022 Annual Conference on Neural Information Processing Systems (NeurIPS), pp. 34077-34090. New Orleans, USA. |
| [45] | Zhang YF, Kang BY, Hooi B, Yan SC, Feng JS (2023) Deep long-tailed learning: A survey. IEEE Transactions on Pattern Analysis and Machine Intelligence, 45, 10795-10816. |
| [46] | Zhao QH, Dai YL, Li H, Hu W, Zhang F, Liu J (2024) LTGC: Long-tail recognition via leveraging LLMs-driven generated content. In: 2024 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 19510-19520. Seattle, USA. |
| [47] | Zhu HW, Tian Y, Zhang JG (2022) Class incremental learning for wildlife biodiversity monitoring in camera trap images. Ecological Informatics, 71, 101760. |
| [48] | Zhu JY, Park T, Isola P, Efros AA (2017) Unpaired image-to-image translation using cycle-consistent adversarial networks. In:2017 Proceedings of the IEEE International Conference on Computer Vision (ICCV), pp. 2242-2251. Venice, Italy. |
| [1] | 孔孜亦, 王德港, 王建涛, 裴志永, 孙晶, 张长春, 张军国. 基于SCD-HRNet模型的野生动物姿态估计方法: 以内蒙古赛罕乌拉地区为例[J]. 生物多样性, 2026, 34(4): 25287-. |
| [2] | 郑俊妮, 尚袁凌博, 罗堯, 魏营, 高志伟, 周宗泽, 廖凌娟, 杨道德. 地方重点保护野生动物名录调整方法探究: 以湖南省陆生脊椎动物为例[J]. 生物多样性, 2025, 33(8): 25055-. |
| [3] | 田璐瑶, 尹豪. 基于生物多样性保护的我国生态铁路现状和对策研究[J]. 生物多样性, 2025, 33(8): 24495-. |
| [4] | 毛静, 王婧, 黄杰, 熊姝红, 张自亮, 张佑祥, 吴涛. 湖南高望界国家级自然保护区2021-2023年鸟兽多样性监测数据集[J]. 生物多样性, 2025, 33(6): 24489-. |
| [5] | 付梦娣, 朱彦鹏, 任月恒, 李爽, 秦乐, 谢正君, 王清春, 张立博. 新疆野生动物通道空间布局优化[J]. 生物多样性, 2025, 33(3): 24346-. |
| [6] | 王大伟, 程帅, 冯佳伟, 王天明. 东北地区张广才岭2015-2020年野生动物红外相机监测数据集[J]. 生物多样性, 2025, 33(2): 24384-. |
| [7] | 安家宁, 张长春, 王建涛, 裴志永, 白丹丹, 张军国. 融合对抗解耦与特征对齐的野生动物图像开集域适应方法[J]. 生物多样性, 2025, 33(12): 25283-. |
| [8] | 蒋承汛, 张塔星, 权子豪, 刘郢, 柴璐艳, 冉江洪. 青藏高原国家重点保护野生鸟类丰富度空间格局及热点区域[J]. 生物多样性, 2025, 33(11): 25171-. |
| [9] | 李佳琪, 冯一迪, 王蕾, 潘盆艳, 刘潇如, 李雪阳, 王怡涵, 王放. 上海城市环境中貉的食性分析及家域范围内的栖息地选择[J]. 生物多样性, 2024, 32(8): 24131-. |
| [10] | 巴苏艳, 赵春艳, 刘媛, 方强. 通过虫体花粉识别构建植物‒传粉者网络: 人工模型与AI模型高度一致[J]. 生物多样性, 2024, 32(6): 24088-. |
| [11] | 李柏灿, 张军国, 张长春, 王丽凤, 徐基良, 刘利. 基于TC-YOLO模型的北京珍稀鸟类识别方法[J]. 生物多样性, 2024, 32(5): 24056-. |
| [12] | 鲁彬悦, 李坤, 王晨溪, 李晟. 基于传感器标记的野生动物追踪技术在中国的应用现状与展望[J]. 生物多样性, 2024, 32(5): 23497-. |
| [13] | 秦涛, 崔荣赫, 宋蕊, 富丽莎. 我国野生动物肇事公众责任保险: 发展模式、现实困境与优化策略[J]. 生物多样性, 2024, 32(5): 23431-. |
| [14] | 朱建国, 王林, 任国鹏. 《国家重点保护野生动物名录》调整的评估方法探讨[J]. 生物多样性, 2023, 31(8): 23045-. |
| [15] | 陈金锋, 吴欣静, 林海, 崔国发. 《国家重点保护野生动物名录》和其他保护名录对比分析[J]. 生物多样性, 2023, 31(6): 22639-. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
备案号:京ICP备16067583号-7
Copyright © 2026 版权所有 《生物多样性》编辑部
地址: 北京香山南辛村20号, 邮编:100093
电话: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn
![]()