



Biodiv Sci ›› 2023, Vol. 31 ›› Issue (8): 23146. DOI: 10.17520/biods.2023146 cstr: 32101.14.biods.2023146
• Original Papers: Microbial Diversity • Previous Articles Next Articles
Chunling Wu1, Zhuhui Luo1, Yide Li2, Han Xu2, Dexiang Chen2, Qiong Ding1,*( )
)
Received:2023-05-08
Accepted:2023-06-30
Online:2023-08-20
Published:2023-07-10
Contact:
*E-mail: dingqiong@hainu.edu.cn
Chunling Wu, Zhuhui Luo, Yide Li, Han Xu, Dexiang Chen, Qiong Ding. Foliar endophytic bacterial communities of woody Fabaceae and Lauraceae plants in tropical mountain rainforests: Understanding species and functional diversity and their driving factors[J]. Biodiv Sci, 2023, 31(8): 23146.
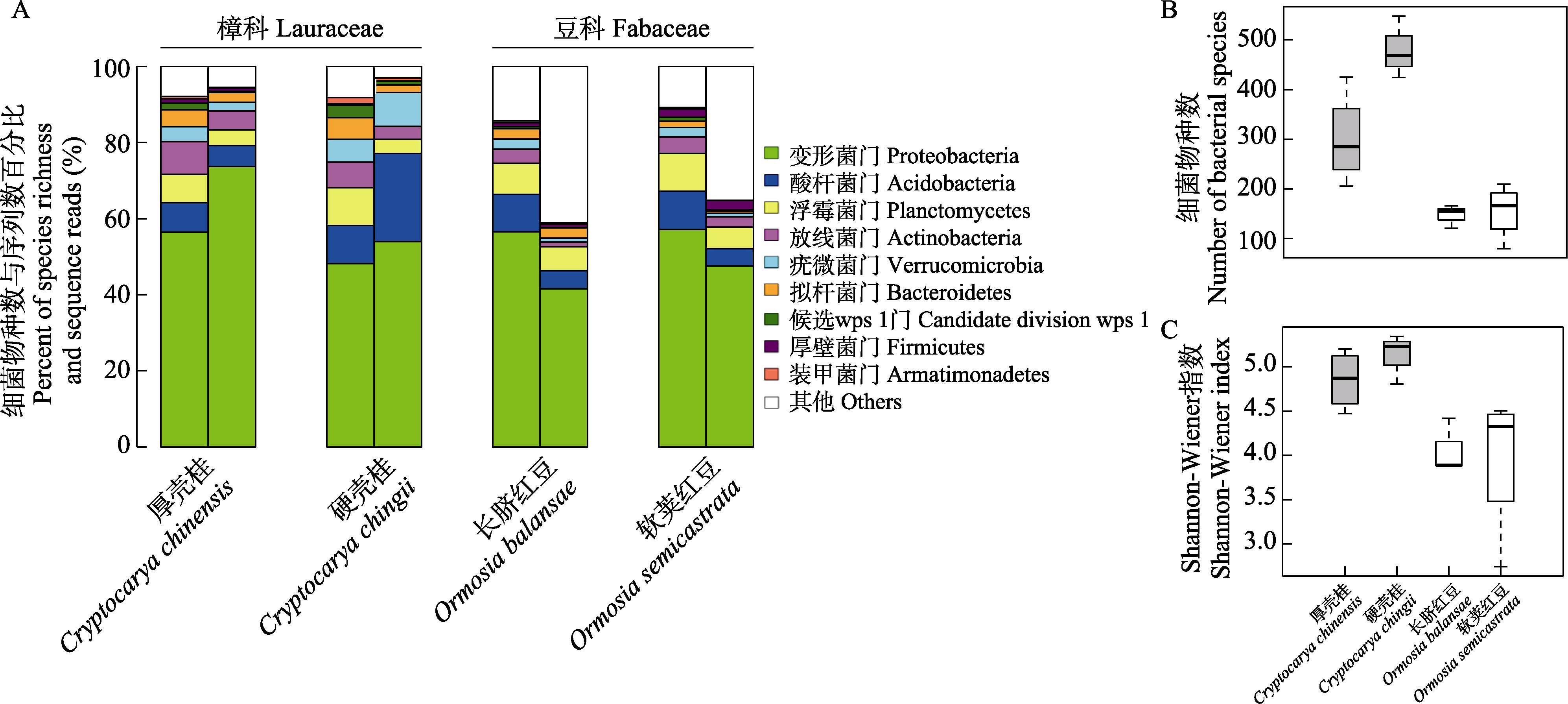
Fig. 1 A comparison of foliar endophytic bacteria diversity among the 4 species of Fabaceae and Lauraceae. A, Percent species richness (left column) and sequence reads (right column) at phylum level; B, Species richness; C, Shannon-Wiener’s index.
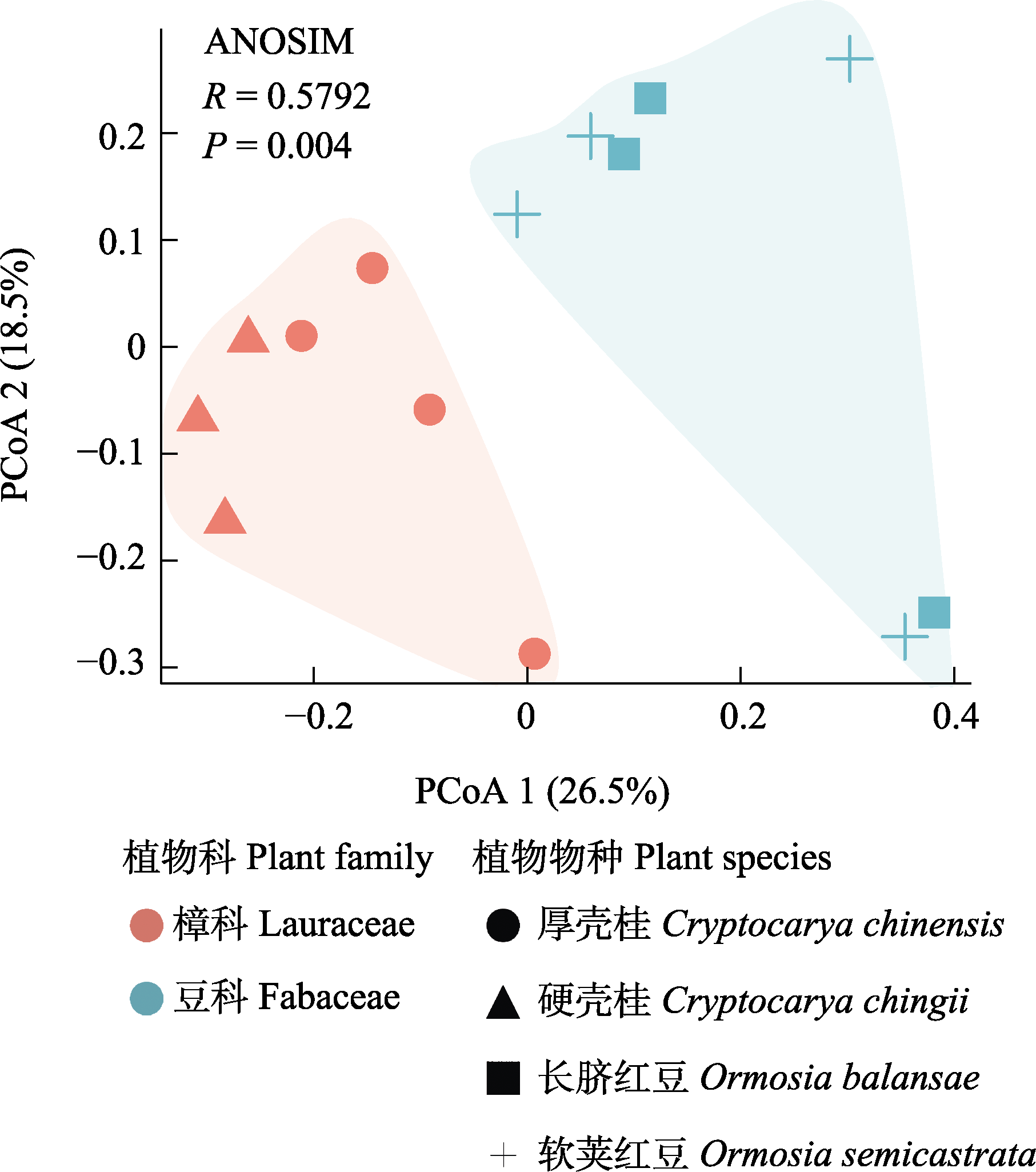
Fig. 2 Principal coordinate analysis (PCoA) and analysis of similarity (ANOSIM) of foliar endophytic bacterial community of four Fabaceae and Lauraceae plants
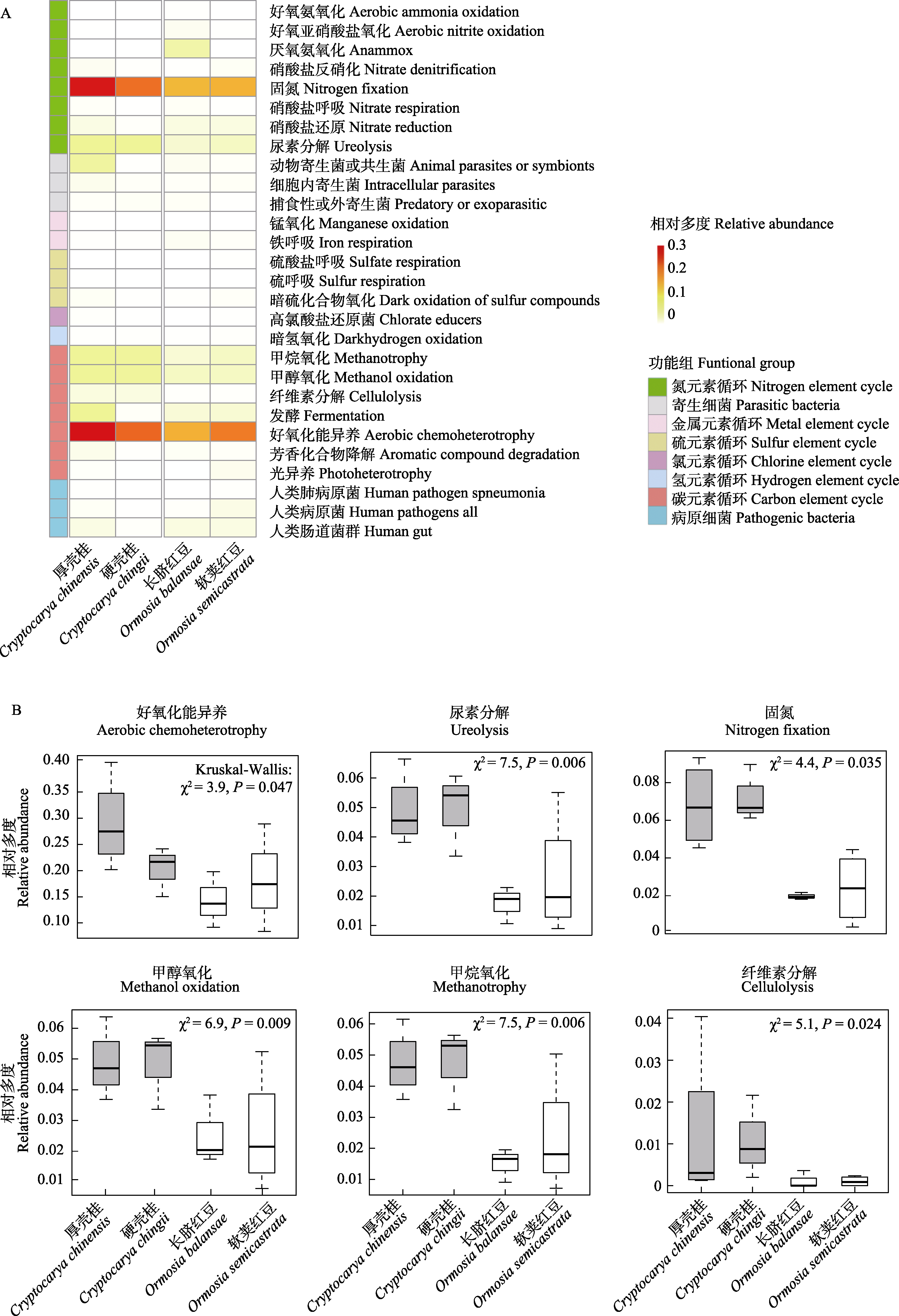
Fig. 3 Heatmap of functional group profiles (A) and key functional groups (B) of foliar endophytic bacteria in the Lauraceae and Fabaceae plant species
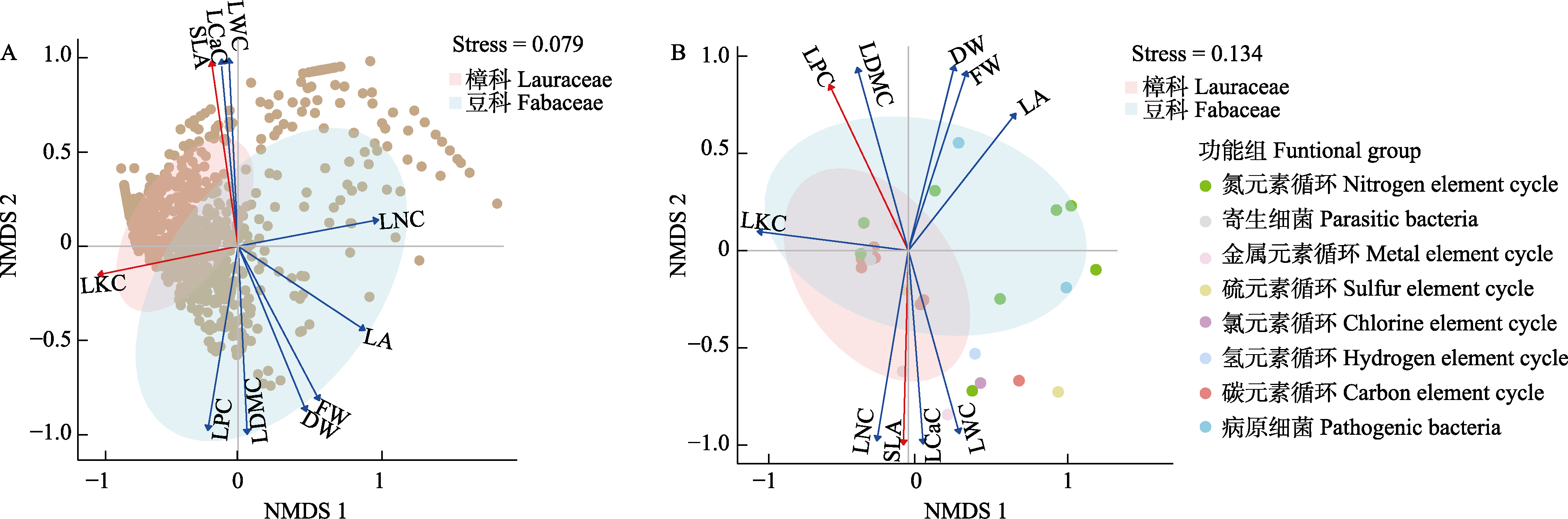
Fig. 4 Correlation between host plant leaf traits and foliar endophytic bacterial community species (A) and functional group (B) compositions. SLA, Specific leaf area; FW, Fresh weight; DW, Dry weight; LDMC, Leaf dry matter content; LWC, Leaf water content; LA, Leaf area; LKC, Leaf potassium content; LNC, Leaf nitrogen content; LPC, Leaf phosphorus content; LCaC, Leaf calcium content.
| [1] |
Abadi VAJM, Sepehri M, Rahmani HA, Dolatabad HK, Shamshiripour M, Khatabi B (2021) Diversity and abundance of culturable nitrogen-fixing bacteria in the phyllosphere of maize. Journal of Applied Microbiology, 131, 898-912.
DOI PMID |
| [2] | Bao SD (2000) Soil and Agricultural Chemistry Analysis, 3nd edn. China Agricultural Press, Beijing. (in Chinese) |
| [ 鲍士旦 (2000) 土壤农化分析(第三版). 中国农业出版社, 北京.] | |
| [3] |
Bao ZH, Okubo T, Kubota K, Kasahara Y, Tsurumaru H, Anda M, Ikeda S, Minamisawa K (2014) Metaproteomic identification of diazotrophic methanotrophs and their localization in root tissues of field-grown rice plants. Applied and Environmental Microbiology, 80, 5043-5052.
DOI PMID |
| [4] | Barman D, Dkhar MS (2019) Plant growth-promoting potential of endophytic bacteria isolated from Costus speciosus in tropical deciduous forest of eastern Himalaya. Proceedings of the National Academy of Sciences, India Section B: Biological Sciences, 89, 841-852. |
| [5] |
Batterman SA, Hedin LO, van Breugel M, Ransijn J, Craven DJ, Hall JS (2013) Key role of symbiotic dinitrogen fixation in tropical forest secondary succession. Nature, 502, 224-227.
DOI |
| [6] | Bērtiņš M, Klūga A, Dubova L, Petrēvics P, Alsiņa I, Vīksna A (2021) Study of rhizobia impact on nutritional element concentration in legumes. Proceedings of the Latvian Academy of Sciences Section B Natural, Exact, and Applied Sciences, 75, 457-462. |
| [7] |
Cornelissen J, Lavorel S, Garnier E, Díaz S, Buchmann N, Gurvich DE, Reich PB, Steege HT, Morgan HD, Heijden M (2003) A handbook of protocols for standardised and easy measurement of plant functional traits worldwide. Australian Journal of Botany, 51, 335-380.
DOI URL |
| [8] |
Cunha HFV, Andersen KM, Lugli LF, Santana FD, Aleixo IF, Moraes AM, Garcia S, Di Ponzio R, Mendoza EO, Brum B, Rosa JS, Cordeiro AL, Portela BTT, Ribeiro G, Coelho SD, de Souza ST, Silva LS, Antonieto F, Pires M, Salomão AC, Miron AC, de Assis RL, Domingues TF, Aragão LEOC, Meir P, Camargo JL, Manzi AO, Nagy L, Mercado LM, Hartley IP, Quesada CA (2022) Direct evidence for phosphorus limitation on Amazon forest productivity. Nature, 608, 558-562.
DOI |
| [9] | Deng FL, Xu H, Chen J, Lin MX, Li YD (2022) Effects of leguminous trees on neighboring tree species in the tropical mountainous rainforest of Jianfengling. Forest Research, 35(3), 1-8. (in Chinese with English abstract) |
| [ 邓方立, 许涵, 陈洁, 林明献, 李意德 (2022) 尖峰岭热带山地雨林豆科树木对邻体树种的影响. 林业科学研究, 35(3), 1-8.] | |
| [10] | Ding T, Ulrich M, Lorenzo B (2016) Influences of plant species, season and location on leaf endophytic bacterial communities of non-cultivated plants. PLoS ONE, 11, e0150895. |
| [11] |
Donald J, Roy M, Suescun U, Iribar A, Manzi S, Péllissier L, Gaucher P, Chave J, Singh B (2020) A test of community assembly rules using foliar endophytes from a tropical forest canopy. Journal of Ecology, 108, 1605-1616.
DOI URL |
| [12] |
Dudeja SS, Giri R, Saini R, Suneja-Madan P, Kothe E (2012) Interaction of endophytic microbes with legumes. Journal of Basic Microbiology, 52, 248-260.
DOI PMID |
| [13] | Epihov DZ, Saltonstall K, Batterman SA, Hedin LO, Hall JS, van Breugel M, Leake JR, Beerling DJ (2021) Legume-microbiome interactions unlock mineral nutrients in regrowing tropical forests. Proceedings of the National Academy of Sciences, USA, 118, e2022241118. |
| [14] |
Fang JY, Li YD, Zhu B, Liu GH, Zhou GY (2004) Community structure, species diversity, and position in the global rainforest of Jianfengling mountainous rainforest on Hainan Island in China. Biodiversity Science, 12, 29-43. (in Chinese with English abstract)
DOI URL |
|
[ 方精云, 李意德, 朱彪, 刘国华, 周光益 (2004) 海南岛尖峰岭山地雨林的群落结构、物种多样性以及在世界雨林中的地位. 生物多样性, 12, 29-43.]
DOI |
|
| [15] |
Fürnkranz M, Wanek W, Richter A, Abell G, Rasche F, Sessitsch A (2008) Nitrogen fixation by phyllosphere bacteria associated with higher plants and their colonizing epiphytes of a tropical lowland rainforest of Costa Rica. The ISME Journal, 2, 561-570.
DOI |
| [16] |
Holland-Moritz H, Stuart JEM, Lewis LR, Miller SN, Mack MC, Ponciano JM, McDaniel SF, Fierer N (2021) The bacterial communities of Alaskan mosses and their contributions to N-fixation. Microbiome, 9, 53.
DOI PMID |
| [17] |
Hong CE, Jo SH, Moon JY, Lee JS, Kwon SY, Park JM (2015) Isolation of novel leaf-inhabiting endophytic bacteria in Arabidopsis thaliana and their antagonistic effects on phytophathogens. Plant Biotechnology Reports, 9, 451-458.
DOI URL |
| [18] |
Houlton BZ, Wang YP, Vitousek PM, Field CB (2008) A unifying framework for dinitrogen fixation in the terrestrial biosphere. Nature, 454, 327-330.
DOI |
| [19] | Huang CW, Liao YH, Ding Q (2017) Two sample pooling strategies revealed different root-associated fungal diversity of Rhododendron species. Acta Microbiologica Sinica, 57, 571-518. (in Chinese with English abstract) |
| [ 黄彩微, 廖映辉, 丁琼 (2017) 两种混合样品策略对揭示杜鹃花根部真菌多样性的影响. 微生物学报, 57, 571-581.] | |
| [20] |
Ikeda S, Sasaki K, Okubo T, Yamashita A, Terasawa K, Bao ZH, Liu DY, Watanabe T, Murase J, Asakawa S, Eda SM, Mitsui H, Sato T, Minamisawa K (2014) Low nitrogen fertilization adapts rice root microbiome to low nutrient environment by changing biogeochemical functions. Microbes and Environments, 29, 50-59.
PMID |
| [21] |
Jung SW, Kang JS, Park JS, Joo HM, Suh SS, Kang D, Lee TK, Kim HJ (2021) Dynamic bacterial community response to Akashiwo sanguinea (Dinophyceae) bloom in indoor marine microcosms. Scientific Reports, 11, 6983.
DOI |
| [22] |
Kandasamy GD, Kathirvel P (2023) Insights into bacterial endophytic diversity and isolation with a focus on their potential applications—A review. Microbiological Research, 266, 127256.
DOI URL |
| [23] | Kembel SW, O’Connor TK, Arnold HK, Hubbell SP, Wright SJ, Green JL (2014) Relationships between phyllosphere bacterial communities and plant functional traits in a neotropical forest. Proceedings of the National Academy of Sciences, USA, 111, 13715-13720. |
| [24] |
Kim M, Singh D, Lai-Hoe A, Go R, Rahim RA, Ainuddin AN, Chun J, Adams JM (2012) Distinctive phyllosphere bacterial communities in tropical trees. Microbial Ecology, 63, 674-681.
DOI PMID |
| [25] |
Li S, Wu CB, Liu H, Lyu XC, Xiao FS, Zhao SH, Ma CM, Yan C, Liu ZL, Li HY, Wang XL, Gong ZP (2023) Systemic regulation of nodule structure and assimilated carbon distribution by nitrate in soybean. Frontiers in Plant Science, 14, 1101074.
DOI URL |
| [26] |
Louca S, Parfrey LW, Doebeli M (2016) Decoupling function and taxonomy in the global ocean microbiome. Science, 353, 1272-1277.
DOI PMID |
| [27] | Lu RK (2000) Methods of Soil Agricultural Chemical Analysis. China Agricultural Science and Technology Press, Beijing. (in Chinese) |
| [ 鲁如坤 (2000) 土壤农业化学分析方法. 中国农业科技出版社, 北京.] | |
| [28] |
Luo D, Cheng RM, Shi ZM, Wang WX, Xu GX, Liu SR (2016) Impacts of nitrogen-fixing and non-nitrogen-fixing tree species on soil respiration and microbial community composition during forest management in subtropical China. Ecological Research, 31, 683-693.
DOI URL |
| [29] |
Moreira JCF, Brum M, de Almeida LC, Barrera-Berdugo S, de Souza AA, de Camargo PB, Oliveira RS, Alves LF, Rosado BHP, Lambais MR (2021) Asymbiotic nitrogen fixation in the phyllosphere of the Amazon forest: Changing nitrogen cycle paradigms. Science of the Total Environment, 773, 145066.
DOI URL |
| [30] |
Nurbolat S, Lv GH, Jiang LM, Zhang L (2022) Convergent variation in the leaf traits of desert plants in the Ebinur Lake basin. Frontiers in Environmental Science, 10, 927572.
DOI URL |
| [31] |
Rivett DW, Bell T (2018) Abundance determines the functional role of bacterial phylotypes in complex communities. Nature Microbiology, 3, 767-772.
DOI PMID |
| [32] | Shi LL, Luo TS, Xu H, Lin MX, Yang H, Chen DX, Li YD (2012) The fine scale spatial heterogeneity of soil physical properties in a primary tropical montane rainforest of Jianfengling, Hainan Island, China. Forest Research, 25, 285-293. (in Chinese with English abstract) |
| [ 时雷雷, 骆土寿, 许涵, 林明献, 杨怀, 陈德祥, 李意德 (2012) 尖峰岭热带山地雨林土壤物理性质小尺度空间异质性研究. 林业科学研究, 25, 285-293.] | |
| [33] | Sinclair L, Osman OA, Bertilsson S, Eiler A (2015) Microbial community composition and diversity via 16S rRNA gene amplicons: Evaluating the Illumina platform. PLoS ONE, 10, e0116955. |
| [34] |
Sprent JI, Ardley J, James EK (2017) Biogeography of nodulated legumes and their nitrogen-fixing symbionts. New Phytologist, 215, 40-56.
DOI PMID |
| [35] |
Sun F, Song CJ, Wang M, Lai DYF, Tariq A, Zeng FJ, Zhong QP, Wang FM, Li ZA, Peng CL (2020) Long-term increase in rainfall decreases soil organic phosphorus decomposition in tropical forests. Soil Biology and Biochemistry, 151, 108056.
DOI URL |
| [36] | Tang X, Liu SR, Xu H, Zhang YG (2019) Estimation of soil microbial species in the 60-hm2 dynamic monitoring plot of tropical mountain rainforest in Jianfengling, Hainan Island. Scientia Silvae Sinicae, 55(12), 84-92. (in Chinese with English abstract) |
| [ 唐欣, 刘世荣, 许涵, 张于光 (2019) 海南尖峰岭热带山地雨林60 hm2动态监测样地土壤微生物物种估算. 林业科学, 55(12), 84-92.] | |
| [37] |
Trivedi P, Leach JE, Tringe SG, Sa TM, Singh BK (2020) Plant-microbiome interactions: From community assembly to plant health. Nature Reviews Microbiology, 18, 607-621.
DOI |
| [38] |
Wei YQ, Lan GY, Wu ZX, Chen BQ, Quan F, Li MM, Sun SQ, Du HN (2022) Phyllosphere fungal communities of rubber trees exhibited biogeographical patterns, but not bacteria. Environmental Microbiology, 24, 3777-3790.
DOI PMID |
| [39] |
Westoby M, Falster DS, Moles AT, Vesk PA, Wright IJ (2002) Plant ecological strategies: Some leading dimensions of variation between species. Annual Review of Ecology and Systematics, 33, 125-159.
DOI URL |
| [40] |
Xu H, Detto M, Fang S, Chazdon RL, Li Y, Hau BCH, Fischer GA, Weiblen GD, Hogan JA, Zimmerman JK, Uriarte M, Thompson J, Lian J, Cao K, Kenfack D, Alonso A, Bissiengou P, Memiaghe HR, Valencia R, Yap SL, Davies SJ, Mi X, Yao TL (2020) Soil nitrogen concentration mediates the relationship between leguminous trees and neighbor diversity in tropical forests. Communications Biology, 3, 317.
DOI PMID |
| [41] |
Xu H, Li YD, Lin MX, Wu JH, Luo TS, Zhou Z, Chen DX, Yang H, Li GJ, Liu SR (2015) Community structure and characteristics of the 60 ha dynamic monitoring plot of tropical mountain rainforest in Jianfengling, Hainan Island. Biodiversity Science, 23, 192-201. (in Chinese with English abstract)
DOI URL |
|
[ 许涵, 李意德, 林明献, 吴建辉, 骆土寿, 周璋, 陈德祥, 杨怀, 李广建, 刘世荣 (2015) 海南尖峰岭热带山地雨林60 ha动态监测样地群落结构特征. 生物多样性, 23, 192-201.]
DOI |
|
| [42] |
Xu NH, Zhao QQ, Zhang ZY, Zhang Q, Wang Y, Qin GY, Ke MJ, Qiu DY, Peijnenburg WJGM, Lu T, Qian HF (2022) Phyllosphere microorganisms: Sources, drivers, and their interactions with plant hosts. Journal of Agricultural and Food Chemistry, 70, 4860-4870.
DOI PMID |
| [43] |
Yoneyama T, Terakado-Tonooka J, Bao Z, Minamisawa K (2019) Molecular analyses of the distribution and function of diazotrophic rhizobia and methanotrophs in the tissues and rhizosphere of non-leguminous plants. Plants, 8, 408.
DOI URL |
| [44] | Zhu YG, Peng JJ, Chen C, Xiong C, Li SL, Ge AH, Wang ET Liesack W, (2023) Harnessing biological nitrogen fixation in plant leaves. Trends in Plant Science, doi: 10.1016/j.tplants.2023.05.009. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn