



Biodiv Sci ›› 2012, Vol. 20 ›› Issue (6): 693-702. DOI: 10.3724/SP.J.1003.2012.10062 cstr: 32101.14.SP.J.1003.2012.10062
• Research Report • Previous Articles Next Articles
Ni Xiang1,2, Yannong Xiao2, Canxing Duan1, Xiaoming Wang1, Zhendong Zhu1,*( )
)
Received:2012-03-02
Accepted:2012-07-16
Online:2012-11-20
Published:2013-01-04
Contact:
Zhendong Zhu
Ni Xiang, Yannong Xiao, Canxing Duan, Xiaoming Wang, Zhendong Zhu. Genetic diversity in Fusarium solani f. sp. pisi based on SSR markers[J]. Biodiv Sci, 2012, 20(6): 693-702.
| 来源 Location | 数量 Amount | 分离物 Isolates |
|---|---|---|
| 安徽 Anhui | 17 | AH-1a、AH-2、AH-3、AH-4、AH-5、AH-6、AH-7、AH-8、AH-9、AH-10、AH-11、AH-12、AH-13、AH-14b、AH-15a、AH-16、AH-17 |
| 北京 Beijing | 13 | BJ-1、BJ-2、BJ-3、BJ-4、BJ-5、BJ-6、BJ-7、BJ-8、BJ-9、BJ-10、BJ-11、BJ-12、BJ-13 |
| 福建 Fujian | 4 | FJ-1b、FJ-2a、FJ-3b、FJ-4a |
| 甘肃 Gansu | 20 | LZ-1、LZ-2、LZ-3b、LZ-4、LZ-5、LZ-6、DX-1、DX-2、DX-3、DX-4a、DX-5、DX-6、DX-7、DX-8、DX-9、DX-10、DX-11、DX-12、DX-13、DX-14 |
| 河北 Hebei | 10 | ZB-1、ZB-2、ZB-3、ZB-4、ZB-5、ZB-6、ZB-7、ZB-8、ZB-9、ZB-10 |
| 内蒙古 Inner Mongolia | 7 | NM-1、NM-2、NM-3、NM-4b、NM-5、NM-6b、NM-7 |
| 吉林 Jilin | 1 | BC-1a |
| 宁夏 Ningxia | 12 | SB-1、SB-2、SB-3、SB-4、SB-5、SB-6、SB-7、SB-8、XB-1、XB-2、PC-1、PC-2 |
| 青海 Qinghai | 5 | QH-1、QH-2、QH-3b、QH-4b、QH-5b |
| 四川 Sichuan | 3 | SC-1、SC-2、SC-3 |
| 云南 Yunnan | 4 | YN-1a、YN-2a、YN-3、YN-4 |
Table 1 Ninety-six isolates of Fusarium solani f. sp. pisi used in this study
| 来源 Location | 数量 Amount | 分离物 Isolates |
|---|---|---|
| 安徽 Anhui | 17 | AH-1a、AH-2、AH-3、AH-4、AH-5、AH-6、AH-7、AH-8、AH-9、AH-10、AH-11、AH-12、AH-13、AH-14b、AH-15a、AH-16、AH-17 |
| 北京 Beijing | 13 | BJ-1、BJ-2、BJ-3、BJ-4、BJ-5、BJ-6、BJ-7、BJ-8、BJ-9、BJ-10、BJ-11、BJ-12、BJ-13 |
| 福建 Fujian | 4 | FJ-1b、FJ-2a、FJ-3b、FJ-4a |
| 甘肃 Gansu | 20 | LZ-1、LZ-2、LZ-3b、LZ-4、LZ-5、LZ-6、DX-1、DX-2、DX-3、DX-4a、DX-5、DX-6、DX-7、DX-8、DX-9、DX-10、DX-11、DX-12、DX-13、DX-14 |
| 河北 Hebei | 10 | ZB-1、ZB-2、ZB-3、ZB-4、ZB-5、ZB-6、ZB-7、ZB-8、ZB-9、ZB-10 |
| 内蒙古 Inner Mongolia | 7 | NM-1、NM-2、NM-3、NM-4b、NM-5、NM-6b、NM-7 |
| 吉林 Jilin | 1 | BC-1a |
| 宁夏 Ningxia | 12 | SB-1、SB-2、SB-3、SB-4、SB-5、SB-6、SB-7、SB-8、XB-1、XB-2、PC-1、PC-2 |
| 青海 Qinghai | 5 | QH-1、QH-2、QH-3b、QH-4b、QH-5b |
| 四川 Sichuan | 3 | SC-1、SC-2、SC-3 |
| 云南 Yunnan | 4 | YN-1a、YN-2a、YN-3、YN-4 |
| 位点a Locusa | 引物序列 Primer sequence(5'-3') | 重复基元 Repeat motif | 基因组位置b Location in the genome b | 期望大小 Expected size (bp) | 片段大小 Allele size range (bp) | 退火温度 Annealing tem- perature (℃) |
|---|---|---|---|---|---|---|
| Fsp1.7 | F: TGTTCCCATAGTCCGATTCCG | (AC)17 | Contig 1: 3457867-3458072 | 206 | 180-220 | 54 |
| R: GCCGCTTGACTTGCTAACCA | ||||||
| Fsp1.10 | F: CGATTCGTTCTTCCCAACACC | (TCT)14 | Contig 1/: 709046-709225 | 180 | 147-242 | 54 |
| R: AAACCTGCCTCAAGCCCTGA | ||||||
| Fsp1.17 | F: AATCCCATTGTTTCGGTGCG | (GT)21 | Contig 1/: 245108-245324 | 217 | 165-217 | 54 |
| R: CCGTTGAGGAATAGAGAGGCA | ||||||
| Fsp2.2 | F: GCTTTGGGTCCAGGTATTACG | (AC)11 | Contig 15/: 2408632-2409004 | 373 | 600-622 | 54 |
| R: TTCTCTCTCTCCCGTCACCA | ||||||
| Fsp2.3 | F: TGCTCCTACACCTCCGTCTC | (TG)12 | Contig 15: 1878221-1878425 | 205 | 160-209 | 54 |
| R: CACGTTTTATCTGCCTTGCTG | ||||||
| Fsp2.4 | F: GATGATTGTGCTGGTAGCCTG | (TTTC)8 | Contig 15: 802092-802323 | 232 | 195-238 | 54 |
| R: CGCTTGACGCCATAATTGAG | ||||||
| Fsp3.1 | F: AACAAGGAGGGTGACGGGAT | (AC)13 | Contig 2/: 1062296-1062559 | 264 | 210-290 | 54 |
| R: AGTGACAAATAGGGAGGTGGG | ||||||
| Fsp3.6 | F: CTTGCCCAAACGCATTGAAG | (TG)16 | Contig 2/: 2066491-2066782 | 292 | 260-320 | 54 |
| R: ACTAACACACACCCAGCCTC | ||||||
| Fsp3.7 | F: CCATTTCCATTCCACCTCACC | (AGC)9 | Contig 2/: 1170545-1170752 | 208 | 180-217 | 52 |
| R: CAGTCTGCTGTTATGAAGGGC | ||||||
| Fsp4.2 | F: TTGACCTCTGCACACCATCAG | (AC)15 | Contig 3/: 2330520-2330703 | 184 | 160-200 | 54 |
| R: CTCACTCTCGGCTCTCGTTAT | ||||||
| Fsp4.4 | F: CGGTCACAGGCAAAGCAAT | (TG)15 | Contig 3/: 2288923-2289149 | 227 | 190-250 | 54 |
| R: GGGATAGAGCAAGCAGCGTT | ||||||
| Fsp4.7 | F: ACACACACACAGGCACAAG | (AC)10 | Contig 4/: 1510008-1510245 | 238 | 201-260 | 54 |
| R: TGAGTAAGGTAGGCTGGGTCT | ||||||
| Fsp5.3 | F: TACACTCCCGTTCCTGGCTA | (TC)15 | Contig 4/: 161679-161833 | 155 | 90-147 | 54 |
| R: CCAAATCTCAACCGTCGTGG | ||||||
| Fsp5.4 | F: TGAATCTGACTGTGGCTGGAG | (TG)11 | Contig 4/: 330399-330579 | 181 | 160-210 | 54 |
| R: ATGGTTCGCCTTGGCTCTGA | ||||||
| Fsp5.8 | F: CAGTTAGGTTCCACTCCAGGT | (ACT)14 | Contig 4/: 216245-216574 | 330 | 260-375 | 54 |
| R: TGGGCGGTCATTGTGTAGGT | ||||||
| Fsp6.3 | F: AGCACGAGGAACGAGTCTA | (AC)15 | Contig 18/: 1864740-1864874 | 135 | 110-150 | 52 |
| R: CGCTACTTGTTTGTGTCCG | ||||||
| Fsp6.7 | F: TGACCGAGAACTGAAGCCA | (TTC)9 | Contig 18/: 2280272-2280530 | 259 | 250-350 | 52 |
| R: GACGACGAAACCTTTGAAGAG | ||||||
| Fsp7.3 | F: CTCTCATTATGTAGCGACGAC | (TCG)10 | Contig 7/: 808299-808470 | 173 | 150-250 | 52 |
| R: TCGATACCCCTGAGTTCTGTG | ||||||
| Fsp9.6 | F: GCATCCAATGTCTTTCCCAAT | (TG)17 | Contig 5/: 189772--190021 | 250 | 230-275 | 52 |
| R: GACTCGATATTCATCAACACCA | ||||||
| Fsp9.8 | F: TTGGTTTCCCTGCCTTGT | (TG)20 | Contig 5/: 1802083-1802348 | 266 | 240-300 | 52 |
| R: TGTTGGGTCATCTTGGTTTC | ||||||
| Fsp9.9 | F: ACGCCACTCTTCCATACTCAG | (AAC)15 | Contig 5/: 764399-764632 | 234 | 200-280 | 54 |
| R: GTTGACGGTCTTGTGGCATAC | ||||||
| Fsp9.10 | F: TCAACAACACCAACCACCTG | (AGC)9 | Contig 5/: 509965-510211 | 247 | 200-300 | 54 |
| R: TTCGTCCGAGTCGCCTTCTA | ||||||
| Fsp9.15 | F: GCTTGTGCTGGAGTTGACCT | (GAAT)8 | Contig 5/: 168328-168492 | 165 | 123-155 | 52 |
| R: CAGGAGCATAGGAGGAATAC | ||||||
| Fsp10.2 | F: CGAGAAAGGTGAGGAAGGGA | (GGA)8 | Contig 11/: 258852-259033 | 182 | 175-200 | 52 |
| R: TATCTGTGAGGTGTGGCGA |
Table 2 Twenty-four polymorphic SSR loci in Fusarium solani f. sp. pisi isolates
| 位点a Locusa | 引物序列 Primer sequence(5'-3') | 重复基元 Repeat motif | 基因组位置b Location in the genome b | 期望大小 Expected size (bp) | 片段大小 Allele size range (bp) | 退火温度 Annealing tem- perature (℃) |
|---|---|---|---|---|---|---|
| Fsp1.7 | F: TGTTCCCATAGTCCGATTCCG | (AC)17 | Contig 1: 3457867-3458072 | 206 | 180-220 | 54 |
| R: GCCGCTTGACTTGCTAACCA | ||||||
| Fsp1.10 | F: CGATTCGTTCTTCCCAACACC | (TCT)14 | Contig 1/: 709046-709225 | 180 | 147-242 | 54 |
| R: AAACCTGCCTCAAGCCCTGA | ||||||
| Fsp1.17 | F: AATCCCATTGTTTCGGTGCG | (GT)21 | Contig 1/: 245108-245324 | 217 | 165-217 | 54 |
| R: CCGTTGAGGAATAGAGAGGCA | ||||||
| Fsp2.2 | F: GCTTTGGGTCCAGGTATTACG | (AC)11 | Contig 15/: 2408632-2409004 | 373 | 600-622 | 54 |
| R: TTCTCTCTCTCCCGTCACCA | ||||||
| Fsp2.3 | F: TGCTCCTACACCTCCGTCTC | (TG)12 | Contig 15: 1878221-1878425 | 205 | 160-209 | 54 |
| R: CACGTTTTATCTGCCTTGCTG | ||||||
| Fsp2.4 | F: GATGATTGTGCTGGTAGCCTG | (TTTC)8 | Contig 15: 802092-802323 | 232 | 195-238 | 54 |
| R: CGCTTGACGCCATAATTGAG | ||||||
| Fsp3.1 | F: AACAAGGAGGGTGACGGGAT | (AC)13 | Contig 2/: 1062296-1062559 | 264 | 210-290 | 54 |
| R: AGTGACAAATAGGGAGGTGGG | ||||||
| Fsp3.6 | F: CTTGCCCAAACGCATTGAAG | (TG)16 | Contig 2/: 2066491-2066782 | 292 | 260-320 | 54 |
| R: ACTAACACACACCCAGCCTC | ||||||
| Fsp3.7 | F: CCATTTCCATTCCACCTCACC | (AGC)9 | Contig 2/: 1170545-1170752 | 208 | 180-217 | 52 |
| R: CAGTCTGCTGTTATGAAGGGC | ||||||
| Fsp4.2 | F: TTGACCTCTGCACACCATCAG | (AC)15 | Contig 3/: 2330520-2330703 | 184 | 160-200 | 54 |
| R: CTCACTCTCGGCTCTCGTTAT | ||||||
| Fsp4.4 | F: CGGTCACAGGCAAAGCAAT | (TG)15 | Contig 3/: 2288923-2289149 | 227 | 190-250 | 54 |
| R: GGGATAGAGCAAGCAGCGTT | ||||||
| Fsp4.7 | F: ACACACACACAGGCACAAG | (AC)10 | Contig 4/: 1510008-1510245 | 238 | 201-260 | 54 |
| R: TGAGTAAGGTAGGCTGGGTCT | ||||||
| Fsp5.3 | F: TACACTCCCGTTCCTGGCTA | (TC)15 | Contig 4/: 161679-161833 | 155 | 90-147 | 54 |
| R: CCAAATCTCAACCGTCGTGG | ||||||
| Fsp5.4 | F: TGAATCTGACTGTGGCTGGAG | (TG)11 | Contig 4/: 330399-330579 | 181 | 160-210 | 54 |
| R: ATGGTTCGCCTTGGCTCTGA | ||||||
| Fsp5.8 | F: CAGTTAGGTTCCACTCCAGGT | (ACT)14 | Contig 4/: 216245-216574 | 330 | 260-375 | 54 |
| R: TGGGCGGTCATTGTGTAGGT | ||||||
| Fsp6.3 | F: AGCACGAGGAACGAGTCTA | (AC)15 | Contig 18/: 1864740-1864874 | 135 | 110-150 | 52 |
| R: CGCTACTTGTTTGTGTCCG | ||||||
| Fsp6.7 | F: TGACCGAGAACTGAAGCCA | (TTC)9 | Contig 18/: 2280272-2280530 | 259 | 250-350 | 52 |
| R: GACGACGAAACCTTTGAAGAG | ||||||
| Fsp7.3 | F: CTCTCATTATGTAGCGACGAC | (TCG)10 | Contig 7/: 808299-808470 | 173 | 150-250 | 52 |
| R: TCGATACCCCTGAGTTCTGTG | ||||||
| Fsp9.6 | F: GCATCCAATGTCTTTCCCAAT | (TG)17 | Contig 5/: 189772--190021 | 250 | 230-275 | 52 |
| R: GACTCGATATTCATCAACACCA | ||||||
| Fsp9.8 | F: TTGGTTTCCCTGCCTTGT | (TG)20 | Contig 5/: 1802083-1802348 | 266 | 240-300 | 52 |
| R: TGTTGGGTCATCTTGGTTTC | ||||||
| Fsp9.9 | F: ACGCCACTCTTCCATACTCAG | (AAC)15 | Contig 5/: 764399-764632 | 234 | 200-280 | 54 |
| R: GTTGACGGTCTTGTGGCATAC | ||||||
| Fsp9.10 | F: TCAACAACACCAACCACCTG | (AGC)9 | Contig 5/: 509965-510211 | 247 | 200-300 | 54 |
| R: TTCGTCCGAGTCGCCTTCTA | ||||||
| Fsp9.15 | F: GCTTGTGCTGGAGTTGACCT | (GAAT)8 | Contig 5/: 168328-168492 | 165 | 123-155 | 52 |
| R: CAGGAGCATAGGAGGAATAC | ||||||
| Fsp10.2 | F: CGAGAAAGGTGAGGAAGGGA | (GGA)8 | Contig 11/: 258852-259033 | 182 | 175-200 | 52 |
| R: TATCTGTGAGGTGTGGCGA |
| SSR位点 SSR locus | 观测等位基因数 Observed number of alleles (Na) | 有效等位基因数 Effective number of alleles (Ne) | 基因多样性指数 Gene diversity index (H) | Shannon指数 Shannon index (I) |
|---|---|---|---|---|
| Fsp1.7 | 5 | 2.8447 | 0.6485 | 1.2162 |
| Fsp1.10 | 15 | 4.0502 | 0.7531 | 1.9836 |
| Fsp1.17 | 5 | 3.6560 | 0.7265 | 1.4019 |
| Fsp2.2 | 6 | 4.8578 | 0.7941 | 1.6700 |
| Fsp2.3 | 5 | 3.3457 | 0.7011 | 1.3637 |
| Fsp2.4 | 5 | 2.1525 | 0.5354 | 0.9979 |
| Fsp3.1 | 4 | 2.3027 | 0.5657 | 1.0762 |
| Fsp3.6 | 3 | 2.7466 | 0.6359 | 1.0521 |
| Fsp3.7 | 5 | 4.4211 | 0.7738 | 1.5415 |
| Fsp4.2 | 4 | 1.9438 | 0.4855 | 0.8826 |
| Fsp4.4 | 4 | 3.5333 | 0.7170 | 1.3187 |
| Fsp4.7 | 4 | 2.5645 | 0.6101 | 1.0413 |
| Fsp5.3 | 6 | 4.3986 | 0.7727 | 1.5779 |
| Fsp5.4 | 5 | 4.2793 | 0.7663 | 1.5211 |
| Fsp5.8 | 6 | 5.1658 | 0.8064 | 1.7170 |
| Fsp6.3 | 4 | 2.5908 | 0.6140 | 1.1040 |
| Fsp6.7 | 7 | 5.2721 | 0.8103 | 1.7429 |
| Fsp7.3 | 5 | 2.7909 | 0.6417 | 1.2545 |
| Fsp9.6 | 5 | 4.0180 | 0.7511 | 1.4874 |
| Fsp9.8 | 4 | 3.1232 | 0.6798 | 1.2192 |
| Fsp9.9 | 8 | 5.7601 | 0.8264 | 1.8905 |
| Fsp9.10 | 7 | 5.3629 | 0.8135 | 1.7745 |
| Fsp9.15 | 4 | 3.1419 | 0.6817 | 1.2351 |
| Fsp10.2 | 6 | 4.5338 | 0.7794 | 1.6062 |
| 平均 Mean | 5.5 | 3.7023 | 0.7038 | 1.4031 |
Table 3 Allelic frequencies and other diversity indices of 24 SSR loci in 96 isolates of Fusarium solani f. sp. pisi
| SSR位点 SSR locus | 观测等位基因数 Observed number of alleles (Na) | 有效等位基因数 Effective number of alleles (Ne) | 基因多样性指数 Gene diversity index (H) | Shannon指数 Shannon index (I) |
|---|---|---|---|---|
| Fsp1.7 | 5 | 2.8447 | 0.6485 | 1.2162 |
| Fsp1.10 | 15 | 4.0502 | 0.7531 | 1.9836 |
| Fsp1.17 | 5 | 3.6560 | 0.7265 | 1.4019 |
| Fsp2.2 | 6 | 4.8578 | 0.7941 | 1.6700 |
| Fsp2.3 | 5 | 3.3457 | 0.7011 | 1.3637 |
| Fsp2.4 | 5 | 2.1525 | 0.5354 | 0.9979 |
| Fsp3.1 | 4 | 2.3027 | 0.5657 | 1.0762 |
| Fsp3.6 | 3 | 2.7466 | 0.6359 | 1.0521 |
| Fsp3.7 | 5 | 4.4211 | 0.7738 | 1.5415 |
| Fsp4.2 | 4 | 1.9438 | 0.4855 | 0.8826 |
| Fsp4.4 | 4 | 3.5333 | 0.7170 | 1.3187 |
| Fsp4.7 | 4 | 2.5645 | 0.6101 | 1.0413 |
| Fsp5.3 | 6 | 4.3986 | 0.7727 | 1.5779 |
| Fsp5.4 | 5 | 4.2793 | 0.7663 | 1.5211 |
| Fsp5.8 | 6 | 5.1658 | 0.8064 | 1.7170 |
| Fsp6.3 | 4 | 2.5908 | 0.6140 | 1.1040 |
| Fsp6.7 | 7 | 5.2721 | 0.8103 | 1.7429 |
| Fsp7.3 | 5 | 2.7909 | 0.6417 | 1.2545 |
| Fsp9.6 | 5 | 4.0180 | 0.7511 | 1.4874 |
| Fsp9.8 | 4 | 3.1232 | 0.6798 | 1.2192 |
| Fsp9.9 | 8 | 5.7601 | 0.8264 | 1.8905 |
| Fsp9.10 | 7 | 5.3629 | 0.8135 | 1.7745 |
| Fsp9.15 | 4 | 3.1419 | 0.6817 | 1.2351 |
| Fsp10.2 | 6 | 4.5338 | 0.7794 | 1.6062 |
| 平均 Mean | 5.5 | 3.7023 | 0.7038 | 1.4031 |
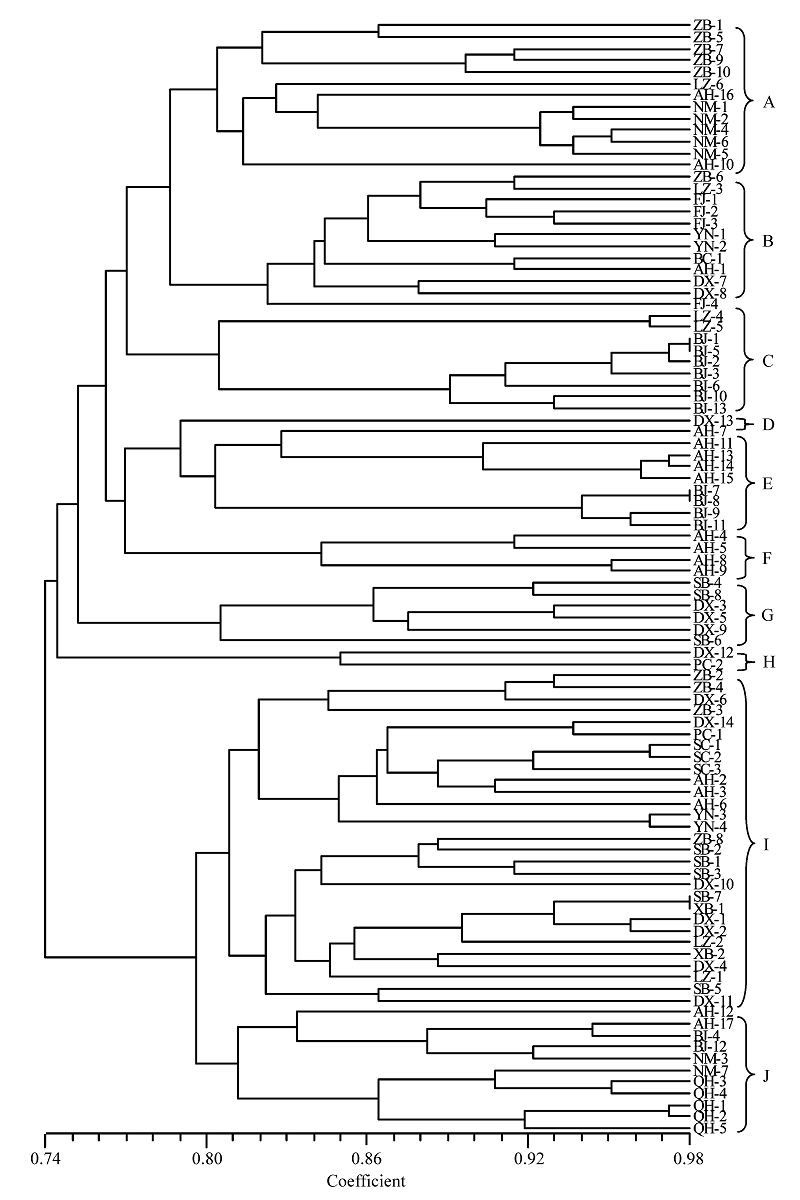
Fig. 1 A UPGMA dendrogram of 96 isolates of Fusarium solani f. sp. pisi based on genetic similarity. A-J indicate different genotype clusters, and isolate codes see Table 1.
| 群体 Population | 菌株数 No. of isolates | 观测等位基因数 Observed number of alleles (Na) | 有效等位基因数 Effective number of alleles (Ne) | 基因多样性指数 Gene diversity Index (H) | Shannon指数 Shannon index (I) |
|---|---|---|---|---|---|
| 河北 Hebei | 10 | 2.8750 | 2.5377 | 0.5925 | 0.9599 |
| 宁夏 Ningxia | 12 | 3.3750 | 2.5005 | 0.5593 | 0.9905 |
| 甘肃 Gansu | 20 | 3.9583 | 2.9460 | 0.6353 | 1.1590 |
| 安徽 Anhui | 17 | 3.8750 | 2.7750 | 0.6040 | 1.1058 |
| 北京 Beijing | 13 | 2.8750 | 2.2937 | 0.5262 | 0.8820 |
| 内蒙古 Inner Mongolia | 7 | 2.1250 | 1.7474 | 0.4007 | 0.6140 |
| 青海 Qinghai | 5 | 1.5000 | 1.3867 | 0.2144 | 0.3091 |
Table 4 Genetic diversity in seven populations of Fusarium solani f. sp. pisi
| 群体 Population | 菌株数 No. of isolates | 观测等位基因数 Observed number of alleles (Na) | 有效等位基因数 Effective number of alleles (Ne) | 基因多样性指数 Gene diversity Index (H) | Shannon指数 Shannon index (I) |
|---|---|---|---|---|---|
| 河北 Hebei | 10 | 2.8750 | 2.5377 | 0.5925 | 0.9599 |
| 宁夏 Ningxia | 12 | 3.3750 | 2.5005 | 0.5593 | 0.9905 |
| 甘肃 Gansu | 20 | 3.9583 | 2.9460 | 0.6353 | 1.1590 |
| 安徽 Anhui | 17 | 3.8750 | 2.7750 | 0.6040 | 1.1058 |
| 北京 Beijing | 13 | 2.8750 | 2.2937 | 0.5262 | 0.8820 |
| 内蒙古 Inner Mongolia | 7 | 2.1250 | 1.7474 | 0.4007 | 0.6140 |
| 青海 Qinghai | 5 | 1.5000 | 1.3867 | 0.2144 | 0.3091 |
| 变异来源 Source of variation | 自由度 df | 平方和 Sum of squares | 方差分量 Variance component | 变异率 Percentage of variation (%) | FST | P |
|---|---|---|---|---|---|---|
| 群体间 Among populations | 6 | 134.733 | 1.25595 | 13.86 | 0.13865 | <0.001 |
| 群体内 Within population | 77 | 600.814 | 7.80278 | 86.14 | - | <0.001 |
| 合计 Total | 83 | 735.548 | 9.05873 | 100 | - | - |
Table 5 AMOVA analysis of genetic variation in seven Fusarium solani f. sp. pisi populations from different geographical regions based on SSR markers
| 变异来源 Source of variation | 自由度 df | 平方和 Sum of squares | 方差分量 Variance component | 变异率 Percentage of variation (%) | FST | P |
|---|---|---|---|---|---|---|
| 群体间 Among populations | 6 | 134.733 | 1.25595 | 13.86 | 0.13865 | <0.001 |
| 群体内 Within population | 77 | 600.814 | 7.80278 | 86.14 | - | <0.001 |
| 合计 Total | 83 | 735.548 | 9.05873 | 100 | - | - |
| 群体 Population | 河北 Hebei | 宁夏 Ningxia | 甘肃 Gansu | 安徽 Anhui | 北京 Beijing | 内蒙古 Inner Mongolia | 青海Qinghai |
|---|---|---|---|---|---|---|---|
| 河北 Heibei | **** | 0.6780 | 0.7550 | 0.6943 | 0.5416 | 0.5859 | 0.5435 |
| 宁夏 Ningxia | 0.3886 | **** | 0.8536 | 0.6432 | 0.4899 | 0.4528 | 0.6226 |
| 甘肃 Gansu | 0.2810 | 0.1583 | **** | 0.7515 | 0.5628 | 0.5260 | 0.5931 |
| 安徽 Anhui | 0.3648 | 0.4413 | 0.2857 | **** | 0.5917 | 0.5285 | 0.4808 |
| 北京 Beijing | 0.6133 | 0.7135 | 0.5748 | 0.5248 | **** | 0.6442 | 0.4073 |
| 内蒙古 Inner Mongolia | 0.5346 | 0.7922 | 0.6425 | 0.6377 | 0.4398 | **** | 0.4802 |
| 青海 Qinghai | 0.6097 | 0.4738 | 0.5224 | 0.7324 | 0.8983 | 0.7336 | **** |
Table 6 Genetic similarity coefficient (above diagonal) and genetic distance (below diagonal) among seven populations of Fusarium solani f. sp. pisi
| 群体 Population | 河北 Hebei | 宁夏 Ningxia | 甘肃 Gansu | 安徽 Anhui | 北京 Beijing | 内蒙古 Inner Mongolia | 青海Qinghai |
|---|---|---|---|---|---|---|---|
| 河北 Heibei | **** | 0.6780 | 0.7550 | 0.6943 | 0.5416 | 0.5859 | 0.5435 |
| 宁夏 Ningxia | 0.3886 | **** | 0.8536 | 0.6432 | 0.4899 | 0.4528 | 0.6226 |
| 甘肃 Gansu | 0.2810 | 0.1583 | **** | 0.7515 | 0.5628 | 0.5260 | 0.5931 |
| 安徽 Anhui | 0.3648 | 0.4413 | 0.2857 | **** | 0.5917 | 0.5285 | 0.4808 |
| 北京 Beijing | 0.6133 | 0.7135 | 0.5748 | 0.5248 | **** | 0.6442 | 0.4073 |
| 内蒙古 Inner Mongolia | 0.5346 | 0.7922 | 0.6425 | 0.6377 | 0.4398 | **** | 0.4802 |
| 青海 Qinghai | 0.6097 | 0.4738 | 0.5224 | 0.7324 | 0.8983 | 0.7336 | **** |
| [1] | Alves-Santos FM, Benito EP, Eslava AP, Díaz-Mínguez JM (1999) Genetic diversity of Fusarium oxysporum strains from common bean fields in Spain. Applied and Environmental Microbiology, 65, 3335-3340. |
| [2] | Chen QH (陈庆河), Weng QY (翁启勇), He YX (何玉仙), Zhao J (赵健) (2004) Pathogens and pathogenicity of root disease of peas in Fujian Province. Fujian Journal of Agricultural Sciences (福建农业学报), 19, 28-31. (in Chinese with English abstract) |
| [3] | Chen XM, Line RF, Leung H (1993) Relationship between virulence variation and DNA polymorphism in Puccinia striiformis. Phytopathology, 83, 1489-1497. |
| [4] |
Coleman JJ, Rounsley SD, Rodriguez-Carres M, Kuo A, Wasmann CC, Grimwood J, Schmutz J, Taga M, White GJ, Zhou SG, Schwartz DC, Freitag M, Ma LJ, Danchin EGJ, Henrissat B, Coutinho PM, Nelson DR, Straney D, Napoli CA, Barker BM, Gribskov M, Rep M, Kroken S, Molnàr I, Rensing C, Kennell JC, Zamora J, Farman ML, Selker EU, Salamov A, Shapiro H, Shapiro J, Lindquist E, Lamers C, Grigoriev IV, Geiser DM, Covert SF, Temporini E, VanEtten HD (2009) The genome of Nectria haematococca: contribution of supernumerary chromosomes to gene expansion. PLoS Genetics, 5, e1000618.
DOI URL PMID |
| [5] | Diao ZM (刁治民) (1996) Studies on the species and pathogenicity of root disease of peas in Qinghai province. Journal of Microbiology (微生物学杂志), 16(1), 31-34. (in Chinese with English abstract) |
| [6] | Etebu E, Osborn AM (2010) Molecular quantification of the pea footrot disease pathogen (Nectria haematococca) in agricultural soils. Phytoparasitica, 38, 447-454. |
| [7] | Etebu E, Osborn AM (2011) Pea footrot disease depends on the combination of pathogenicity genes in Nectria haematococca. Asian Journal of Agricultural Sciences, 3, 156-161. |
| [8] |
Excoffier L, Laval S, Schneider G (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online, 1, 47-50.
URL PMID |
| [9] | Feng J, Hwang R, Chang KF, Conner RL, Hwang SF, Strelkov SE, Gossen BD, McLaren DL, Xue AG (2011) Identification of microsatellite markers linked to quantitative trait loci controlling resistance to Fusarium root rot in field pea. Canadian Journal of Plant Science, 91, 199-204. |
| [10] | Feng J, Hwang R, Chang KF, Hwang SF, Strelkov SE, Gossen BD, Conner RL, Turnbu GD (2010) Genetic variation in Fusarium avenaceum causing root rot on field pea. Plant Pathology, 59, 845-852. |
| [11] | Fiers M, Edel-Hermann V, Héraud C, Gautheron N, Chatot C, Le Hingrat Y, Bouchek-Mechiche K, Steinberg C (2011) Genetic diversity of Rhizoctonia solani associated with potato tubers in France. Mycologia, 103, 1230-1244. |
| [12] |
Graham PH, Vance CP (2003) Legumes: importance and constraints to greater use. Plant Physiology, 131, 872-877.
DOI URL PMID |
| [13] | Han YN, Liu XG, Benny U, Kistler HC, VanEtten HD (2001) Genes determining pathogenicity to pea are clustered on a supernumerary chromosome in the fungal plant pathogen Nectria haematococca. The Plant Journal, 25, 305-314. |
| [14] | Hasanzade F, Rastegar MF, Jafarpour B, Kermani M (2008) Identification of Fusarium solani f. sp. Pisi: the cause of root rot in chickpea and assessment of its genetic diversity using AFLP in Northeast Iran. Research Journal of Biological Sciences, 3, 737-741. |
| [15] | Hill AL, Reeves PA, Larson RL, Fenwick AL, Hanson LE, Panell L (2011) Genetic variability among isolates of Fusarium oxysporum from sugar beet. Plant Pathology, 60, 496-505. |
| [16] | Infantino A, Kharrat M, Riccioni L, Coyne CJ, McPhee KE, Grünwald NJ (2006) Screening techniques and sources of resistance to root diseases in cool season food legumes. Euphytica, 147, 201-221. |
| [17] | Kalendar R, Lee D, Schulman AH (2009) FastPCR software for PCR primer and probe design and repeat search. Genes, Genomes and Genomics, 3, 1-14. |
| [18] |
Kimura M, Crow JF (1964) The number of alleles that can be maintained in a finite population. Genetics, 49, 725-738.
URL PMID |
| [19] | Lewontin RC (1972) The apportionment of human diversity. Evolutionary Biology, 6, 381-398. |
| [20] |
McDonald BA (1997) The population genetics of fungi: tools and techniques. Phytopathology, 87, 448-453.
DOI URL PMID |
| [21] |
McDonald BA, Lindem C (2002) Pathogen population genetics, evolutionary potential, and durable resistance. Annual Review of Phytopathology, 40, 349-379.
DOI URL PMID |
| [22] | Mwang’ombe AW, Kipsumbai PK, Kiprop EK, Olubayo FM, Ochieng JW (2008) Analysis of Kenyan isolates of Fusarium solani f. sp. phaseoli from common bean using colony characteristics, pathogenicity and microsatellite DNA. African Journal of Biotechnology, 7, 1662-1671. |
| [23] | Nei M (1972) Genetic distance between populations. The American Naturalist, 106, 283-292. |
| [24] | Nei M, Li WH (1979) Mathematical model for studying genetic variation in terms of restriction endonucleases. Proceedings of the National Academy of Sciences, USA, 76, 5269-5273. |
| [25] | Ren X (任旭), Zhu ZD (朱振东), Li HJ (李洪杰), Duan CX (段灿星), Wang XM (王晓鸣) (2012) SSR marker development and analysis of genetic diversity of Fusarium verticillioides isolated from maize in China. Scientia Agricultura Sinica (中国农业科学), 45, 52-66. (in Chinese with English abstract) |
| [26] |
Rep M, Kistler HC (2010) The genomic organization of plant pathogenicity in Fusarium species. Current Opinion in Plant Biology, 13, 420-426.
URL PMID |
| [27] | Rohlf FJ (1997) NTSYSpc Version 2.1. Numerical Taxonomy and Multivariate Analysis System. Exeter Software, Setauket, New York. |
| [28] | Rush CM, Kraft JM (1986) Effects of inoculum density and placement on Fusarium root rot of peas. Phytopathology, 76, 1325-1329. |
| [29] | Skovgaard K, Bødker L, Rosendahl S (2002) Population structure and pathogenicity of members of the Fusarium oxysporum complex isolated from soil and root necrosis of pea (Pisum sativum L.). FEMS Microbiology Ecology, 42, 367-374. |
| [30] | Tang DZ (唐德志), He SQ (何苏琴), Li YQ (李玉奇), Zhu RS (朱润身) (1993) Studies on the species and pathogenicity of root disease of peas in Gansu Province. Acta Agriculturae Boreali-Occidentalis Sinica (西北农业学报), (2), 37-39. (in Chinese with English abstract) |
| [31] |
Temporini ED, VanEtten HD (2002) Distribution of the pea pathogenicity (PEP) genes in the fungus Nectria haematococca mating population VI. Current Genetics, 41, 107-114.
URL PMID |
| [32] | VanEtten HD (1978) Identification of additional habitats of Nectria haematococca mating population. VI. Phytopathology, 68, 1552-1556. |
| [33] |
VanEtten HD, Funnel-Baerg D, Wasmann C, McCluskey K (1994) Location of pathogenicity genes on dispensable chromosomes in Nectria haematococca MPVI. Antonie van Leeuwenhoek, 65, 263-267.
URL PMID |
| [34] | VanEtten HD, Matthews PS, Tegtmeier KJ, Dietert MF, Stein JI (1980) The association of pisatin tolerance and demethylation with virulence on pea in Nectria haematococca. Physiological Plant Pathology, 16, 257-268. |
| [35] | Wang KC (王宽仓), Zhang ZS (张宗山), Chen JN (陈渐宁), Fan ZQ (樊仲庆), Niu BS (牛宝山), Zhao M (赵明), Xie CJ (谢成君) (1995) Studies on occurrence regulation and the integrated prevention and control techniques of Fusarium root rot of pea. Ningxia Journal of Agriculture and Forestry Science and Technology (宁夏农林科技), (5), 1-6. (in Chinese with English abstract) |
| [36] | Wang MC (王梅春), Lian RF (连荣芳), Mo JP (墨金萍), Wang SH (王思慧) (2008) Research of the pea root rot and resistant breeding in Gansu Province. Rain Fed Crops (杂粮作物), 28, 272-273. (in Chinese with English abstract) |
| [37] | Wu KJ (伍克俊), Xie ZT (谢正团), Li XJ (李秀君) (1992) Study on the pathogens of root rot of pea in the central region of Gansu Province. Journal of Gansu Agricultural University (甘肃农业大学学报), 27, 225-231. (in Chinese with English abstract) |
| [38] | Xiang N (向妮), Duan CX (段灿星), Xiao YN (肖炎农), Wang XM (王晓鸣), Zhu ZD (朱振东) (2012) Identification of the pathogen causing Fusarium root rot of pea and diversity of pathogenicity genes. Scientia Agricultura Sinica (中国农业科学), 45, 2838-2847. (in Chinese with English abstract) |
| [39] | Xu JJ (徐静静), Lin Y (蔺宇), Zhu ZD (朱振东) (2008) Development and applications of SSR markers in plant pathogen. Plant Protection (植物保护), 34, 14-21. (in Chinese with English abstract) |
| [40] | Yu TF (俞大绂) (1955) A preliminary list of Fusaria in China. Acta Phytopathologica Sinica (植物病理学报), 1(1), 1-18. (in Chinese with English abstract) |
| [41] | Zong XX (宗绪晓), Guan JP (关建平), Wang SM (王述民), Liu QC (刘庆昌) (2008) Genetic diversity among Chinese pea (Pisum sativum L.) landraces revealed by SSR markers. Acta Agronomica Sinica (作物学报), 34, 1330-1338. (in Chinese with English abstract) |
| [1] | Weng Zhuoxian, Huang Jiaqiong, Zhang Shihao, Yu Kaichun, Zhong Fusheng, Huang Xunhe, Zhang Bin. Genetic diversity and population structure of black-bone chickens in China revealed by mitochondrial COI gene sequences [J]. Biodiv Sci, 2019, 27(6): 667-676. |
| [2] | Xingtong Wu, Lu Chen, Minqiu Wang, Yuan Zhang, Xueying Lin, Xinyu Li, Hong Zhou, Yafeng Wen. Population structure and genetic divergence in Firmiana danxiaensis [J]. Biodiv Sci, 2018, 26(11): 1168-1179. |
| [3] | Shaoshuai Yu, Qicong Xu, Caili Lin, Shengjie Wang, Guozhong Tian. Genetic diversity of phytoplasmas: research status and prospects [J]. Biodiv Sci, 2016, 24(2): 205-215. |
| [4] | Gangbiao Xu,Yan Liang,Yan Jiang,Xiongsheng Liu,Shangli Hu,Yufei Xiao,Bobo Hao. Genetic diversity and population structure of Bretschneidera sinensis, an endangered species [J]. Biodiv Sci, 2013, 21(6): 723-731. |
| [5] | Bo Zhou, Haidong Jiang, Xiuxin Zhang, Jingqi Xue, Yantong Shi. Morphological diversity of some introduced tree peony cultivars [J]. Biodiv Sci, 2011, 19(5): 543-550. |
| [6] | Biyun Chen, Qiong Hu, Christina Dixelius, Guoqing Li, Xiaoming Wu. Genetic diversity in Sclerotinia sclerotiorum assessed with SRAP markers [J]. Biodiv Sci, 2010, 18(5): 509-515. |
| [7] | Changzhong Wang, Zhong Li, Hongwei Liang, Guangfu Hu, Qinchao Wu, Guiwei Zou, Xiangzhong Luo. Genetic diversity in fourProcambarus clarkii populations in the lower reaches of the Yangtze River [J]. Biodiv Sci, 2009, 17(5): 518-523. |
| [8] | Zhengfeng Wang, Xuejun Ge. Not only genetic diversity: advances in plant conservation genetics [J]. Biodiv Sci, 2009, 17(4): 330-339. |
| [9] | Yinghui Ling, Yuejiao Cheng, Yanping Wang, Weijun Guan, Jianlin Han, Baoling Fu, Qianjun Zhao, Xiaohong He, Yabin Pu, Yuehui Ma. Genetic diversity of 23 Chinese indigenous horse breeds revealed by microsatellite markers [J]. Biodiv Sci, 2009, 17(3): 240-247. |
| [10] | Hongwei Liang, Zhong Li, Xiangzhong Luo, Changzhong Wang, Guangfu Hu, Guiwei Zou, Yongquan Yang. Genetic diversity based on microsatellite markers in five Nile tilapia strains [J]. Biodiv Sci, 2009, 17(1): 82-87. |
| [11] | Yanlai Ding, Tuanjie Zhao, Junyi Gai. Genetic diversity and ecological differentiation of Chinese annual wild soybean (Glycine soja) [J]. Biodiv Sci, 2008, 16(2): 133-142. |
| [12] | Xueying Lu, Daoyuan Zhang, Wenbao Ma. Genetic variation and clonal diversity in fragmented populations of the desert plant Eremosparton songoricum based on ISSR markers [J]. Biodiv Sci, 2007, 15(3): 282-291. |
| [13] | Yongfa Luo, Zhigang Wang, Jiaqi Li, Guixiang Zhang, Yaosheng Chen, Yong Liang, Fuqing Yu, Weitao Song, Zifu Zhang . Genetic variation and genetic relationship among 13 Chinese and intro-duced cattle breeds using microsatellite DNA markers [J]. Biodiv Sci, 2006, 14(6): 498-507. |
| [14] | Huifang Wu, Zuozhou Li, Hongwen Huang. Genetic differentiation among natural populations of Gastrodia elata (Orchidaceae) in Hubei and germplasm assessment of the cultivated populations [J]. Biodiv Sci, 2006, 14(4): 315-326. |
| [15] | Yingzhe Xia, Yan Sheng, Yiyu Chen. DNA sequence variation in the mitochondrial control region of lenok (Brachymystax lenok) populations in China [J]. Biodiv Sci, 2006, 14(1): 48-54. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2022 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn ![]()