



Biodiv Sci ›› 2026, Vol. 34 ›› Issue (1): 25210. DOI: 10.17520/biods.2025210 cstr: 32101.14.biods.2025210
• Special Feature: Methods for Ecological Data Analysis • Previous Articles Next Articles
Yafei Shi1,*( )(
)( ), Furong Niu2, Xiaomin Huang3, Xing Hong1, Xiangwen Gong4, Yanli Wang2, Dong Lin1, Xiaoni Liu1
), Furong Niu2, Xiaomin Huang3, Xing Hong1, Xiangwen Gong4, Yanli Wang2, Dong Lin1, Xiaoni Liu1
Received:2025-06-06
Accepted:2025-09-05
Online:2026-01-20
Published:2026-01-21
Contact:
Yafei Shi
Supported by:Yafei Shi, Furong Niu, Xiaomin Huang, Xing Hong, Xiangwen Gong, Yanli Wang, Dong Lin, Xiaoni Liu. Interpretable machine learning and its applications in ecology[J]. Biodiv Sci, 2026, 34(1): 25210.
| 解释需求 Interpretation domain | 核心目标 Core objective | 解释类型 Interpretation type |
|---|---|---|
| 自变量贡献(重要)度 Predictor importance assessment | 哪些自变量更重要? Which predictors contribute most significantly to the model’s output? | 全局解释 Global interpretability |
| 自变量与因变量的关系 Predictor-response relationships | 某个自变量如何影响因变量? How does a given predictor influence the response variable across the dataset? | 全局解释 Global interpretability |
| 单个案例解释 Case-level interpretation | 为何对某一案例有如此的预测结果? What explains the prediction outcome for a specific observation or case? | 局部解释 Local interpretability |
| 生态学意义提炼 Ecological synthesis and insight | 模型结果揭示了什么生态学机制或意义? What ecological mechanisms or implications are inferred from the model outputs? | 抽象提炼、概念集成 Abstract conceptualization and theoretical framing |
Table 1 Main contents of ecological interpretation
| 解释需求 Interpretation domain | 核心目标 Core objective | 解释类型 Interpretation type |
|---|---|---|
| 自变量贡献(重要)度 Predictor importance assessment | 哪些自变量更重要? Which predictors contribute most significantly to the model’s output? | 全局解释 Global interpretability |
| 自变量与因变量的关系 Predictor-response relationships | 某个自变量如何影响因变量? How does a given predictor influence the response variable across the dataset? | 全局解释 Global interpretability |
| 单个案例解释 Case-level interpretation | 为何对某一案例有如此的预测结果? What explains the prediction outcome for a specific observation or case? | 局部解释 Local interpretability |
| 生态学意义提炼 Ecological synthesis and insight | 模型结果揭示了什么生态学机制或意义? What ecological mechanisms or implications are inferred from the model outputs? | 抽象提炼、概念集成 Abstract conceptualization and theoretical framing |
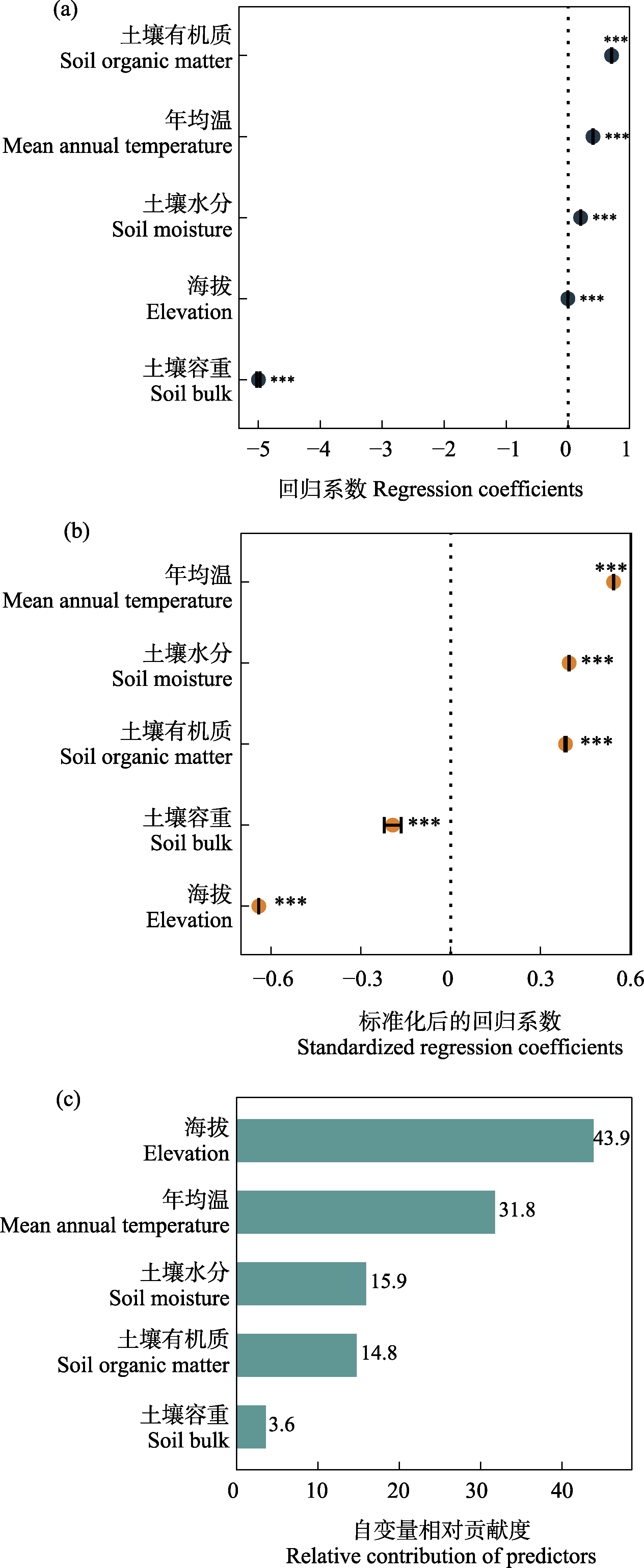
Fig. 1 Interpretation of linear regression model results. (a) Linear regression coefficients; (b) Standardized linear regression coefficients; (c) Variable importance ranking based on variation partitioning and hierarchical partitioning. *** P < 0.001.
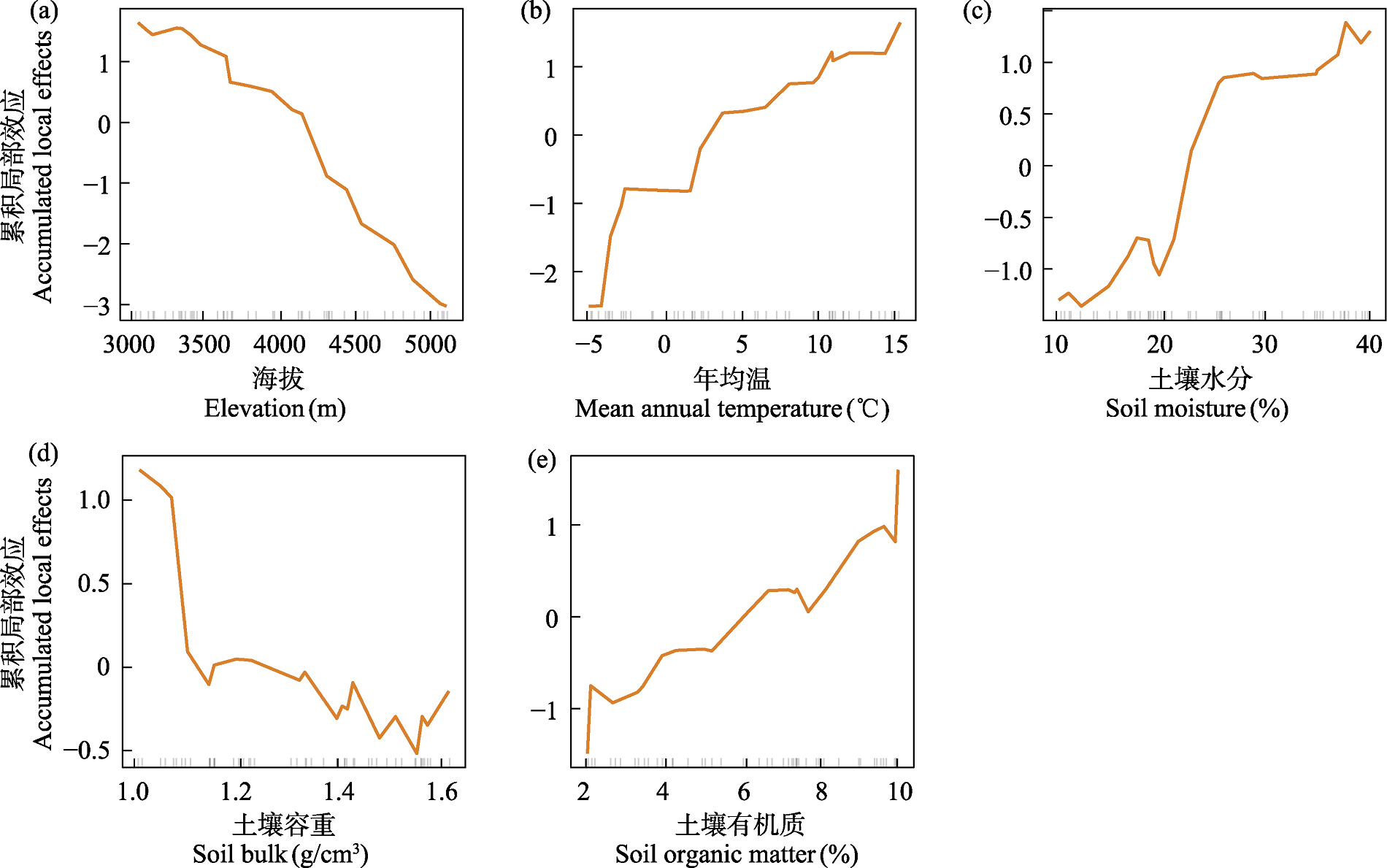
Fig. 5 Accumulated local effects plot (ALE) based on the random forest model. The small vertical ticks on the x-axis indicate the distribution of the variable in the dataset.
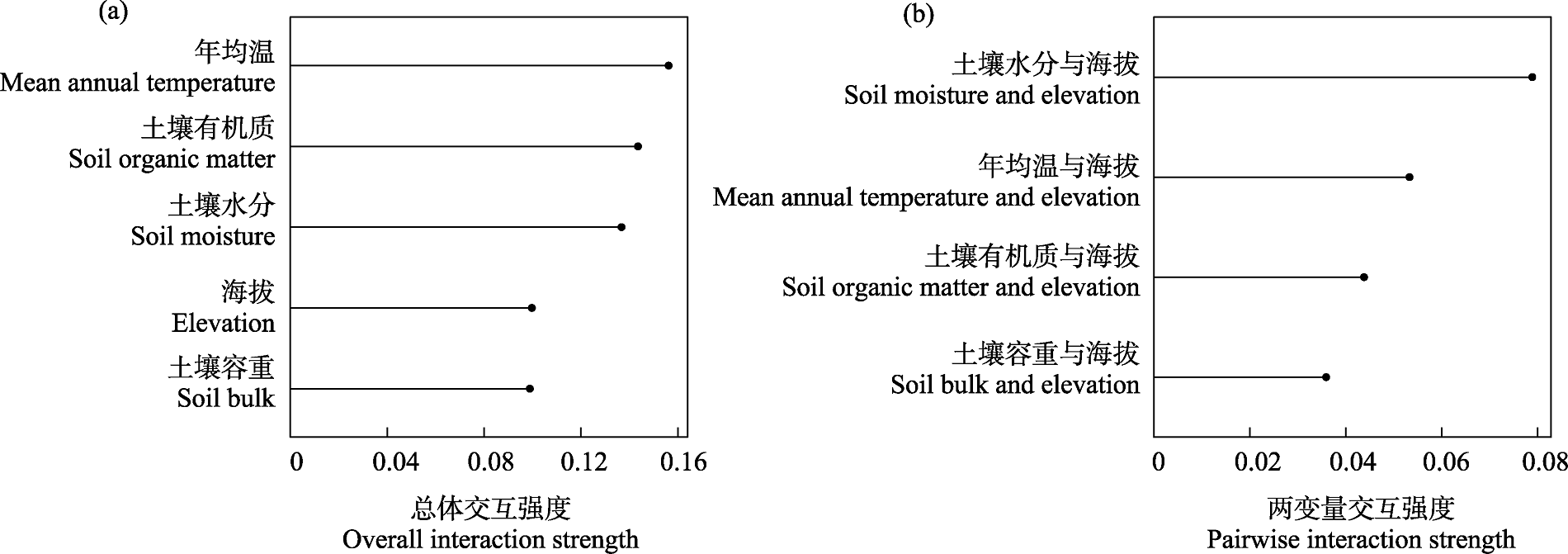
Fig. 6 Feature interactions based on H-statistic. (a) The overall interaction strength between a single variable and all other variables; (b) The pairwise interaction strength between variables.
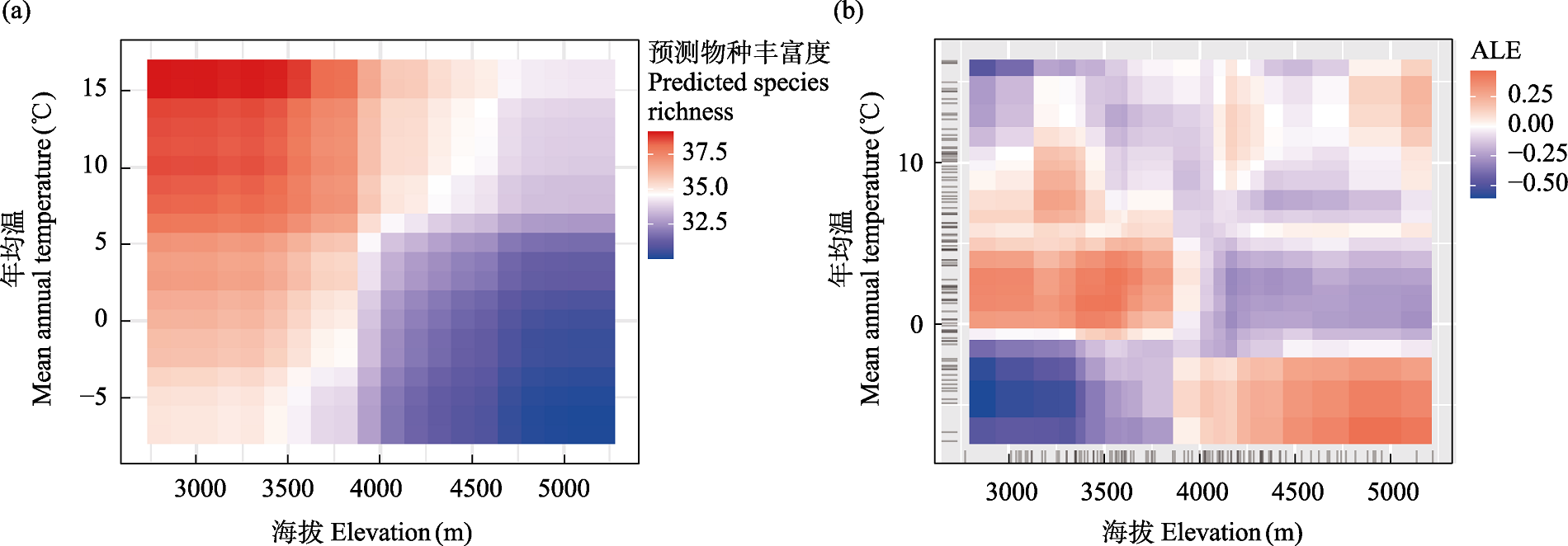
Fig. 7 Two-dimensional interaction plots of elevation and mean annual temperature. (a) Partial dependence plot; (b) Accumulated local effects plot (ALE).
| 方法 Methods | 适合模型 Applicable models | 是否模型无关 Whether it is model-agnostic | 解释层级 Interpretation level | 优点 Advantages | 局限性/注意事项 Limitations/notes |
|---|---|---|---|---|---|
| 回归系数 Regression coefficients | 线性模型(白盒) Linear models (white-box) | 否 No | 全局 Global | 直观、易于理解 Intuitive and easy to interpret | 依赖模型假设; 适用于线性关系 Dependent on model assumptions; suitable only for linear relationships |
| 决策树 Decision tree | 决策树类模型(白盒) Decision tree models (white-box) | 否 No | 全局 Global | 可视化清晰, 展示变量优先级 Clearly visualizes structure; reveals variable hierarchy | 易过拟合, 稳定性差 Overfitting-prone and unstable |
| 置换特征重要性 Permutation importance | 黑盒模型 Black-box models | 是 Yes | 全局 Global | 简单直观, 适用于多数模型 Simple and widely applicable; supports many models | 易受变量相关性影响, 结果不稳定 Sensitive to variable correlation; results may be unstable |
| Gini重要性 Gini importance | 随机森林等树模型 Tree-based models such as random forest | 否 No | 全局 Global | 计算快速、内置于模型 Fast computation; embedded in model | 偏向取值范围大的变量, 不适用于所有模型 Biased toward variables with large ranges; lacks generalizability |
| 部分依赖图 Partial dependence plots (PDP) | 黑盒/白盒模型 Black-/white-box models | 是 Yes | 全局 Global | 图形表达直观, 适合展示非线性关系 Visual and intuitive; useful for nonlinear patterns | 假设变量独立, 存在不现实组合风险 Assumes feature independence; may generate unrealistic combinations |
| 累积局部效应图 Accumulated local effects (ALE) | 黑盒/白盒模型 Black-/white-box models | 是 Yes | 全局 Global | 不依赖变量独立性假设, 解释更稳定 Free from variable independence assumptions, yielding more robust interpretations | Y轴是特征变化的局部平均效应值, 不易直观理解 Y-axis shows local average effect, which lacks intuitive clarity |
| H-statistic交互分析 H-statistic interaction analysis | 黑盒/白盒模型 Black-/white-box models | 是 Yes | 全局 Global | 可量化变量交互强度, 补充单变量解释的不足 Quantifies feature interaction strength; supplements univariate interpretations | 解释难度略高, 计算复杂度大 Interpretation complexity increases; relatively high computational cost |
| 全局代理模型 Global surrogate models | 黑盒模型 Black-box models | 是 Yes | 全局 Global | 简化复杂模型, 借助可解释模型进行间接解释 Approximates complex models with simpler ones for interpretation | 逼近可能不完全, 解释不一 定准确 Approximation may be incomplete; explanation only partially faithful |
| 局部模型无关解释 Local interpretable model-agnostic explanations | 黑盒模型 Black-box models | 是 Yes | 局部 Local | 适合个体预测的解释 Suitable for explaining individual predictions | 局部解释不稳定, 依赖邻域构建 Unstable local explanations; neighborhood-dependent |
| Shapley加性解释 Shapley additive explanations | 黑盒/白盒模型 Black-/white-box models | 是 Yes | 全局/局部 Global/Local | 理论稳健、解释公平性强 Theoretically sound; ensures fair feature attribution | 计算复杂度大 High computational cost |
Table 2 Comparative summary of selected interpretable machine learning methods
| 方法 Methods | 适合模型 Applicable models | 是否模型无关 Whether it is model-agnostic | 解释层级 Interpretation level | 优点 Advantages | 局限性/注意事项 Limitations/notes |
|---|---|---|---|---|---|
| 回归系数 Regression coefficients | 线性模型(白盒) Linear models (white-box) | 否 No | 全局 Global | 直观、易于理解 Intuitive and easy to interpret | 依赖模型假设; 适用于线性关系 Dependent on model assumptions; suitable only for linear relationships |
| 决策树 Decision tree | 决策树类模型(白盒) Decision tree models (white-box) | 否 No | 全局 Global | 可视化清晰, 展示变量优先级 Clearly visualizes structure; reveals variable hierarchy | 易过拟合, 稳定性差 Overfitting-prone and unstable |
| 置换特征重要性 Permutation importance | 黑盒模型 Black-box models | 是 Yes | 全局 Global | 简单直观, 适用于多数模型 Simple and widely applicable; supports many models | 易受变量相关性影响, 结果不稳定 Sensitive to variable correlation; results may be unstable |
| Gini重要性 Gini importance | 随机森林等树模型 Tree-based models such as random forest | 否 No | 全局 Global | 计算快速、内置于模型 Fast computation; embedded in model | 偏向取值范围大的变量, 不适用于所有模型 Biased toward variables with large ranges; lacks generalizability |
| 部分依赖图 Partial dependence plots (PDP) | 黑盒/白盒模型 Black-/white-box models | 是 Yes | 全局 Global | 图形表达直观, 适合展示非线性关系 Visual and intuitive; useful for nonlinear patterns | 假设变量独立, 存在不现实组合风险 Assumes feature independence; may generate unrealistic combinations |
| 累积局部效应图 Accumulated local effects (ALE) | 黑盒/白盒模型 Black-/white-box models | 是 Yes | 全局 Global | 不依赖变量独立性假设, 解释更稳定 Free from variable independence assumptions, yielding more robust interpretations | Y轴是特征变化的局部平均效应值, 不易直观理解 Y-axis shows local average effect, which lacks intuitive clarity |
| H-statistic交互分析 H-statistic interaction analysis | 黑盒/白盒模型 Black-/white-box models | 是 Yes | 全局 Global | 可量化变量交互强度, 补充单变量解释的不足 Quantifies feature interaction strength; supplements univariate interpretations | 解释难度略高, 计算复杂度大 Interpretation complexity increases; relatively high computational cost |
| 全局代理模型 Global surrogate models | 黑盒模型 Black-box models | 是 Yes | 全局 Global | 简化复杂模型, 借助可解释模型进行间接解释 Approximates complex models with simpler ones for interpretation | 逼近可能不完全, 解释不一 定准确 Approximation may be incomplete; explanation only partially faithful |
| 局部模型无关解释 Local interpretable model-agnostic explanations | 黑盒模型 Black-box models | 是 Yes | 局部 Local | 适合个体预测的解释 Suitable for explaining individual predictions | 局部解释不稳定, 依赖邻域构建 Unstable local explanations; neighborhood-dependent |
| Shapley加性解释 Shapley additive explanations | 黑盒/白盒模型 Black-/white-box models | 是 Yes | 全局/局部 Global/Local | 理论稳健、解释公平性强 Theoretically sound; ensures fair feature attribution | 计算复杂度大 High computational cost |
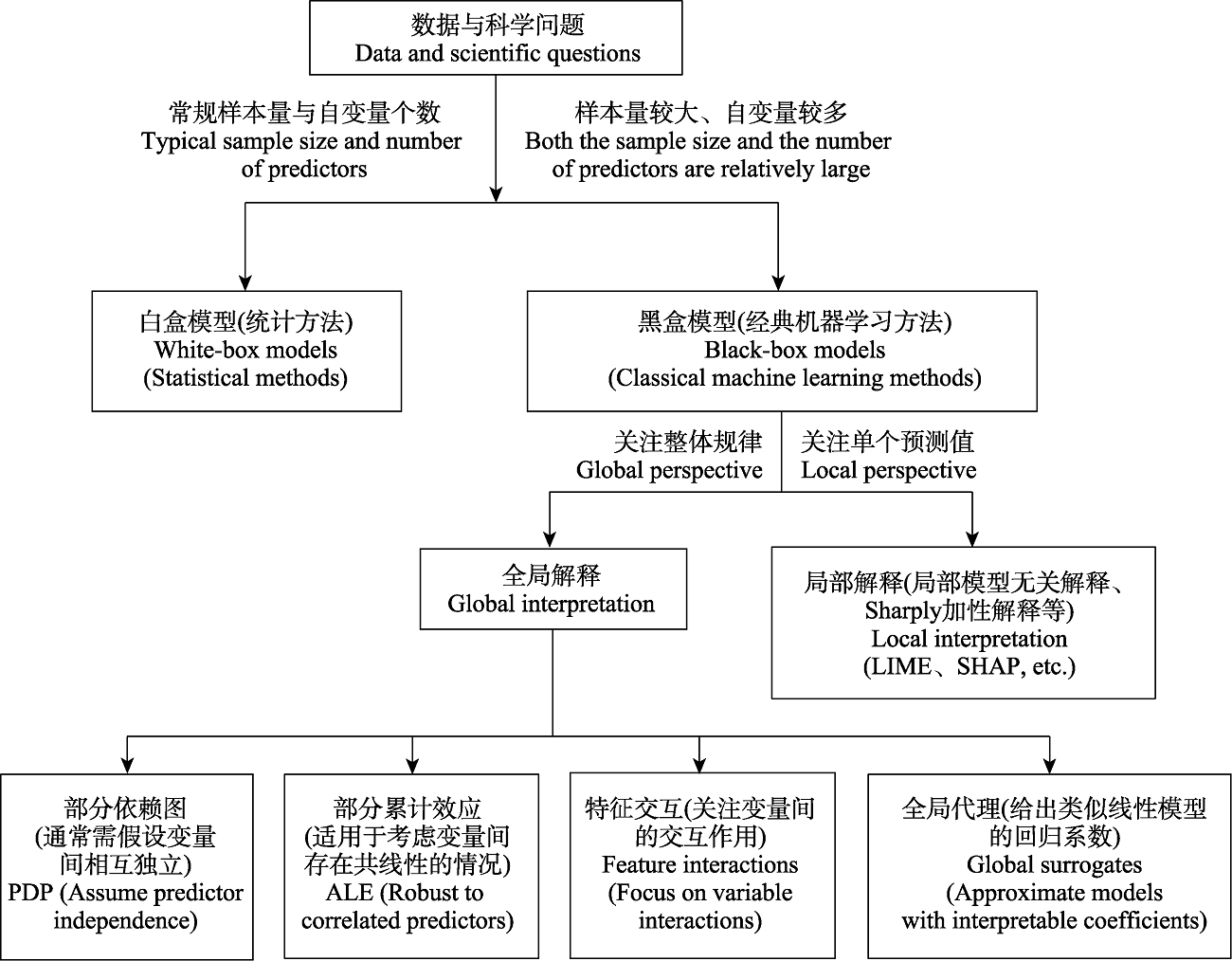
Fig. 11 Recommendations for selecting interpretable machine learning methods. LIME, Local interpretable model-agnostic explanations; SHAP, Shapley additive explanations; PDP, Partial dependence plots; ALE, Accumulated local effects.
| [1] | Apley DW, Zhu JY (2020) Visualizing the effects of predictor variables in black box supervised learning models. Journal of the Royal Statistical Society B: Statistical Methodology, 82, 1059-1086. |
| [2] | Beery S, Cole E, Parker J, Perona P, Winner K (2021) Species distribution modeling for machine learning practitioners: A review. ACM SIGCAS Conference on Computing and Sustainable Societies (COMPASS’21), New York, USA. |
| [3] |
Breiman L (2001) Statistical modeling: The two cultures. Statistical Science, 16, 199-231.
DOI URL |
| [4] | Christoph M (2022) Interpretable Machine Learning: A Guide for Making Black Box Models Explainable, 2nd edn. Independently published. |
| [5] |
Davison CW, Assmann JJ, Normand S, Rahbek C, Morueta-Holme N (2023) Vegetation structure from LiDAR explains the local richness of birds across Denmark. Journal of Animal Ecology, 92, 1332-1344.
DOI PMID |
| [6] |
De’ath G, Fabricius KE (2000) Classification and regression trees: A powerful yet simple technique for ecological data analysis. Ecology, 81, 3178-3192.
DOI URL |
| [7] |
Ding Y, Zang RG (2021) Determinants of aboveground biomass in forests across three climatic zones in China. Forest Ecology Management, 482, 118805.
DOI URL |
| [8] | Friedman JH (2001) Greedy function approximation: A gradient boosting machine. Annals of Statistics, 29, 1189-1232. |
| [9] | Friedman JH, Popescu BE (2008) Predictive learning via rule ensembles. Annals of Applied Statistics, 2, 916-954. |
| [10] |
Gobeyn S, Mouton AM, Cord AF, Kaim A, Volk M, Goethals P (2019) Evolutionary algorithms for species distribution modelling: A review in the context of machine learning. Ecological Modelling, 392, 179-195.
DOI |
| [11] |
Guo QH, Jin SC, Li M, Yang QL, Xu KX, Ju YZ, Zhang J, Xuan J, Liu J, Su YJ, Xu Q, Liu Y (2020) Application of deep learning in ecological resource research: Theories, methods, and challenges. Science China Earth Sciences, 63, 1457-1474.
DOI |
| [12] | Jiang SJ, Sweet L, Blougouras G, Brenning A, Li WT, Reichstein M, Denzler J, SG W, Yu G, Huang FN (2024) How interpretable machine learning can benefit process understanding in the geosciences. Earth’s Future, 12, e2024EF004540. |
| [13] |
Lai JS, Tang J, Li TY, Zhang AY, Mao LF (2024) Evaluating the relative importance of predictors in generalized additive models using the gam.hp R package. Plant Diversity, 46, 542-546.
DOI URL |
| [14] |
Lai JS, Zou Y, Zhang JL, Peres-Neto PR (2022a) Generalizing hierarchical and variation partitioning in multiple regression and canonical analyses using the rdacca.hp R package. Methods in Ecology Evolution, 13, 782-788.
DOI URL |
| [15] |
Lai JS, Zou Y, Zhang S, Zhang XG, Mao LF (2022b) glmm.hp: An R package for computing individual effect of predictors in generalized linear mixed models. Journal of Plant Ecology, 15, 1302-1307.
DOI URL |
| [16] |
Li HJ, Wang B, Niu X, Liang YL, Li JY (2023) Application of machine learning technology in ecology. Chinese Journal of Ecology, 42, 2767-2775. (in Chinese with English abstract)
DOI |
| [李慧杰, 王兵, 牛香, 梁咏亮, 李静尧 (2023) 机器学习技术在生态学中的应用进展. 生态学杂志, 42, 2767-2775.] | |
| [17] |
Liu ZL, Peng CH, Work T, Candau J, DesRochers A, Kneeshaw D (2018) Application of machine-learning methods in forest ecology: Recent progress and future challenges. Environmental Reviews, 26, 339-350.
DOI URL |
| [18] |
Lu SZ, Qiao XG, Zhao LQ, Wang Z, Gao CG, Wang J, Guo K (2020) Basic characteristics of Stipa sareptana var. krylovii communities in China. Chinese Journal of Plant Ecology, 44, 1087-1094. (in Chinese with English Abstract)
DOI URL |
| [陆帅志, 乔鲜果, 赵利清, 王孜, 高趁光, 王静, 郭柯 (2020) 中国西北针茅草原的基本群落特征. 植物生态学报, 44, 1087-1094.] | |
| [19] | Lucas T (2020) A translucent box: Interpretable machine learning in ecology. Ecological Monographs, 90, e01422. |
| [20] |
Lundberg SM, Erion G, Chen H, DeGrave A, Prutkin JM, Nair B, Katz R, Himmelfarb J, Bansal N, Lee SI (2020) From local explanations to global understanding with explainable AI for trees. Nature Machine Intelligence, 2, 56-67.
DOI PMID |
| [21] | Lundberg SM, Lee SI (2017) A unified approach to interpreting model predictions. Conference on Neural Information Processing Systems (NIPS 2017), Long Beach, USA |
| [22] | Mac Aodha O, Gibb R, Barlow KE, Browning E, Firman M, Freeman R, Harder B, Kinsey L, Mead GR, Newson SE (2018) Bat detective—Deep learning tools for bat acoustic signal detection. PLoS Computational Biology, 14, e1005995. |
| [23] | Norberg A, Abrego N, Blanchet FG, Adler FR, Anderson BJ, Anttila J, Araújo MB, Dallas T, Dunson D, Elith J, Foster SD, Fox R, Franklin J, Godsoe W, Guisan A, O’Hara B, Hill NA, Holt RD, Hui FKC, Husby M, Kålås JA, Lehikoinen A, Luoto M, Mod HK, Newell G, Renner I, Roslin T, Soininen J, Thuiller W, Vanhatalo J, Warton D, White M, Zimmermann NE, Gravel D, Ovaskainen O (2019) A comprehensive evaluation of predictive performance of 33 species distribution models at species and community levels. Ecological Monographs, 89, e01370. |
| [24] |
Pichler M, Hartig F (2023) Machine learning and deep learning—A review for ecologists. Methods in Ecology and Evolution, 14, 994-1016.
DOI URL |
| [25] |
Pollock L J, O’Connor LMJ, Mokany K, Rosauer DF, Talluto L, Thuiller W (2020) Protecting biodiversity (in all its complexity): New models and methods. Trends in Ecology & Evolution, 35, 1119-1128.
DOI URL |
| [26] |
Reichstein M, Camps-Valls G, Stevens B, Jung M, Denzler J, Carvalhais N, Prabhat F (2019) Deep learning and process understanding for data-driven Earth system science. Nature, 566, 195-204.
DOI |
| [27] | Ribeiro MT, Singh S, Guestrin C (2016) “Why should I trust you?” Explaining the predictions of any classifier. Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, USA. |
| [28] | Rudin C, Chen CF, Chen Z, Huang HY, Semenova L, Zhong C (2022) Interpretable machine learning: Fundamental principles and 10 grand challenges. Statistic Surveys, 16, 1-85. |
| [29] |
Shamir L, Yerby C, Simpson R, von Benda-Beckmann AM, Tyack P, Samarra F, Miller P, Wallin J (2014) Classification of large acoustic datasets using machine learning and crowdsourcing: Application to whale calls. The Journal of the Acoustical Society of America, 135, 953-962.
DOI URL |
| [30] |
Teng QY, Liu Z, Song YQ, Han K, Lu Y (2022) A survey on the interpretability of deep learning in medical diagnosis. Multimedia Systems, 28, 2335-2355.
DOI |
| [31] | Thampi A (2022) Interpretable AI:Building Explainable Machine Learning Systems. Manning Publications, New York. |
| [32] |
Tuia D, Kellenberger B, Beery S, Costelloe BR, Zuffi S, Risse B, Mathis A, Mathis MW, van Langevelde F, Burghardt T, Kays R, Klinck H, Wikelski M, Couzin ID, van Horn G, Crofoot MC, Stewart CV, Berger WT (2022) Perspectives in machine learning for wildlife conservation. Nature Communications, 13, 792.
DOI PMID |
| [33] |
Wäldchen J, Mäder P (2018) Machine learning for image based species identification. Methods in Ecology and Evolution, 9, 2216-2225.
DOI URL |
| [34] |
Welchowski T, Maloney KO, Mitchell R, Schmid M (2022) Techniques to improve ecological interpretability of black-box machine learning models: Case study on biological health of streams in the United States with gradient boosted trees. Journal of Agricultural, Biological Environmental Statistics, 27, 175-197.
DOI |
| [35] |
Xue YF, Wang TJ, Skidmore AK (2017) Automatic counting of large mammals from very high resolution panchromatic satellite imagery. Remote Sensing, 9, 878.
DOI URL |
| [36] |
Yu QY, Ji WJ, Prihodko L, Ross CW, Anchang JY, Hanan NP (2021) Study becomes insight: Ecological learning from machine learning. Methods in Ecology and Evolution, 12, 2117-2128.
DOI PMID |
| [37] |
Yu SJ, Wang J, Zhang CY, Zhao XH (2022) Impact and mechanism of maintaining biomass stability in a temperate coniferous and broadleaved mixed forest. Chinese Journal of Plant Ecology, 46, 632-641. (in Chinese with English abstract)
DOI URL |
|
[于水今, 王娟, 张春雨, 赵秀海 (2022) 温带针阔混交林生物量稳定性影响机制. 植物生态学报, 46, 632-641.]
DOI |
|
| [38] |
Zhao YF, Zhu WW, Wei PP, Fang P, Zhang XW, Yan NN, Liu WJ, Zhao H, Wu QR (2022) Classification of Zambian grasslands using random forest feature importance selection during the optimal phenological period. Ecological Indicators, 135, 108529.
DOI URL |
| [39] | Zheng LL, Xu JY, Wang XL, Liu BG (2020) Study on relationships between vegetation community distribution and topsoil factors based on random forests in shoaly wetlands of Poyang Lake. Soils, 52, 378-385. (in Chinese with English Abstract) |
| [郑利林, 徐金英, 王晓龙, 刘宝贵 (2020) 基于随机森林方法研究鄱阳湖典型洲滩植被群落分布与表层土壤因子耦合关系. 土壤, 52, 378-385.] |
| [1] | Wenqi Gao, Jingrong Xiang, Yao Zhao, Lingshuang Fan, Yuan Gu, Weihan Shao, Gaojun Li, Guangjun Zhao, Mingbin Chen, Xingwei Cai, Kai Chen. Stream fish community characteristics and their response to land use in Limushan and Jianfengling of Hainan Tropical Rainforest National Park [J]. Biodiv Sci, 2026, 34(2): 25374-. |
| [2] | Shuai Li, Weihua Liu, Yudan Xu, Xiaobo Tian, Houjuan Song, Xiaoting Yue, Lingling Wu, Qing Zhang, Tieliang Shanguan. A dataset on inventory and geographical distributions of vascular plants in Shanxi, China [J]. Biodiv Sci, 2025, 33(7): 24317-. |
| [3] | Qinwen Lin, Na Zhang, Qiang Wang. A dataset on inventory and geographical distribution of halophytes of China [J]. Biodiv Sci, 2025, 33(7): 25030-. |
| [4] | Quanfeng Yang, Yanjie Tang, Haijun Xiao, Ying Wang, Rong Zhang, Fang Ouyang, Shuhua Wei. Cascading effects of plant diversity-grasshoppers-carabids and their impacts on primary productivity across grassland types of Ningxia, China [J]. Biodiv Sci, 2025, 33(6): 25021-. |
| [5] | Song Wei, Cheng Cai, Wang Jiawei, Wu Jihua. Soil microbes regulate the relationships between plant diversity and ecosystem functions [J]. Biodiv Sci, 2025, 33(4): 24579-. |
| [6] | Zhang Haobin, Xiao Lu, Liu Yanjie. Effects of artificial light at night on the diversity and growth of invasive alien and native plants [J]. Biodiv Sci, 2025, 33(4): 24553-. |
| [7] | Weiyu Lu, Xu Chen, Riqian Zhang, Yunhai Zhang, Zhao Li. Effects of nitrogen and cow dung input on plant species-area relationship in a typical grassland [J]. Biodiv Sci, 2025, 33(12): 25251-. |
| [8] | Junhan Shen, Haiyang Gao, Song Sun, Fei Wu, Nan He, He Wang, Yan Hua, Zhengjun Wu. Microhabitat selection characteristics and conservation implications of the Chinese crocodile lizard (Shinisaurus crocodilurus) [J]. Biodiv Sci, 2025, 33(11): 25203-. |
| [9] | Jiali Lian, Jing Chen, Xueqin Yang, Ying Zhao, Xu Luo, Cui Han, Yaxin Zhao, Jianping Li. Responses of desert steppe plant diversity and microbial diversity to precipitation change [J]. Biodiv Sci, 2024, 32(6): 24044-. |
| [10] | Fengming Wan, Huawei Wan, Zhiru Zhang, Jixi Gao, Chenxi Sun, Yongcai Wang. The application potential of unmanned aerial vehicle surveys in grassland plant diversity [J]. Biodiv Sci, 2024, 32(3): 23381-. |
| [11] | Naipeng Zhang, Hongru Liang, Yan Zhang, Chao Sun, Yong Chen, Lulu Wang, Jiangbao Xia, FangLei Gao. Effects of soil type and groundwater depth on spatial differentiation of typical salt marsh plant communities in the Yellow River Delta [J]. Biodiv Sci, 2024, 32(2): 23370-. |
| [12] | Chen-Kun Jiang, Wen-Bin Yu, Guang-Yuan Rao, Huaicheng Li, Julien B. Bachelier, Hartmut H. Hilger, Theodor C. H. Cole. Plant Phylogeny Posters—An educational project on plant diversity from an evolutionary perspective [J]. Biodiv Sci, 2024, 32(11): 24210-. |
| [13] | Yun Han, Xiaofeng Chi, Jingya Yu, Xujie Ding, Shilong Chen, Faqi Zhang. A checklist of wild vascular plants in Qinghai, China [J]. Biodiv Sci, 2023, 31(9): 23280-. |
| [14] | Yousheng Chen, Zhuqiu Song, Ran Wei, Yan Luo, Wenli Chen, Fusheng Yang, Lianming Gao, Yuan Xu, Zhuoxin Zhang, Pengcheng Fu, Chunlei Xiang, Huanchong Wang, Jiachen Hao, Shiyong Meng, Lei Wu, Bo Li, Shengxiang Yu, Shuren Zhang, Li He, Xinqiang Guo, Wenguang Wang, Yihua Tong, Qi Gao, Wenqun Fei, Youpai Zeng, Lin Bai, Zichao Jin, Xingjie Zhong, Buyun Zhang, Siyi Du. A dataset on inventory and geographical distribution of vascular plants in Xizang, China [J]. Biodiv Sci, 2023, 31(9): 23188-. |
| [15] | Zhuqiu Song, Wen Ye, Shiyong Dong, Zichao Jin, Xingjie Zhong, Zhen Wang, Buyun Zhang, Yechun Xu, Wenli Chen, Shijin Li, Gang Yao, Zhoufeng Xu, Shuai Liao, Yihua Tong, Youpai Zeng, Yunbao Zeng, Yousheng Chen. A dataset on inventory and geographical distributions of higher plants in Guangdong, China [J]. Biodiv Sci, 2023, 31(9): 23177-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn