



Biodiv Sci ›› 2025, Vol. 33 ›› Issue (12): 25233. DOI: 10.17520/biods.2025233 cstr: 32101.14.biods.2025233
• Original Papers:Genetic Diversity • Previous Articles Next Articles
Jinshan Wu1, Changle Yang2, Yufeng Ma3, Yaxuan Li1, Wenjia Gao2, Ye Kusili2, Lianghong Ye2, Yujiao Yang1, Mengqi Xu1, Tingqiong Liao1, Linqiang Zhong1,*( ), Wenjuan Shan1,*(
), Wenjuan Shan1,*( )
)
Received:2025-06-18
Accepted:2025-11-18
Online:2025-12-20
Published:2026-01-09
Supported by:Jinshan Wu, Changle Yang, Yufeng Ma, Yaxuan Li, Wenjia Gao, Ye Kusili, Lianghong Ye, Yujiao Yang, Mengqi Xu, Tingqiong Liao, Linqiang Zhong, Wenjuan Shan. Genetic diversity and genetic structure of red deer in the Ebinur Lake Wetland National Nature Reserve[J]. Biodiv Sci, 2025, 33(12): 25233.
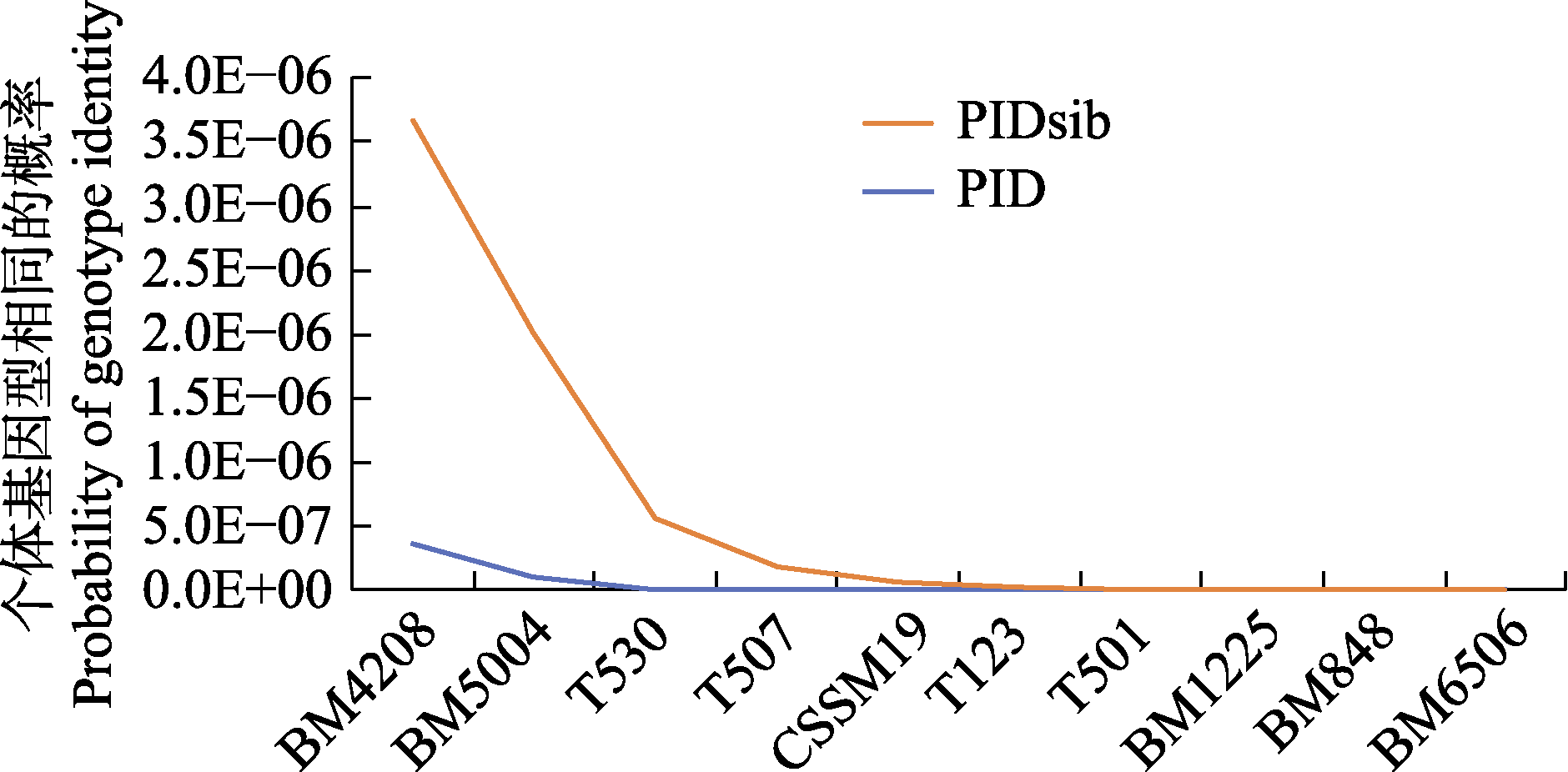
Fig. 1 Results of cumulative gene locus identity consistency resolution testing for the protection of red deer in Ebinur Lake Wetland National Nature Reserve. PIDsib, Probability of identity for siblings; PID, Probability of identity for individuals.
| Locus | N | Na | Ne | I | Ho | He | PIC | Fis | P |
|---|---|---|---|---|---|---|---|---|---|
| BM4208 | 73 | 16 | 6.952 | 2.154 | 0.589 | 0.856 | 0.799 | 0.312 | 0.000*** |
| BM5004 | 73 | 7 | 2.100 | 0.972 | 0.110 | 0.524 | 0.416 | 0.791 | 0.000*** |
| T530 | 73 | 16 | 9.716 | 2.439 | 0.767 | 0.897 | 0.871 | 0.145 | 0.000*** |
| T507 | 73 | 11 | 6.741 | 2.048 | 0.822 | 0.852 | 0.823 | 0.035 | 0.013* |
| CSSM19 | 73 | 12 | 7.506 | 2.166 | 0.890 | 0.867 | 0.855 | -0.027 | 0.000*** |
| T123 | 73 | 14 | 4.320 | 1.872 | 0.808 | 0.769 | 0.685 | -0.052 | 0.000*** |
| T501 | 73 | 23 | 4.860 | 2.192 | 0.589 | 0.794 | 0.662 | 0.258 | 0.000*** |
| BM1225 | 73 | 19 | 7.706 | 2.342 | 0.671 | 0.870 | 0.849 | 0.229 | 0.000*** |
| BM848 | 73 | 13 | 2.771 | 1.339 | 0.603 | 0.639 | 0.536 | 0.057 | 0.000*** |
| BM6506 | 73 | 10 | 3.332 | 1.554 | 0.767 | 0.700 | 0.619 | -0.096 | 0.000*** |
| 平均值 Mean | 73 | 14.1 | 5.600 | 1.908 | 0.662 | 0.777 | 0.712 | 0.165 |
Table 1 Genetic diversity parameters of 10 pairs of microsatellite loci
| Locus | N | Na | Ne | I | Ho | He | PIC | Fis | P |
|---|---|---|---|---|---|---|---|---|---|
| BM4208 | 73 | 16 | 6.952 | 2.154 | 0.589 | 0.856 | 0.799 | 0.312 | 0.000*** |
| BM5004 | 73 | 7 | 2.100 | 0.972 | 0.110 | 0.524 | 0.416 | 0.791 | 0.000*** |
| T530 | 73 | 16 | 9.716 | 2.439 | 0.767 | 0.897 | 0.871 | 0.145 | 0.000*** |
| T507 | 73 | 11 | 6.741 | 2.048 | 0.822 | 0.852 | 0.823 | 0.035 | 0.013* |
| CSSM19 | 73 | 12 | 7.506 | 2.166 | 0.890 | 0.867 | 0.855 | -0.027 | 0.000*** |
| T123 | 73 | 14 | 4.320 | 1.872 | 0.808 | 0.769 | 0.685 | -0.052 | 0.000*** |
| T501 | 73 | 23 | 4.860 | 2.192 | 0.589 | 0.794 | 0.662 | 0.258 | 0.000*** |
| BM1225 | 73 | 19 | 7.706 | 2.342 | 0.671 | 0.870 | 0.849 | 0.229 | 0.000*** |
| BM848 | 73 | 13 | 2.771 | 1.339 | 0.603 | 0.639 | 0.536 | 0.057 | 0.000*** |
| BM6506 | 73 | 10 | 3.332 | 1.554 | 0.767 | 0.700 | 0.619 | -0.096 | 0.000*** |
| 平均值 Mean | 73 | 14.1 | 5.600 | 1.908 | 0.662 | 0.777 | 0.712 | 0.165 |
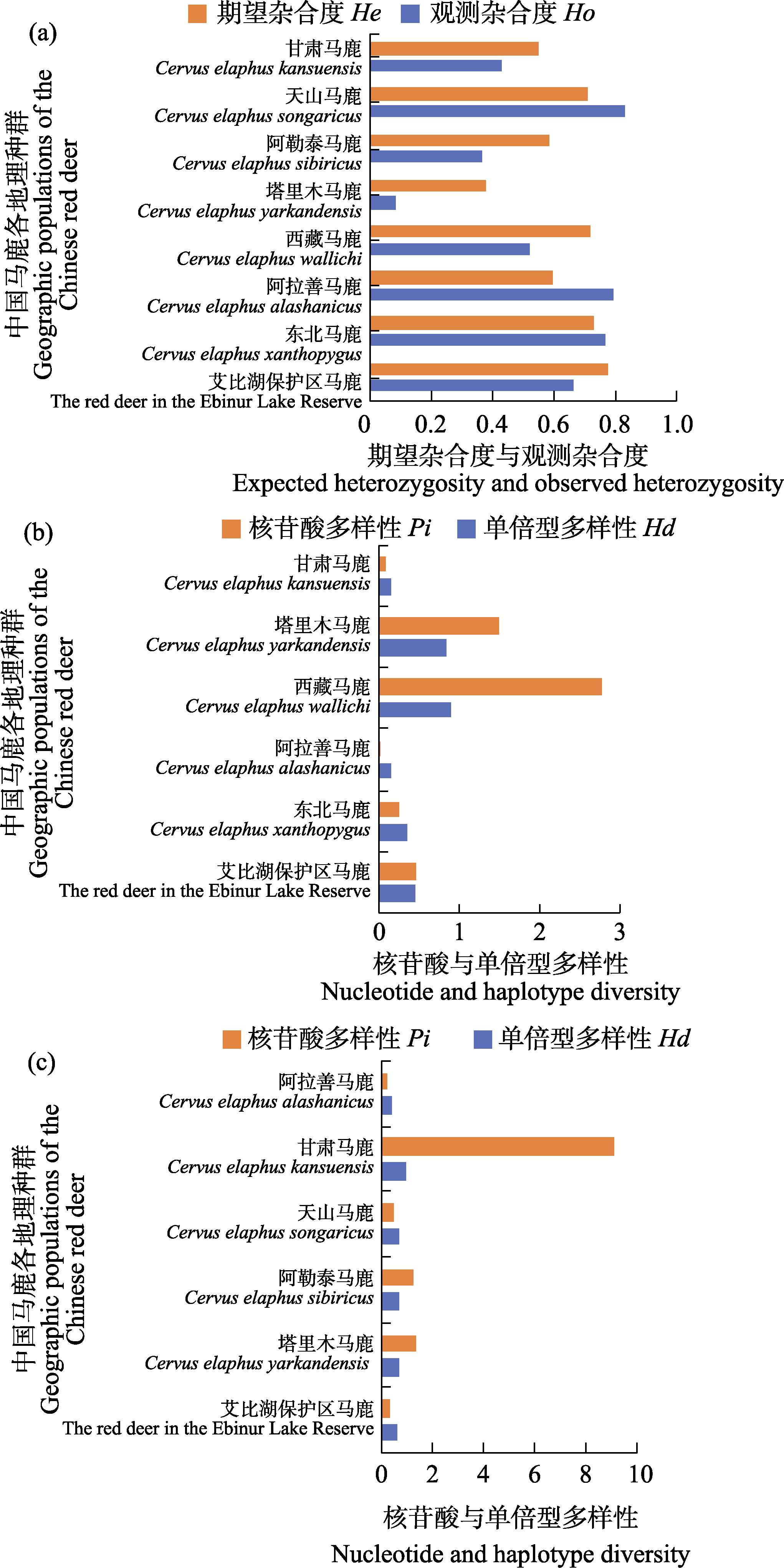
Fig. 2 Comparative analysis of genetic diversity in red deer from in the Ebinur Lake Wetland National Nature Reserve (Ebinur Lake Reserve) and other red deer populations. (a) Microsatellite genetic diversity; (b) Cytb genetic diversity (c) D-loop genetic diversity. Ho, Observed heterozygosity; He, Expected heterozygosity; Pi, Nucleotide diversity; Hd, Haplotype diversity.
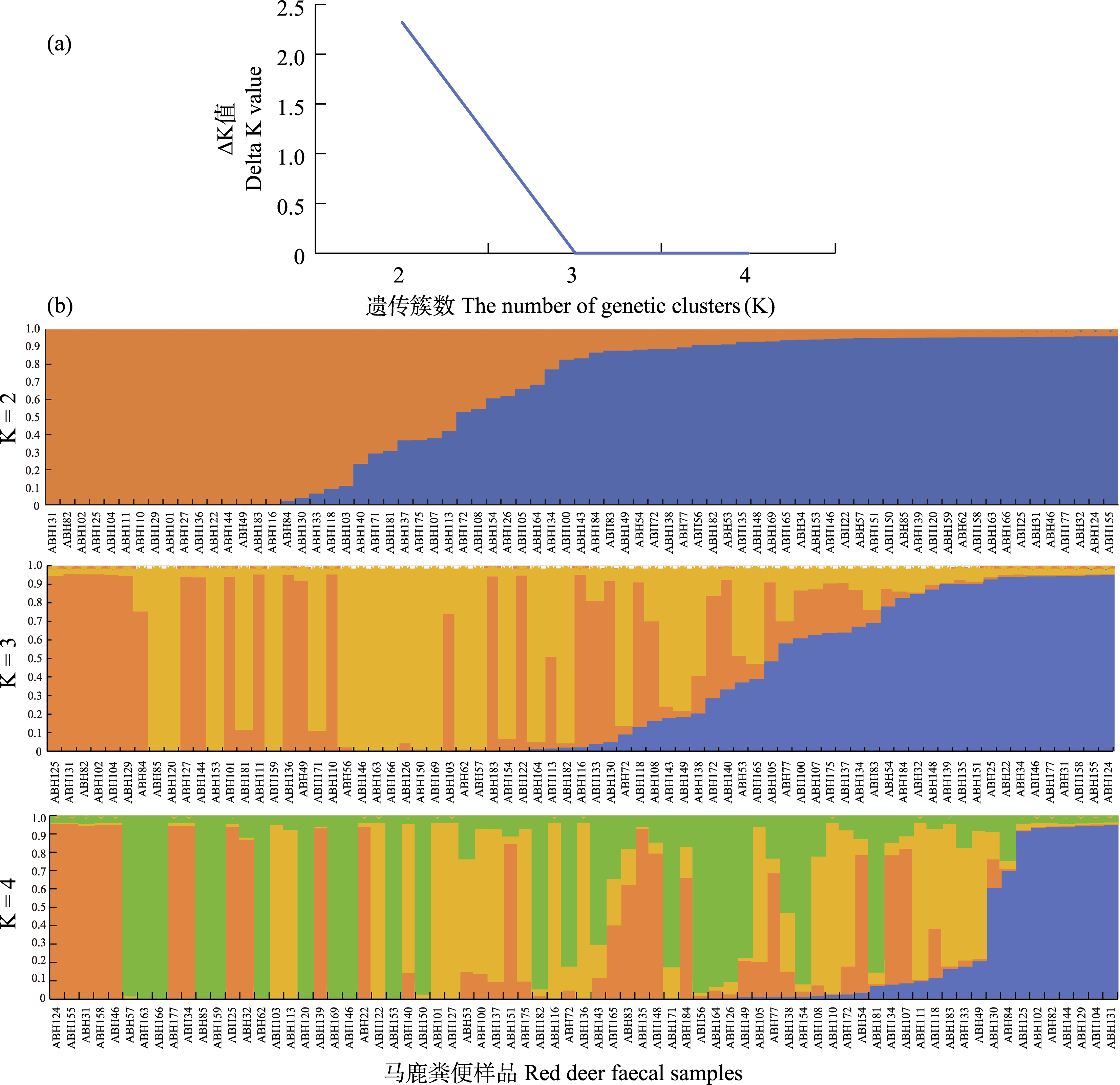
Fig. 3 The ΔK results (a) and the corresponding internal genetic structure (b) of the red deer population in Ebinur Lake Wetland National Nature Reserve under different K values
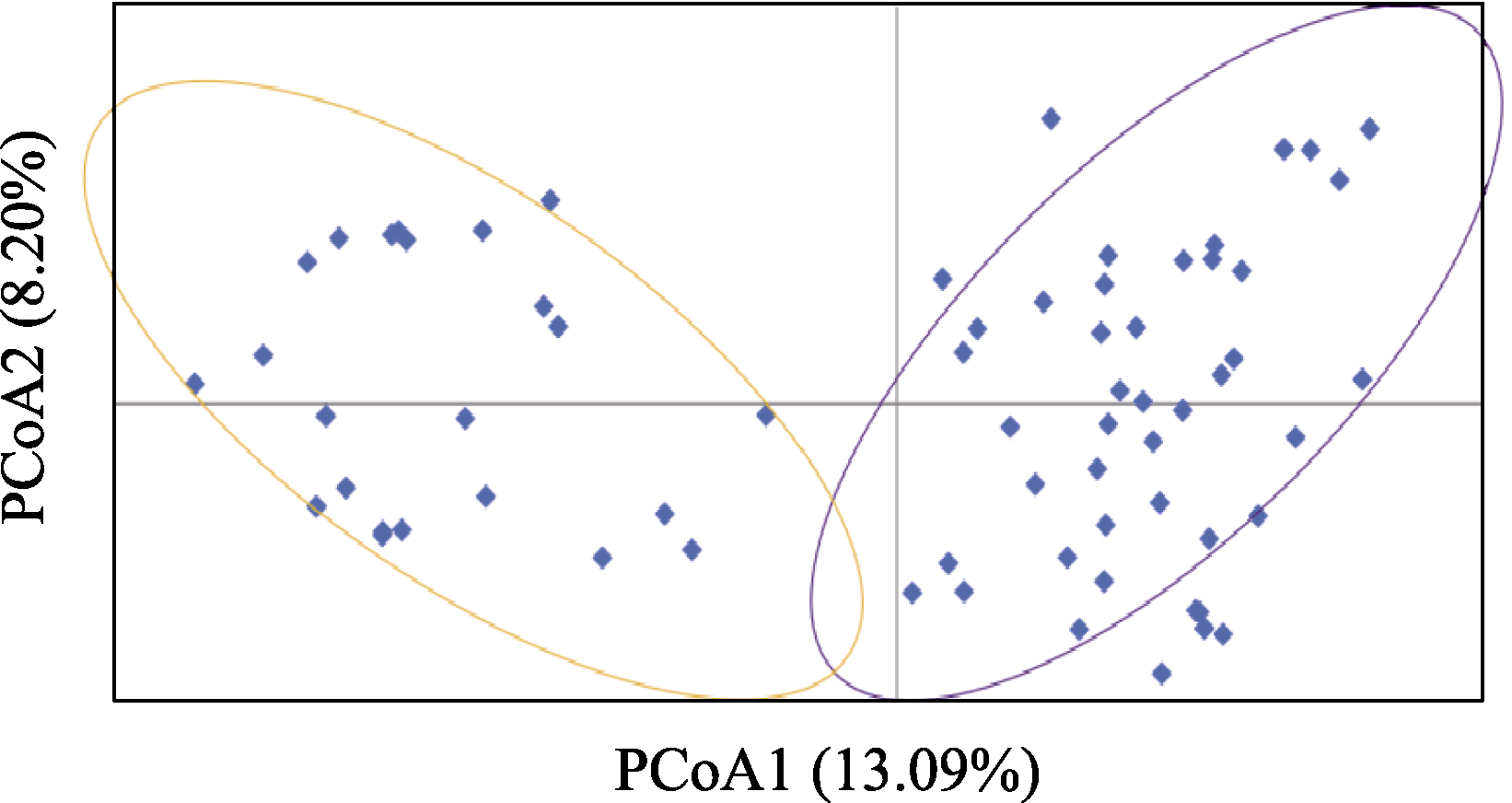
Fig. 4 Principal coordinates analysis (PCoA) genetic structure analysis of red deer in Ebinur Lake Wetland National Nature Reserve based on microsatellites
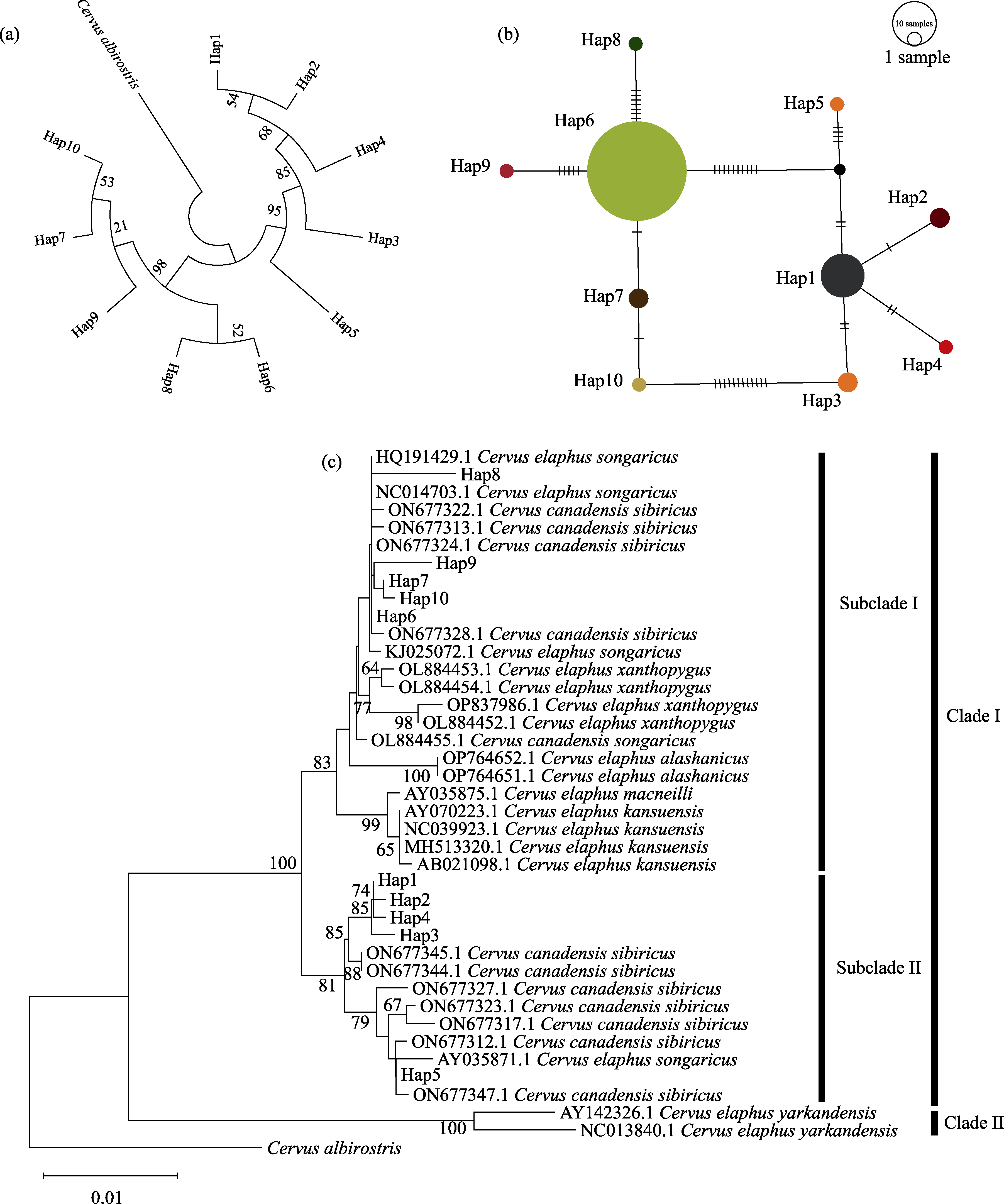
Fig. 5 Genetic structure and phylogenetic relationship of red deer populations in Ebinur Lake Wetland National Nature Reserve. (a) Phylogenetic tree of Cytb haplotypes; (b) Haplotype network diagram of Cytb haplotypes; (c) Phylogenetic tree of Cytb haplotypes in red deer from Ebinur Lake Wetland National Nature Reserve and Chinese red deer subspecies; In Figure c, Clade I, Clade II, Subclade I, and Subclade II correspond sequentially to the red deer populations of the Tianshan-Altai-Northeast-Alashan- Sichuan-Gansu region and the Ebinur Lake Wetland National Nature Reserve, the Tarim red deer population, the Tianshan-Altai-Northeast-Alashan-Sichuan-Gansu region and the Ebinur Lake Wetland National Nature Reserve red deer population, and the Tianshan-Altai and Ebinur Lake Wetland National Nature Reserve red deer populations.
| ABH | ALT | TS | DB | ALS | GS | SC | |
|---|---|---|---|---|---|---|---|
| ALT | 0.010 | ||||||
| TS | 0.009 | 0.009 | |||||
| DB | 0.010 | 0.011 | 0.008 | ||||
| ALS | 0.014 | 0.014 | 0.010 | 0.011 | |||
| GS | 0.012 | 0.013 | 0.009 | 0.011 | 0.014 | ||
| SC | 0.012 | 0.012 | 0.009 | 0.010 | 0.014 | 0.002 | |
| TLM | 0.054 | 0.051 | 0.052 | 0.053 | 0.053 | 0.052 | 0.052 |
Table 2 Cytb genetic distance between red deer and Chinese red deer subspecies in Ebinur Lake Wetland National Nature Reserve
| ABH | ALT | TS | DB | ALS | GS | SC | |
|---|---|---|---|---|---|---|---|
| ALT | 0.010 | ||||||
| TS | 0.009 | 0.009 | |||||
| DB | 0.010 | 0.011 | 0.008 | ||||
| ALS | 0.014 | 0.014 | 0.010 | 0.011 | |||
| GS | 0.012 | 0.013 | 0.009 | 0.011 | 0.014 | ||
| SC | 0.012 | 0.012 | 0.009 | 0.010 | 0.014 | 0.002 | |
| TLM | 0.054 | 0.051 | 0.052 | 0.053 | 0.053 | 0.052 | 0.052 |
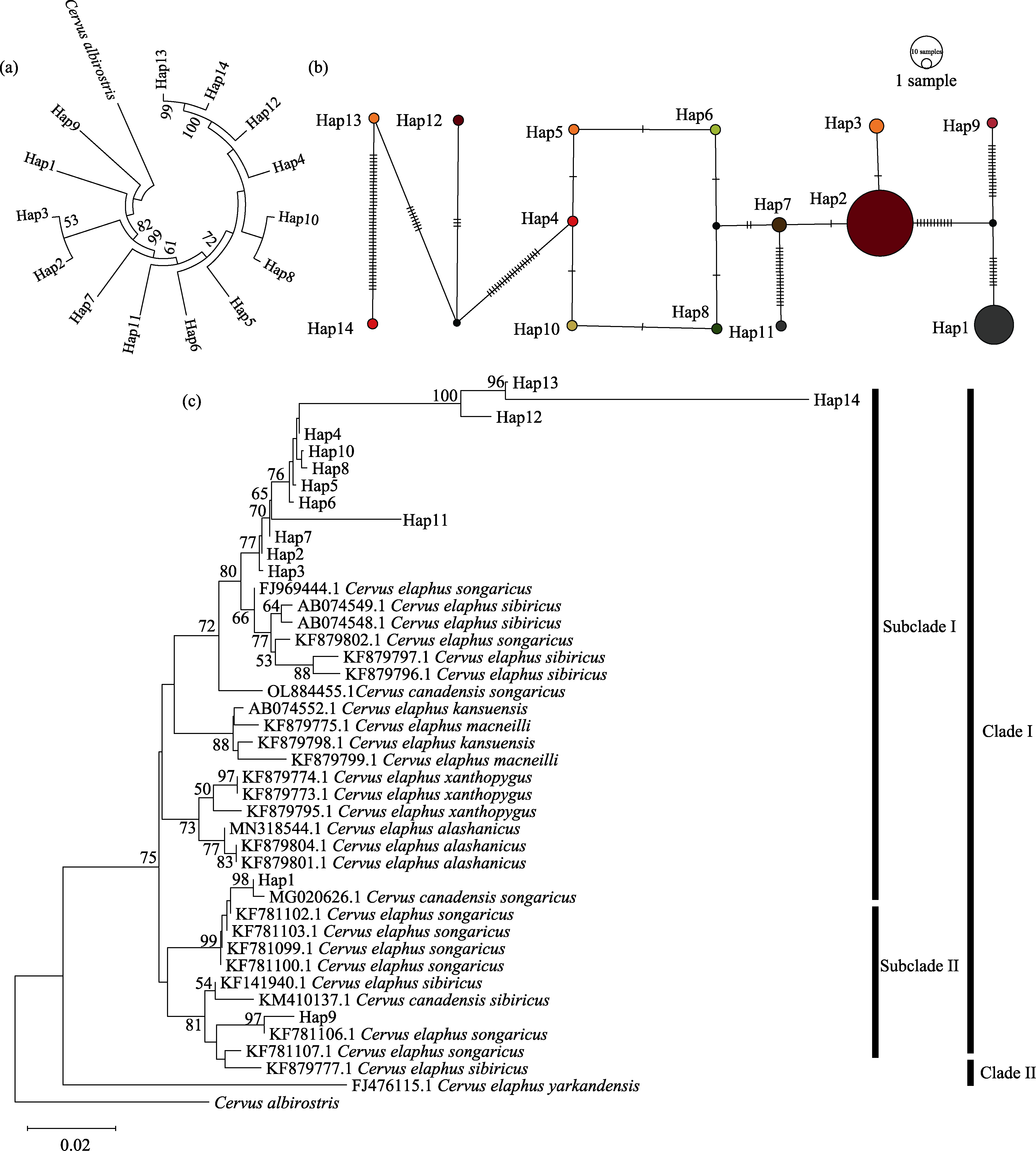
Fig. 6 Genetic structure and phylogenetic relationships of the red deer population in the Ebinur Lake Wetland National Nature Reserve. (a) Phylogenetic tree of the D-loop haplotype system; (b) Network diagram of the D-loop haplotype system; (c) Phylogenetic tree of D-loop haplotype in red deer from Ebinur Lake Wetland National Nature Reserve and Chinese red deer subspecies; In Figure c, Clade I, Clade II, Subclade I, and Subclade II correspond sequentially to the red deer populations of the Tianshan-Altai-Gansu-Sichuan-Northeast-Alashan and Ebinur Lake Wetland National Nature Reserve, the Tarim red deer, the red deer populations of the Tianshan-Altai-Gansu-Sichuan-Northeast-Alashan and Ebinur Lake Wetland National Nature Reserve, and the red deer populations of the Tianshan-Altai and Ebinur Lake Wetland National Nature Reserve.
| ABH | ALT | TS | DB | ALS | GS | SC | |
|---|---|---|---|---|---|---|---|
| ALT | 0.013 | ||||||
| TS | 0.015 | 0.005 | |||||
| DB | 0.032 | 0.018 | 0.014 | ||||
| ALS | 0.046 | 0.018 | 0.022 | 0.007 | |||
| GS | 0.036 | 0.018 | 0.023 | 0.033 | 0.025 | ||
| SC | 0.039 | 0.019 | 0.025 | 0.031 | 0.022 | -0.001 | |
| TLM | 0.120 | 0.091 | 0.093 | 0.080 | 0.087 | 0.087 | 0.082 |
Table 3 D-loop genetic distance between red deer and Chinese red deer subspecies in Ebinur Lake Wetland National Nature Reserve
| ABH | ALT | TS | DB | ALS | GS | SC | |
|---|---|---|---|---|---|---|---|
| ALT | 0.013 | ||||||
| TS | 0.015 | 0.005 | |||||
| DB | 0.032 | 0.018 | 0.014 | ||||
| ALS | 0.046 | 0.018 | 0.022 | 0.007 | |||
| GS | 0.036 | 0.018 | 0.023 | 0.033 | 0.025 | ||
| SC | 0.039 | 0.019 | 0.025 | 0.031 | 0.022 | -0.001 | |
| TLM | 0.120 | 0.091 | 0.093 | 0.080 | 0.087 | 0.087 | 0.082 |
| 位点 Locus | 基因数 N | 等位基因数 Na | 期望杂合度 He | 无限等位基因突变模型 I.A.M. | 两阶段突变模型 T.P.M. | 逐步突变模型 S.M.M. | |||
|---|---|---|---|---|---|---|---|---|---|
| 平均期望杂合度 Heq | P | 平均期望杂合度 Heq | P | 平均期望杂合度 Heq | P | ||||
| BM4208 | 146 | 16 | 0.862 | 0.817 | 0.250 | 0.871 | 0.315 | 0.904 | 0.033* |
| BM5004 | 146 | 7 | 0.527 | 0.600 | 0.259 | 0.687 | 0.066 | 0.762 | 0.007* |
| T530 | 146 | 16 | 0.903 | 0.822 | 0.014* | 0.871 | 0.122 | 0.903 | 0.449 |
| T507 | 146 | 11 | 0.858 | 0.735 | 0.039* | 0.803 | 0.142 | 0.854 | 0.465 |
| CSSM19 | 146 | 12 | 0.873 | 0.749 | 0.020* | 0.821 | 0.090 | 0.868 | 0.494 |
| T123 | 146 | 14 | 0.774 | 0.797 | 0.284 | 0.851 | 0.039* | 0.888 | 0.000** |
| T501 | 146 | 23 | 0.800 | 0.885 | 0.035* | 0.919 | 0.002* | 0.936 | 0.002* |
| BM1225 | 146 | 19 | 0.876 | 0.855 | 0.400 | 0.896 | 0.166 | 0.920 | 0.008* |
| BM848 | 146 | 13 | 0.644 | 0.776 | 0.072 | 0.839 | 0.000** | 0.878 | 0.000** |
| BM6506 | 146 | 10 | 0.705 | 0.715 | 0.368 | 0.783 | 0.097 | 0.839 | 0.003* |
Table 4 Detection of population bottlenecks of red deer in the Ebinur Lake Wetland National Nature Reserve
| 位点 Locus | 基因数 N | 等位基因数 Na | 期望杂合度 He | 无限等位基因突变模型 I.A.M. | 两阶段突变模型 T.P.M. | 逐步突变模型 S.M.M. | |||
|---|---|---|---|---|---|---|---|---|---|
| 平均期望杂合度 Heq | P | 平均期望杂合度 Heq | P | 平均期望杂合度 Heq | P | ||||
| BM4208 | 146 | 16 | 0.862 | 0.817 | 0.250 | 0.871 | 0.315 | 0.904 | 0.033* |
| BM5004 | 146 | 7 | 0.527 | 0.600 | 0.259 | 0.687 | 0.066 | 0.762 | 0.007* |
| T530 | 146 | 16 | 0.903 | 0.822 | 0.014* | 0.871 | 0.122 | 0.903 | 0.449 |
| T507 | 146 | 11 | 0.858 | 0.735 | 0.039* | 0.803 | 0.142 | 0.854 | 0.465 |
| CSSM19 | 146 | 12 | 0.873 | 0.749 | 0.020* | 0.821 | 0.090 | 0.868 | 0.494 |
| T123 | 146 | 14 | 0.774 | 0.797 | 0.284 | 0.851 | 0.039* | 0.888 | 0.000** |
| T501 | 146 | 23 | 0.800 | 0.885 | 0.035* | 0.919 | 0.002* | 0.936 | 0.002* |
| BM1225 | 146 | 19 | 0.876 | 0.855 | 0.400 | 0.896 | 0.166 | 0.920 | 0.008* |
| BM848 | 146 | 13 | 0.644 | 0.776 | 0.072 | 0.839 | 0.000** | 0.878 | 0.000** |
| BM6506 | 146 | 10 | 0.705 | 0.715 | 0.368 | 0.783 | 0.097 | 0.839 | 0.003* |
| [1] | Avise JC, Arnold J, Ball RM, Bermingham E, Lamb T, Neigel JE, Reeb CA, Saunders NC (1987) Intraspecific phylogeography: The mitochondrial DNA bridge between population genetics and systematics.Annual Review of Ecology and Systematics, 18, 489-522. |
| [2] |
Bellemain E, Swenson JE, Tallmon D, Brunberg S, Taberlet P (2005) Estimating population size of elusive animals with DNA from hunter-collected feces: Four methods for brown bears. Conservation Biology, 19, 150-161.
DOI URL |
| [3] |
Bishop MD, Kappes SM, Keele JW, Stone RT, Sunden SL, Hawkins G A, Toldo SS, Fries R, Grosz MD, Yoo J (1994) A genetic linkage map for cattle. Genetics, 136, 619-639.
DOI PMID |
| [4] |
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. American Journal of Human Genetics, 32, 314-331.
PMID |
| [5] |
Brehm A, Harris DJ, Hernández M, Cabrera VM, Larruga JM, Pinto FM, González AM (2001) Structure and evolution of the mitochondrial DNA complete control region in the Drosophila subobscura subgroup. Insect Molecular Biology, 10, 573-578.
PMID |
| [6] | Chen SJ, Hou P, Li WH, Li H (2006) Comprehensive Scientific Survey of the Ebinur Lake Wetland Nature Reserve in Xinjiang. Xinjiang Science and Technology Press, Urumqi. (in Chinese) |
| [陈蜀江, 侯平, 李文华, 李虎 (2006) 新疆艾比湖湿地自然保护区综合科学考察. 新疆科学技术出版社, 乌鲁木齐.] | |
| [7] | Cheng L, Mahmut H, Shi QD, Chen LH, Wang DH, Rahmutulla A, Ezizian N, Peng JB (2016) Habitat suitability assessment for gazellasubgutturosa in Ebinur Lake Wetland Nature Reserve. Acta Ecologica Sinica, 36, 6583-6590. (in Chinese with English abstract) |
| [程亮, 马合木提·哈力克, 师庆东, 陈丽华, 王德虎, 热木图拉·阿卜杜克热木, 艾孜孜江·乃比, 彭佳宾 (2016) 艾比湖湿地自然保护区鹅喉羚生境适宜性评价. 生态学报, 36, 6583-6590.] | |
| [8] |
Ellegren H (2004) Microsatellites: Simple sequences with complex evolution. Nature Reviews Genetics, 5, 435-445.
DOI PMID |
| [9] |
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: A simulation study. Molecular Ecology, 14, 2611-2620.
DOI PMID |
| [10] |
Galtier N, Nabholz B, Glémin S, Hurst GDD (2009) Mitochondrial DNA as a marker of molecular diversity: A reappraisal. Molecular Ecology, 18, 4541-4550.
DOI PMID |
| [11] | Gao H, Qiao FJ, Teng LW, Li JL, Yu MQ, Liu ZS (2020) Genetic diversity and structure of the Alashan red deer (Cervus elaphus alashanicus) in Helan Mountains, China. Acta Theriologica Sinica, 40, 458-466. (in Chinese with English abstract) |
| [高惠, 乔付杰, 滕丽微, 李俊乐, 余梦琦, 刘振生 (2020) 马鹿阿拉善亚种遗传多样性和群体结构. 兽类学报, 40, 458-466.] | |
| [12] | Hu HJ (2016) Studies on Molecular Ecology and Nutritional Ecology of Tibetan Red Deer (Cervus wallichii) in China. PhD dissertation, Northeast Forestry University, Harbin. (in Chinese with English abstract) |
| [胡贺娇 (2016) 西藏马鹿(Cervus wallichii)分子生态学与营养生态学研究. 博士学位论文, 东北林业大学, 哈尔滨.] | |
| [13] |
Hu HJ, Xing B, Yang M, Mpemba H, Lv ZH, Zhang MH (2018) Population and genetic diversity of Tibetan red deer based on fecal DNA. Journal of Forestry Research, 29, 227-232.
DOI URL |
| [14] |
Jakobsson M, Rosenberg NA (2007) CLUMPP: A cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics, 23, 1801-1806.
DOI PMID |
| [15] | Johns GC, Avise JC (1998) A comparative summary of genetic distances in the vertebrates from the mitochondrial cytochrome b gene. Molecular Biology and Evolution, 15, 1481-1490. |
| [16] |
Kalinowski ST, Taper ML, Marshall TC (2007) Revising how the computer program Cervus accommodates genotyping error increases success in paternity assignment. Molecular Ecology, 16, 1099-1106.
DOI PMID |
| [17] |
Kim KI, Lee JH, Li K, Zhang YP, Lee SS, Gongora J, Moran C (2002) Phylogenetic relationships of Asian and European pig breeds determined by mitochondrial DNA D-loop sequence polymorphism. Animal Genetics, 33, 19-25.
DOI PMID |
| [18] |
Kuehn R, Schroeder W, Pirchner F, Rottmann O (2003) Genetic diversity, gene flow and drift in Bavarian red deer populations (Cervus elaphus). Conservation Genetics, 4, 157-166.
DOI |
| [19] | Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Custal X version 2.0. Bioinformatics, 23, 2947-2948. |
| [20] |
Leigh JW, Bryant D (2015) POPART: Full-feature software for haplotype network construction. Methods in Ecology and Evolution, 6, 1110-1116.
DOI URL |
| [21] |
Lenney WC, Serfass TL, Cogan R, Rhodes OE Jr (2002) Microsatellite variation in the reintroduced Pennsylvania elk herd. Molecular Ecology, 11, 1299-1310.
PMID |
| [22] | Li YH, Chu XZ, Jin HL (2006) Study on changes of hydrological characteristics of Ebinur Lake basin in Xinjiang. Journal of China Hydrology, 26(5), 68-71. (in Chinese with English abstract) |
| [李艳红, 楚新正, 金海龙 (2006) 新疆艾比湖流域水文特征分析. 水文, 26(5), 68-71.] | |
| [23] | Li Z, Guo JH, Liu XX, Zhang MH (2023) Population genetic diversity of red deer in the southern of the Greater Khingan Mountains. Chinese Journal of Wildlife, 44, 548-555. (in Chinese with English abstract) |
| [李峥, 郭金昊, 刘鑫鑫, 张明海 (2023) 大兴安岭南麓东北马鹿种群遗传多样性分析. 野生动物学报, 44, 548-555.] | |
| [24] | Liu YH, Zhang MH (2011) Population genetic diversity in Tibet red deer (Cervus elaphus wallichi) revealed by mitochondrial Cytb gene analysis. Acta Ecologica Sinica, 31, 1976-1981. (in Chinese with English abstract) |
| [刘艳华, 张明海 (2011) 基于线粒体Cytb基因的西藏马鹿种群遗传多样性研究. 生态学报, 31, 1976-1981.] | |
| [25] |
Luikart G, Allendorf FW, Cornuet JM, Sherwin WB (1998) Distortion of allele frequency distributions provides a test for recent population bottlenecks. The Journal of Heredity, 89, 238-247.
DOI URL |
| [26] | Mahmut H (2012) Xinjiang Tarim Red Deer. Xinjiang University Press, Urumqi. (in Chinese) |
| [ 马合木提·哈力克 (2012) 新疆塔里木马鹿. 新疆大学出版社, 乌鲁木齐.] | |
| [27] | Mahmut H, Ganzorig S, Onuma M, Masuda R, Suzuki M, Ohtaishi N (2001) A preliminary study of the genetic diversity of Xinjiang Tarim red deer (Cervus elaphus yarkandensis) using the microsatellite DNA method. The Japanese Journal of Veterinary Research, 49, 231-237. |
| [28] |
Miller WL, Edson J, Pietrandrea P, Miller-Butterworth C, Walter WD (2019) Identification and evaluation of a core microsatellite panel for use in white-tailed deer (Odocoileus virginianus). BMC Genetics, 20, 49.
DOI PMID |
| [29] | Park SDE (2001) Trypanotolerance in West African Cattle and the Population Genetic Effects of Selection. PhD dissertation, University of Dublin, Dublin. |
| [30] |
Peakall R, Smouse PE (2012) GenAlEx 6.5: Genetic analysis in Excel. Population genetic software for teaching and research—An update. Bioinformatics, 28, 2537-2539.
PMID |
| [31] |
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics, 155, 945-959.
DOI PMID |
| [32] |
Putman AI, Carbone I (2014) Challenges in analysis and interpretation of microsatellite data for population genetic studies. Ecology and Evolution, 4, 4399-4428.
DOI PMID |
| [33] |
Raymond M, Rousset F (1995) GENEPOP (version 1.2): Population genetics software for exact tests and ecumenicism. Journal of Heredity, 86, 248-249.
DOI URL |
| [34] |
Rosenberg NA (2004) Distruct: A program for the graphical display of population structure. Molecular Ecology Notes, 4, 137-138.
DOI URL |
| [35] | Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6:DNA sequence polymorphism analysis of large data sets. Molecular Biology and Evolution, 34, 3299-3302. |
| [36] |
Selkoe KA, Toonen RJ (2006) Microsatellites for ecologists: A practical guide to using and evaluating microsatellite markers. Ecology Letters, 9, 615-629.
DOI PMID |
| [37] | Shamshinur M, Shamshidin A, Tajigvl T, Ning LQ, Mahmut H (2016) Population estimate of red deer in Ebinur Lake National Wetland Nature Reserve, Xinjiang. Journal of Wildlife Biology, 37, 287-292. (in Chinese with English abstract) |
| [先木西努·莫合德, 夏米西丁·阿不都热依木, 塔吉古丽·吐热甫, 宁礼群, 马合木提·哈力克(2016) 艾比湖湿地国家级自然保护区马鹿种群资源调查. 野生动物学报, 37, 287-292.] | |
| [38] |
Taberlet P, Griffin S, Goossens B, Questiau S, Manceau V, Escaravage N, Waits LP, Bouvet J (1996) Reliable genotyping of samples with very low DNA quantities using PCR. Nucleic Acids Research, 24, 3189-3194.
DOI PMID |
| [39] | Tayerjan M, Tajigul T, Buweihailigiemu A, Subinur E, Mahmut H (2018) Influence of environmental factors on genetic diversity of Tarim red deer. Chinese Journal of Wildlife, 39, 754-760. (in Chinese with English abstract) |
| [塔依尔江·麦麦提, 塔吉古丽·吐热甫, 布威海丽且姆·阿巴拜科日, 苏比奴尔·艾力, 马合木提·哈力克(2018) 环境因子对塔里木马鹿种群遗传多样性的影响. 野生动物学报, 39, 754-760.] | |
| [40] |
Tian XM, Zhang MH (2023) Genetic diversity of wapiti in Northeast China based on fecal DNA. Acta Theriologica Sinica, 43, 41-49. (in Chinese with English abstract)
DOI |
|
[田新民, 张明海 (2023) 基于粪便DNA的东北马鹿遗传多样性. 兽类学报, 43, 41-49.]
DOI |
|
| [41] |
Tu JF, Xu JP, Wang HL, Li YQ, Xing XM (2018) Genetic differentiation of five subspecies wapiti in China based on mitochondrial DNA control region sequences. Acta Agriculturae Boreali-Sinica, 33(S1), 79-83. (in Chinese)
DOI |
|
[涂剑锋, 徐佳萍, 王洪亮, 李一清, 邢秀梅 (2018) 基于线粒体DNA控制区序列分析我国马鹿5个亚种的遗传分化. 华北农学报, 33(S1), 79-83.]
DOI |
|
| [42] |
Van Oosterhout C, Hutchinson WF, Wills DPM, Shipley P (2004) Micro-checker: Software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes, 4, 535-538.
DOI URL |
| [43] | Wu JL, Liu JJ, Wang SM (2004) Climatic change record from stable isotopes in Lake Aibi, Xinjiang during the past 1500 years. Quaternary Sciences, 24, 585-590. (in Chinese with English abstract) |
| [吴敬禄, 刘建军, 王苏民 (2004) 近1,500年来新疆艾比湖同位素记录的气候环境演化特征. 第四纪研究, 24, 585-590.] | |
| [44] | Yang XD, Lü GH, Tian YH, Yang J, Zhang XM (2009) Ecological groups of plants in Ebinur Lake Wetland Nature Reserve of Xinjiang. Chinese Journal of Ecology, 28, 2489-2494. (in Chinese with English abstract) |
| [杨晓东, 吕光辉, 田幼华, 杨军, 张雪梅 (2009) 新疆艾比湖湿地自然保护区植物的生态分组. 生态学杂志, 28, 2489-2494.] | |
| [45] | Zhang L, Gun SB, Lei TY, Liu LX, Qin DW, Zhao SF (2010) Analysis of genetic diversity and phylogeny of five Chinese subspecies of wapiti using mitochondrial Cytb complete sequence. Acta Agriculturae Boreali-Sinica, 25(4), 12-16. (in Chinese with English abstract) |
|
[张丽, 滚双宝, 雷天云, 刘丽霞, 秦大伟, 赵世峰 (2010) 应用mtDNA Cytb基因全序列分析中国5个马鹿群体的遗传多样性和系统发育. 华北农学报, 25(4), 12-16.]
DOI |
|
| [46] | Zhang ZX, Chen YX, Xi XY, Xiao RR, Zhang Q (2022) Genetic diversity analysis of Gansu red deer using microsatellite markers. Plateau Science Research, 6(1), 63-75. (in Chinese) |
| [张振祥, 陈艳霞, 习向玉, 肖荣荣, 张琪 (2022) 应用微卫星标记分析山丹军马场甘肃马鹿遗传多样性. 高原科学研究, 6(1), 63-75.] | |
| [47] | Zheng SW (2003) Morphological variation and taxonomic status of red deer in the Tianshan Mountains. Chinese Journal of Zoology, 38(2), 58-63. (in Chinese with English abstract) |
| [郑生武 (2003) 天山马鹿的形态变异及其分类地位探讨. 动物学杂志, 38(2), 58-63.] | |
| [48] | Zhou CL (2015) The Study on Population Size, Genetic Structure, Home Range and Phylogeny of Tianshan Red Deer (Cervus elaphus songaricus). PhD dissertation, Xinjiang University, Urumqi. (in Chinese with English abstract) |
| [周璨林 (2015) 天山马鹿种群数量、遗传结构、家域及系统发育关系研究. 博士学位论文, 新疆大学, 乌鲁木齐.] |
| [1] | Yuan Jiang, Beixi Huang, Xueyuan Jia, Si Liang, Yutong Xie, Ping Fan, Gang Song. An approach for estimating haplotype richness from DNA sequences with unequal lengths [J]. Biodiv Sci, 2026, 34(1): 25263-. |
| [2] | Ruxiao Wang, Boyang Shi, Da Pan, Hongying Sun. Biodiversity patterns and conservation gaps of the endemic freshwater crab genus Sinopotamon in China [J]. Biodiv Sci, 2025, 33(8): 25123-. |
| [3] | Zhicheng Zhou, Tianling Cao, Ruyao Liu, Qiqi Ding, Ke Ma, Liping Yang, Chuanjiang Zhou, Guoxing Nie, Yongtao Tang. Genetic diversity and structure of Pseudorasbora parva population in the Yellow River basin based on the mitochondrial COI gene [J]. Biodiv Sci, 2025, 33(8): 24501-. |
| [4] | Qilong Yu, Minhui Hao, Huaijiang He, Chunyu Zhang, Xiuhai Zhao. Relationships of biodiversity and productivity change with forest succession in Changbai Mountains: Insights from species, traits, and phylogeny [J]. Biodiv Sci, 2025, 33(8): 25060-. |
| [5] | Ping Fan, Huan Wang, Zhixin Wen, Gang Song, Fuming Lei. Impact of climatic factors on the genetic diversity-species area relationship of birds [J]. Biodiv Sci, 2025, 33(8): 25072-. |
| [6] | Ruixiang Xue, Xuerong Ma, Jiongwen Wu, Aijun Liu, Xiquan Zhang, Congliang Ji, Yingshan Yin, Weijian Zhu, Qingbin Luo. Genetic diversity and genetic structure of Zhongshan partridge duck populations [J]. Biodiv Sci, 2025, 33(8): 24592-. |
| [7] | Ping Fan, Zhixin Wen, Gang Song. The effect of climatic factors and anthropogenic activities on different genetic diversity indicators of amphibians and mammals [J]. Biodiv Sci, 2025, 33(8): 25022-. |
| [8] | Jiachen Wang, Tangjun Xu, Wei Xu, Gaoji Zhang, Yijin You, Honghua Ruan, Hongyi Liu. Impact of urban landscape pattern on the genetic structure of Thereuopoda clunifera population in Nanjing, China [J]. Biodiv Sci, 2025, 33(1): 24251-. |
| [9] | Kexin Cao, Jingwen Wang, Guo Zheng, Pengfeng Wu, Yingbin Li, Shuyan Cui. Effects of precipitation regime change and nitrogen deposition on soil nematode diversity in the grassland of northern China [J]. Biodiv Sci, 2024, 32(3): 23491-. |
| [10] | Linjun He, Wenjing Yang, Yuhao Shi, Kezhemo Ashuo, Yu Fan, Guoyan Wang, Jingji Li, Songlin Shi, Guihua Yi, Peihao Peng. Effects of plant community phylogeny and functional diversity on Ageratina adenophora invasion under fire disturbance [J]. Biodiv Sci, 2024, 32(11): 24269-. |
| [11] | Shiyi Long, Bobo Zhang, Yuchen Xia, Yangfan Fei, Yani Meng, Bingwei Lü, Yueqing Song, Pu Zheng, Taoran Guo, Jian Zhang, Shaopeng Li. Effects of diversity and temporal stability of native communities on the biomass of invasive species Solidago canadensis [J]. Biodiv Sci, 2024, 32(11): 24263-. |
| [12] | Chen Feng, Jie Zhang, Hongwen Huang. Parallel situ conservation: A new plant conservation strategy to integrate in situ and ex situ conservation of plants [J]. Biodiv Sci, 2023, 31(9): 23184-. |
| [13] | Qingduo Li, Dongmei Li. Analysis for the prevalence of global bat-borne Bartonella [J]. Biodiv Sci, 2023, 31(9): 23166-. |
| [14] | Hailing Qi, Pengzhen Fan, Yuehua Wang, Jie Liu. Genetic diversity and population structure of Juglans regia from six provinces in northern China [J]. Biodiv Sci, 2023, 31(8): 23120-. |
| [15] | Zhenjie Zhan, Chao Zhang, Minhao Chen, Jiadong Wang, Aihua Fu, Yuwei Fan, Xiaofeng Luan. DNA metabarcoding-based winter diet analysis of Eurasian otter (Lutra lutra) in the northern Greater Khingan Mountains [J]. Biodiv Sci, 2023, 31(6): 22586-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn