



生物多样性 ›› 2016, Vol. 24 ›› Issue (3): 296-303. DOI: 10.17520/biods.2015259 cstr: 32101.14.biods.2015259
胡亚妮1,2, 张宗文2,3,*( ), 吴斌2, 高佳2, 李艳琴1,*(
), 吴斌2, 高佳2, 李艳琴1,*( )
)
收稿日期:2015-09-21
接受日期:2016-03-02
出版日期:2016-03-20
发布日期:2016-04-05
通讯作者:
张宗文,李艳琴
基金资助:
Yani Hu1,2, Zongwen Zhang2,3,*( ), Bin Wu2, Jia Gao2, Yanqin Li1,*(
), Bin Wu2, Jia Gao2, Yanqin Li1,*( )
)
Received:2015-09-21
Accepted:2016-03-02
Online:2016-03-20
Published:2016-04-05
Contact:
Zhang Zongwen,Li Yanqin
摘要:
荞麦起源于我国, 演化形成了丰富的物种和遗传多样性。为了有效研究和利用荞麦及其野生种资源, 以从四川、甘肃、贵州等地采集的荞麦属(Fagopyrum)10个种(含变种、亚种和复合体种)共71份材料为对象, 通过ITS和叶绿体ndhF-rpl32序列分析, 利用MEGA5.0构建系统进化树, 探讨了荞麦种内及种间亲缘关系。结果表明, 在ITS序列矩阵中, 序列长度为725 bp, 信息位点为150个, 占序列总长度的20.7%; 在ndhF-rpl32序列矩阵中, 序列长度为940 bp, 信息位点为158个, 占序列总长度的16.8%。由ITS序列和ndhF-rpl32序列构建的两个进化树都可以将71份材料分为大粒荞麦种组和小粒荞麦种组; 其中, 大粒荞麦种组包括栽培苦荞和米苦荞(F. tataricum)、金荞复合体(F. cymosum complex)、栽培甜荞(F. esculentum)和野生甜荞(F. esculentum ssp. ancestralis); 小粒荞麦种组包括齿翅野荞(F. gracilipes var. odontopterum)、疏穗小野荞(F. leptopodum var. grossii)、小野荞(F. leptopodum)、密毛野荞(F. densovillosum)、细柄野荞(F. gracilipes)和硬枝万年荞(F. urophyllum)。而ndhF-rpl32序列构建的系统发育树还能区分栽培甜荞和野生甜荞, 具有更好的聚类效果。另外, 与栽培甜荞相比, 金荞复合体与栽培苦荞的亲缘关系更近。该研究为荞麦属种的分类和条形码研究提供了一定的科学依据。
胡亚妮, 张宗文, 吴斌, 高佳, 李艳琴 (2016) 基于ITS和ndhF-rpl32序列的荞麦种间亲缘关系分析. 生物多样性, 24, 296-303. DOI: 10.17520/biods.2015259.
Yani Hu, Zongwen Zhang, Bin Wu, Jia Gao, Yanqin Li (2016) Genetic relationships of buckwheat species based on the sequence analysis of ITS and ndhF-rpl32. Biodiversity Science, 24, 296-303. DOI: 10.17520/biods.2015259.
| 编号 Collecting number | 原产地 Origin | 代号 Code | 编号 Collecting number | 原产地 Origin | 代号 Code | ||
|---|---|---|---|---|---|---|---|
| 齿翅野荞 Fagopyrum gracilipes var. odontopterum | 细柄野荞 F. gracilipes | ||||||
| YXCC201211030 | 四川越西 Yuexi, Sichuan | a1 | XDXB201211028 | 四川喜德 Xide, Sichuan | g1 | ||
| ZJCC2012102601 | 四川昭觉 Zhaojue, Sichuan | a2 | ZJXB2012102602 | 四川昭觉 Zhaojue, Sichuan | g2 | ||
| ZJCC2012102603 | 四川昭觉 Zhaojue, Sichuan | a3 | ZJXB2012102605 | 四川昭觉 Zhaojue, Sichuan | g3 | ||
| ZJCC2012102604 | 四川昭觉 Zhaojue, Sichuan | a4 | ZJXB2012102703 | 四川昭觉 Zhaojue, Sichuan | g4 | ||
| ZJCC2012102704 | 四川昭觉 Zhaojue, Sichuan | a5 | MGXB2012102901 | 四川美姑 Meigu, Sichuan | g5 | ||
| BTCC2012102706 | 四川布拖 Butuo, Sichuan | a6 | HLXB2012110303 | 四川会理 Huili, Sichuan | g6 | ||
| ZJCC2012102801 | 四川昭觉 Zhaojue, Sichuan | a7 | HLXB2012110306 | 四川会理 Huili, Sichuan | g7 | ||
| MGCC2012102902 | 四川美姑 Meigu, Sichuan | a8 | HDXB2012110404 | 四川会东 Huidong, Sichuan | g8 | ||
| LBCC2012103002 | 四川雷波 Leibo, Sichuan | a9 | HDXB2012110501 | 四川会东 Huidong, Sichuan | g9 | ||
| HLCC2012110304 | 四川会理 Huili, Sichuan | a10 | HDCC2012110504 | 四川会东 Huidong, Sichuan | g10 | ||
| HLCC2012110305 | 四川会理 Huili, Sichuan | a11 | HDXB2012110504 | 四川会东 Huidong, Sichuan | g11 | ||
| HDCC2012110502 | 四川会东 Huidong, Sichuan | a12 | HLXB2012110301 | 四川会理 Huili, Sichuan | g12 | ||
| HDCC2012110503 | 四川会东 Huidong, Sichuan | a13 | MNXB201210006 | 四川冕宁 Mianning, Sichuan | g13 | ||
| YXCC201211040 | 四川越西 Yuexi, Sichuan | a14 | XDXB201210014 | 四川喜德 Xide, Sichuan | g14 | ||
| YXCC201211044 | 四川越西 Yuexi, Sichuan | a15 | XDXB201211046 | 四川喜德 Xide, Sichuan | g15 | ||
| YYCC201210012 | 四川盐源 Yanyuan, Sichuan | a16 | YXXB201211031 | 四川越西 Yuexi, Sichuan | g16 | ||
| HDCC2012110402 | 四川越西 Yuexi, Sichuan | a17 | 金荞复合体 F. cymosum complex | ||||
| 栽培苦荞 F. tataricum | XDJQ201210004 | 四川喜德 Xide, Sichuan | h1 | ||||
| XDKQ201210003 | 四川喜德 Xide, Sichuan | b1 | MNJQ201210007 | 四川冕宁 Mianning, Sichuan | h2 | ||
| XDKQ201210005 | 四川喜德 Xide, Sichuan | b2 | XDKQ201210017 | 西川喜德 Xide, Sichuan | h3 | ||
| XDKQ201210013 | 四川喜德 Xide, Sichuan | b3 | XDJQ201210022 | 西川喜德 Xide, Sichuan | h4 | ||
| XDKQ201210019 | 四川喜德 Xide, Sichuan | b4 | YXJQ201211029 | 四川越西 Yuexi, Sichuan | h5 | ||
| XDKQ201210020 | 四川喜德 Xide, Sichuan | b5 | YXJQ201211039 | 四川越西 Yuexi, Sichuan | h6 | ||
| YXKQ201211032 | 四川越西 Yuexi, Sichuan | b6 | XDJQ201211045 | 四川喜德 Xide, Sichuan | h7 | ||
| YXKQ201211033 | 四川越西 Yuexi, Sichuan | b7 | DCJQ2012110302 | 四川德昌 Dechang, Sichuan | h8 | ||
| 疏穗小野荞 F. leptopodum var.grossii | PGJQ2012110601 | 四川普格 Puge, Sichuan | h9 | ||||
| GLSS201211037 | 四川甘洛 Ganluo, Sichuan | c1 | DCJQ2012110202 | 四川德昌 Dechang, Sichuan | h10 | ||
| GLSS201211038 | 四川甘洛 Ganluo, Sichuan | c2 | HDJQ2012110403 | 四川会东 Huidong, Sichuan | h11 | ||
| GLSS201211050 | 四川甘洛 Ganluo, Sichuan | c3 | XDKQ201210018 | 四川喜德 Xide, Sichuan | h12 | ||
| GLSS201211052 | 四川甘洛 Ganluo, Sichuan | c4 | YXJQ201211043 | 四川越西 Yuexi, Sichuan | h13 | ||
| YXSS201211051 | 四川越西 Yuexi, Sichuan | c5 | 硬枝万年荞 F. urophyllum | ||||
| 野生甜荞 F. esculentum ssp. ancestralis | YYYZ201210011 | 四川盐源 Yanyuan, Sichuan | i1 | ||||
| XCYTQ201210008 | 四川西昌 Xichang, Sichuan | d1 | LBYZ2012103004 | 四川雷波 Leibo, Sichuan | i2 | ||
| LBTJ2012103001 | 四川雷波 Leibo, Sichuan | d2 | MGYZ2012102903 | 四川美姑 Meigu, Sichuan | i3 | ||
| 小野荞 F. leptopodum | 栽培甜荞 F. esculentum | ||||||
| XDXY201210021 | 四川喜德 Xide, Sichuan | e1 | 00000664 | 甘肃武威 Wuwei, Gansu | j1 | ||
| MNXY201210024 | 四川冕宁 Mianning, Sichuan | e2 | 00000906 | 贵州威宁 Weining, Guizhou | j2 | ||
| 密毛野荞 F. densovillosum | 米苦荞 F. tataricum | ||||||
| MGMM2012102804 | 四川美姑 Meigu, Sichuan | f1 | YXXMQ201211034 | 四川越西 Yuexi, Sichuan | k1 | ||
| LBMM2012103003 | 四川雷波 Leibo, Sichuan | f2 | YXXMQ201211042 | 四川越西 Yuexi, Sichuan | k2 | ||
表1 供试的荞麦材料名称和来源
Table 1 The Latin names and sources of the tested Fagopyrum materials
| 编号 Collecting number | 原产地 Origin | 代号 Code | 编号 Collecting number | 原产地 Origin | 代号 Code | ||
|---|---|---|---|---|---|---|---|
| 齿翅野荞 Fagopyrum gracilipes var. odontopterum | 细柄野荞 F. gracilipes | ||||||
| YXCC201211030 | 四川越西 Yuexi, Sichuan | a1 | XDXB201211028 | 四川喜德 Xide, Sichuan | g1 | ||
| ZJCC2012102601 | 四川昭觉 Zhaojue, Sichuan | a2 | ZJXB2012102602 | 四川昭觉 Zhaojue, Sichuan | g2 | ||
| ZJCC2012102603 | 四川昭觉 Zhaojue, Sichuan | a3 | ZJXB2012102605 | 四川昭觉 Zhaojue, Sichuan | g3 | ||
| ZJCC2012102604 | 四川昭觉 Zhaojue, Sichuan | a4 | ZJXB2012102703 | 四川昭觉 Zhaojue, Sichuan | g4 | ||
| ZJCC2012102704 | 四川昭觉 Zhaojue, Sichuan | a5 | MGXB2012102901 | 四川美姑 Meigu, Sichuan | g5 | ||
| BTCC2012102706 | 四川布拖 Butuo, Sichuan | a6 | HLXB2012110303 | 四川会理 Huili, Sichuan | g6 | ||
| ZJCC2012102801 | 四川昭觉 Zhaojue, Sichuan | a7 | HLXB2012110306 | 四川会理 Huili, Sichuan | g7 | ||
| MGCC2012102902 | 四川美姑 Meigu, Sichuan | a8 | HDXB2012110404 | 四川会东 Huidong, Sichuan | g8 | ||
| LBCC2012103002 | 四川雷波 Leibo, Sichuan | a9 | HDXB2012110501 | 四川会东 Huidong, Sichuan | g9 | ||
| HLCC2012110304 | 四川会理 Huili, Sichuan | a10 | HDCC2012110504 | 四川会东 Huidong, Sichuan | g10 | ||
| HLCC2012110305 | 四川会理 Huili, Sichuan | a11 | HDXB2012110504 | 四川会东 Huidong, Sichuan | g11 | ||
| HDCC2012110502 | 四川会东 Huidong, Sichuan | a12 | HLXB2012110301 | 四川会理 Huili, Sichuan | g12 | ||
| HDCC2012110503 | 四川会东 Huidong, Sichuan | a13 | MNXB201210006 | 四川冕宁 Mianning, Sichuan | g13 | ||
| YXCC201211040 | 四川越西 Yuexi, Sichuan | a14 | XDXB201210014 | 四川喜德 Xide, Sichuan | g14 | ||
| YXCC201211044 | 四川越西 Yuexi, Sichuan | a15 | XDXB201211046 | 四川喜德 Xide, Sichuan | g15 | ||
| YYCC201210012 | 四川盐源 Yanyuan, Sichuan | a16 | YXXB201211031 | 四川越西 Yuexi, Sichuan | g16 | ||
| HDCC2012110402 | 四川越西 Yuexi, Sichuan | a17 | 金荞复合体 F. cymosum complex | ||||
| 栽培苦荞 F. tataricum | XDJQ201210004 | 四川喜德 Xide, Sichuan | h1 | ||||
| XDKQ201210003 | 四川喜德 Xide, Sichuan | b1 | MNJQ201210007 | 四川冕宁 Mianning, Sichuan | h2 | ||
| XDKQ201210005 | 四川喜德 Xide, Sichuan | b2 | XDKQ201210017 | 西川喜德 Xide, Sichuan | h3 | ||
| XDKQ201210013 | 四川喜德 Xide, Sichuan | b3 | XDJQ201210022 | 西川喜德 Xide, Sichuan | h4 | ||
| XDKQ201210019 | 四川喜德 Xide, Sichuan | b4 | YXJQ201211029 | 四川越西 Yuexi, Sichuan | h5 | ||
| XDKQ201210020 | 四川喜德 Xide, Sichuan | b5 | YXJQ201211039 | 四川越西 Yuexi, Sichuan | h6 | ||
| YXKQ201211032 | 四川越西 Yuexi, Sichuan | b6 | XDJQ201211045 | 四川喜德 Xide, Sichuan | h7 | ||
| YXKQ201211033 | 四川越西 Yuexi, Sichuan | b7 | DCJQ2012110302 | 四川德昌 Dechang, Sichuan | h8 | ||
| 疏穗小野荞 F. leptopodum var.grossii | PGJQ2012110601 | 四川普格 Puge, Sichuan | h9 | ||||
| GLSS201211037 | 四川甘洛 Ganluo, Sichuan | c1 | DCJQ2012110202 | 四川德昌 Dechang, Sichuan | h10 | ||
| GLSS201211038 | 四川甘洛 Ganluo, Sichuan | c2 | HDJQ2012110403 | 四川会东 Huidong, Sichuan | h11 | ||
| GLSS201211050 | 四川甘洛 Ganluo, Sichuan | c3 | XDKQ201210018 | 四川喜德 Xide, Sichuan | h12 | ||
| GLSS201211052 | 四川甘洛 Ganluo, Sichuan | c4 | YXJQ201211043 | 四川越西 Yuexi, Sichuan | h13 | ||
| YXSS201211051 | 四川越西 Yuexi, Sichuan | c5 | 硬枝万年荞 F. urophyllum | ||||
| 野生甜荞 F. esculentum ssp. ancestralis | YYYZ201210011 | 四川盐源 Yanyuan, Sichuan | i1 | ||||
| XCYTQ201210008 | 四川西昌 Xichang, Sichuan | d1 | LBYZ2012103004 | 四川雷波 Leibo, Sichuan | i2 | ||
| LBTJ2012103001 | 四川雷波 Leibo, Sichuan | d2 | MGYZ2012102903 | 四川美姑 Meigu, Sichuan | i3 | ||
| 小野荞 F. leptopodum | 栽培甜荞 F. esculentum | ||||||
| XDXY201210021 | 四川喜德 Xide, Sichuan | e1 | 00000664 | 甘肃武威 Wuwei, Gansu | j1 | ||
| MNXY201210024 | 四川冕宁 Mianning, Sichuan | e2 | 00000906 | 贵州威宁 Weining, Guizhou | j2 | ||
| 密毛野荞 F. densovillosum | 米苦荞 F. tataricum | ||||||
| MGMM2012102804 | 四川美姑 Meigu, Sichuan | f1 | YXXMQ201211034 | 四川越西 Yuexi, Sichuan | k1 | ||
| LBMM2012103003 | 四川雷波 Leibo, Sichuan | f2 | YXXMQ201211042 | 四川越西 Yuexi, Sichuan | k2 | ||
| 种名 Species | ITS (%) | ndhF-rpl32 (%) |
|---|---|---|
| 齿翅野荞 F. gracilipes var. odontopterum | 65.57 | 24.17 |
| 栽培苦荞 F. tataricum | 67.65 | 22.79 |
| 疏穗小野荞 F. leptopodum var. grossii | 65.96 | 22.01 |
| 野生甜荞 F. esculentum ssp. ancestralis | 66.46 | 24.47 |
| 小野荞 F. leptopodum | 66.82 | 23.43 |
| 密毛野荞 F. densovillosum | 65.34 | 24.77 |
| 细柄野荞 F. gracilipes | 65.81 | 23.17 |
| 金荞复合体 F. cymosum complex | 65.80 | 22.75 |
| 硬枝万年荞 F. urophyllum | 63.93 | 24.68 |
| 栽培甜荞 F. esculentum | 65.47 | 24.13 |
| 米苦荞 F. tataricum | 67.95 | 23.40 |
表2 不同荞麦种(含变种、亚种和复合体种)的ITS序列和ndhF-rpl32序列中G+C含量
Table 2 The G+C contents of ITS and ndhF-rpl32 sequences in Fagopyrum species
| 种名 Species | ITS (%) | ndhF-rpl32 (%) |
|---|---|---|
| 齿翅野荞 F. gracilipes var. odontopterum | 65.57 | 24.17 |
| 栽培苦荞 F. tataricum | 67.65 | 22.79 |
| 疏穗小野荞 F. leptopodum var. grossii | 65.96 | 22.01 |
| 野生甜荞 F. esculentum ssp. ancestralis | 66.46 | 24.47 |
| 小野荞 F. leptopodum | 66.82 | 23.43 |
| 密毛野荞 F. densovillosum | 65.34 | 24.77 |
| 细柄野荞 F. gracilipes | 65.81 | 23.17 |
| 金荞复合体 F. cymosum complex | 65.80 | 22.75 |
| 硬枝万年荞 F. urophyllum | 63.93 | 24.68 |
| 栽培甜荞 F. esculentum | 65.47 | 24.13 |
| 米苦荞 F. tataricum | 67.95 | 23.40 |
| 种名 Species | ITS | ndhF-rpl32 |
|---|---|---|
| 齿翅野荞 F. gracilipes var. odontopterum | 0.001 | 0.182 |
| 栽培苦荞 F. tataricum | 0.003 | 0.000 |
| 疏穗小野荞 F. leptopodum var. grossii | 0.002 | 0.002 |
| 野生甜荞 F. esculentum ssp. ancestralis | 0.107 | 0.001 |
| 小野荞 F. leptopodum | 0.062 | 0.021 |
| 密毛野荞 F. densovillosum | 0.003 | 0.018 |
| 细柄野荞 F. gracilipes | 0.001 | 0.099 |
| 金荞复合体 F. cymosum complex | 0.028 | 0.026 |
| 硬枝万年荞 F. urophyllum | 0.165 | 0.014 |
| 栽培甜荞 F. esculentum | 0.110 | 0.000 |
| 米苦荞 F. tataricum | 0.000 | 0.001 |
表3 基于ITS序列和ndhF-rpl32序列计算的荞麦种内遗传距离
Table 3 Distance within Fagopyrum species based on ITS and ndhF-rpl32 sequences
| 种名 Species | ITS | ndhF-rpl32 |
|---|---|---|
| 齿翅野荞 F. gracilipes var. odontopterum | 0.001 | 0.182 |
| 栽培苦荞 F. tataricum | 0.003 | 0.000 |
| 疏穗小野荞 F. leptopodum var. grossii | 0.002 | 0.002 |
| 野生甜荞 F. esculentum ssp. ancestralis | 0.107 | 0.001 |
| 小野荞 F. leptopodum | 0.062 | 0.021 |
| 密毛野荞 F. densovillosum | 0.003 | 0.018 |
| 细柄野荞 F. gracilipes | 0.001 | 0.099 |
| 金荞复合体 F. cymosum complex | 0.028 | 0.026 |
| 硬枝万年荞 F. urophyllum | 0.165 | 0.014 |
| 栽培甜荞 F. esculentum | 0.110 | 0.000 |
| 米苦荞 F. tataricum | 0.000 | 0.001 |
| CC | KQ | SS | TJ | XY | MM | XB | JQ | YZ | TQ | MKQ | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 齿翅野荞(CC) F. gracilipes var. odontopterum | 0.191 | 0.024 | 0.198 | 0.001 | 0.001 | 0.001 | 0.191 | 0.027 | 0.197 | 0.191 | |
| 栽培苦荞(KQ) F. tataricum | 0.157 | 0.185 | 0.037 | 0.191 | 0.190 | 0.191 | 0.007 | 0.188 | 0.039 | 0.001 | |
| 疏穗小野荞(SS) F. leptopodum var. grossii | 0.034 | 0.169 | 0.191 | 0.025 | 0.024 | 0.025 | 0.186 | 0.021 | 0.194 | 0.186 | |
| 野生甜荞(TJ) F. esculentum ssp. ancestralis | 0.193 | 0.145 | 0.209 | 0.199 | 0.198 | 0.199 | 0.039 | 0.194 | 0.003 | 0.038 | |
| 小野荞(XY) F. leptopodum | 0.031 | 0.176 | 0.038 | 0.204 | 0.001 | 0.000 | 0.192 | 0.028 | 0.198 | 0.192 | |
| 密毛野荞(MM) F. densovillosum | 0.003 | 0.160 | 0.036 | 0.196 | 0.033 | 0.001 | 0.191 | 0.027 | 0.197 | 0.191 | |
| 细柄野荞(XB) F. gracilipes | 0.001 | 0.157 | 0.034 | 0.193 | 0.031 | 0.003 | 0.191 | 0.027 | 0.198 | 0.192 | |
| 金荞复合体(JQ) F. cymosum complex | 0.144 | 0.059 | 0.156 | 0.144 | 0.161 | 0.147 | 0.144 | 0.188 | 0.041 | 0.008 | |
| 硬枝万年荞(YZ) F. urophyllum | 0.111 | 0.238 | 0.117 | 0.267 | 0.118 | 0.114 | 0.112 | 0.217 | 0.197 | 0.189 | |
| 栽培甜荞(TQ) F. esculentum | 0.211 | 0.170 | 0.233 | 0.133 | 0.226 | 0.214 | 0.211 | 0.168 | 0.279 | 0.040 | |
| 米苦荞 (MKQ) F. tataricum | 0.156 | 0.002 | 0.168 | 0.146 | 0.175 | 0.159 | 0.156 | 0.060 | 0.237 | 0.071 |
表4 由ITS序列(对角线下)和ndhF-rpl32序列(对角线上)计算的荞麦种间遗传距离
Table 4 Distance between Fagopyrum species based on ITS (below diagonal) and ndhF-rpl32 (above diagonal) sequences
| CC | KQ | SS | TJ | XY | MM | XB | JQ | YZ | TQ | MKQ | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 齿翅野荞(CC) F. gracilipes var. odontopterum | 0.191 | 0.024 | 0.198 | 0.001 | 0.001 | 0.001 | 0.191 | 0.027 | 0.197 | 0.191 | |
| 栽培苦荞(KQ) F. tataricum | 0.157 | 0.185 | 0.037 | 0.191 | 0.190 | 0.191 | 0.007 | 0.188 | 0.039 | 0.001 | |
| 疏穗小野荞(SS) F. leptopodum var. grossii | 0.034 | 0.169 | 0.191 | 0.025 | 0.024 | 0.025 | 0.186 | 0.021 | 0.194 | 0.186 | |
| 野生甜荞(TJ) F. esculentum ssp. ancestralis | 0.193 | 0.145 | 0.209 | 0.199 | 0.198 | 0.199 | 0.039 | 0.194 | 0.003 | 0.038 | |
| 小野荞(XY) F. leptopodum | 0.031 | 0.176 | 0.038 | 0.204 | 0.001 | 0.000 | 0.192 | 0.028 | 0.198 | 0.192 | |
| 密毛野荞(MM) F. densovillosum | 0.003 | 0.160 | 0.036 | 0.196 | 0.033 | 0.001 | 0.191 | 0.027 | 0.197 | 0.191 | |
| 细柄野荞(XB) F. gracilipes | 0.001 | 0.157 | 0.034 | 0.193 | 0.031 | 0.003 | 0.191 | 0.027 | 0.198 | 0.192 | |
| 金荞复合体(JQ) F. cymosum complex | 0.144 | 0.059 | 0.156 | 0.144 | 0.161 | 0.147 | 0.144 | 0.188 | 0.041 | 0.008 | |
| 硬枝万年荞(YZ) F. urophyllum | 0.111 | 0.238 | 0.117 | 0.267 | 0.118 | 0.114 | 0.112 | 0.217 | 0.197 | 0.189 | |
| 栽培甜荞(TQ) F. esculentum | 0.211 | 0.170 | 0.233 | 0.133 | 0.226 | 0.214 | 0.211 | 0.168 | 0.279 | 0.040 | |
| 米苦荞 (MKQ) F. tataricum | 0.156 | 0.002 | 0.168 | 0.146 | 0.175 | 0.159 | 0.156 | 0.060 | 0.237 | 0.071 |
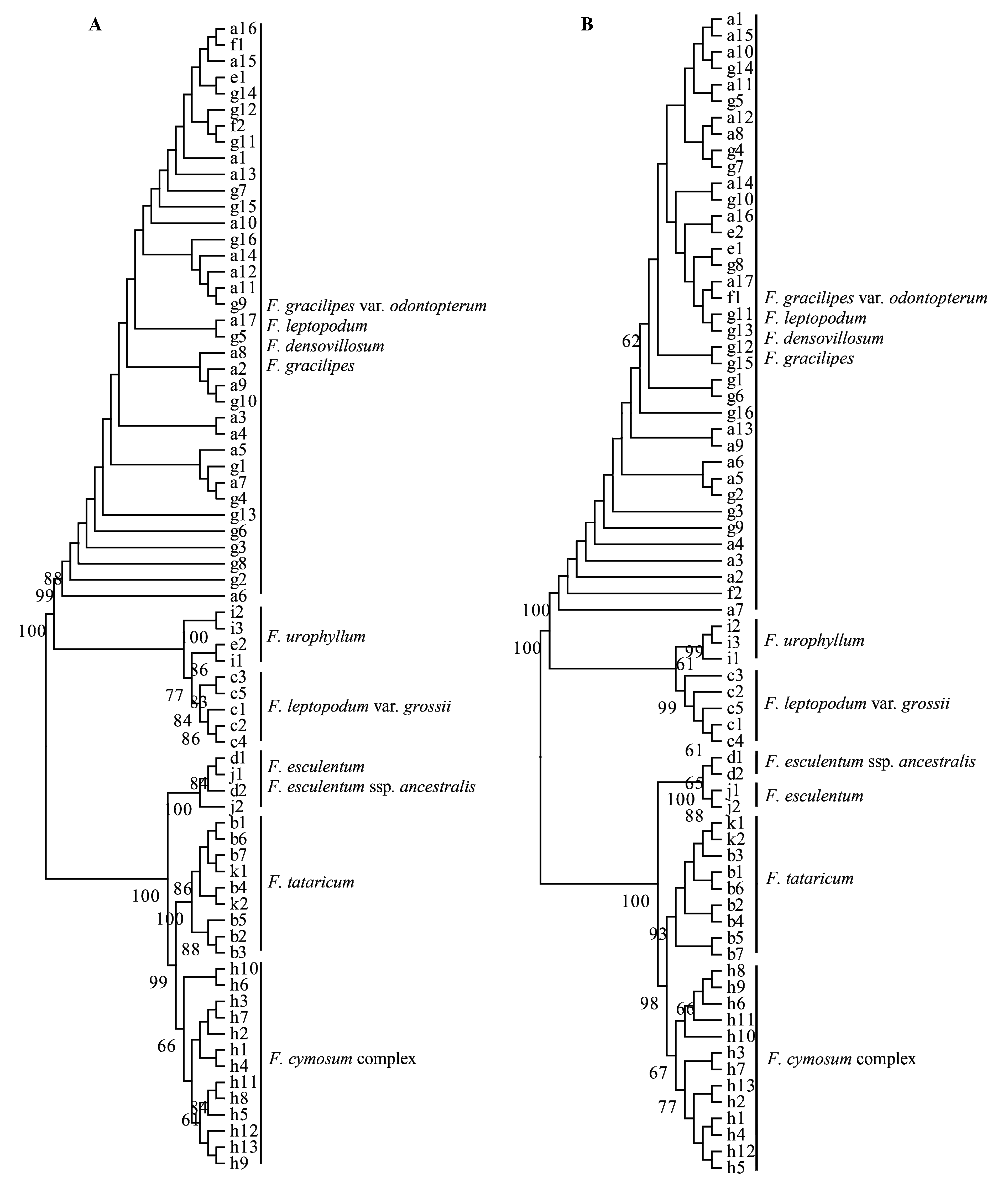
图1 通过邻接法得到的荞麦系统进化树。(A)基于ITS序列; (B)基于ndhF-rpl32序列。
Fig. 1 Neighbor-joining trees of the genus Fagopyrum resulted from (A) ITS sequences; (B) ndhF-rpl32 sequences.
| 1 | Chen QF (1999a) A study of resources of Fagopyrum (Polygonaceae) native to China. Botanical Journal of the Linnean Society, 130, 53-64. |
| 2 | Chen QF (1999b) Hybridization between Fagopyrum (Polygonaceae) species native to China. Botanical Journal of the Linnean Society, 131, 177-185. |
| 3 | Chen QF (2004) A study of isozyme and interspecific hybridization on big-achene group of buckwheat species (Fagopyrum, Polygonaceae). Crop Sciences, 44, 1151-1158. |
| 4 | Chen QF (2012) Plant Sciences on Genus Fagopyrum. Science Press, Beijing.(in Chinese) |
| [陈庆富 (2012) 荞麦属植物科学. 科学出版社, 北京.] | |
| 5 | Doyle JJ (1986) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin, 19, 11-15. |
| 6 | Han JP, Song JY, Liu C, Chen J, Qian J, Zhu YJ, Shi LC, Yao H, Chen SL (2010) Identification of Cistanche species (Orobanchaceae) based on sequences of the plastid psbA- trnH intergenic region. Acta Pharmaceutica Sinica, 45, 126-130. |
| 7 | Hedberg O (1946) Pollen morphology in the genus Polygonees L. (s. lat.) and its taxonomical significance. Svensk Botaisk Tidskrift, 40, 371-404. |
| 8 | Hirose T, Ujihara A, Kitayashi H, Minami M (1995) Pollen tube behavior related to self incompatibility in interspecific crosses of Fagopyrum. Breeding Science, 45, 65-70. |
| 9 | Hou YJ, Zhang ZW, Wu B, Li YQ (2009) Genetic diversity in tartary buckwheat revealed by AFLP analysis. Scientia Agricultura Sinica, 42, 4166-4174.(in Chinese with English abstract) |
| [侯雅君, 张宗文, 吴斌, 李艳琴 (2009) 苦荞种质资源AFLP标记遗传多样性分析. 中国农业科学, 42, 4166-4174.] | |
| 10 | Liu J, He T, Chun Z (2009) Analysis and authentication of chloroplast matK gene sequences of Herba dendrobii. Acta Pharmaceutica Sinica, 44, 1051-1055.(in Chinese with English abstract) |
| [刘静, 何涛, 淳泽 (2009) 药用石斛的叶绿体matK基因序列分析及鉴别. 药学学报, 44, 1051-1055.] | |
| 11 | Ohsako T, Ohishi O (2000) Intra and interspecific phylogeny of wild Fagopyrum (Polygonaceae) species based on nucleotide sequences of noncoding regions in chloroplast DNA. American Journal of Botany, 87, 573-582. |
| 12 | Sharma TR, Jana S (2002a) Random amplified polymorphic DNA (RAPD) variation in Fagopyrum tataricum Gaertn. accessions from China and the Himalayan region. Euphytica, 127, 327-333. |
| 13 | Sharma TR, Jana S (2002b) Species relationships in Fagopyrum revealed by PCR based DNA fingerprinting. Theoretical & Applied Genetics, 105, 306-312. |
| 14 | Shevchuk TE, Gavrilyuk IP, Konarev VG (1981) Immunochemical analysis of the affinities of the genus Fagopyrum and the species of some other genera of the family Polygonaceae. Botanical Journal, 66, 259-267. |
| 15 | Shi JQ, Li YQ, Zhang ZW, Wu B, Wang AH (2015) Genetic diversity of buckwheat and its wild species. Journal of Plant Genetic Resources, 16, 443-450.(in Chinese with English abstract) |
| [史建强, 李艳琴, 张宗文, 吴斌, 王安虎 (2015) 荞麦及其野生种遗传多样性分析. 植物遗传资源学报, 16, 443-450.] | |
| 16 | Steward AN (1930) The Polygonaceae of Eastern Asia. Contributions from the Gray Herbarium of Harvard University, Boston. |
| 17 | Tsuji K, Ohnishi O (2001) Phylogenetic position of east Tibetan natural populations in Tartary buckwheat (Fagopyrum tataricum Gaertn.) revealed by RAPD analyses. Genetic Resources and Crop Evolution, 48, 63-67. |
| 18 | Wang AH, Xia MZ, Cai GZ, Ren YH (2006) Study of the characteristics and geographical distribution of wild buckwheat resources in eastern Liangshan of Sichuan Province. Crops, (5), 25-27.(in Chinese) |
| [王安虎, 夏明忠, 蔡光泽, 任迎虹 (2006) 四川省凉山州东部野生荞麦资源的特征特性和地理分布研究. 作物杂志, (5), 25-27.] | |
| 19 | Wang AH, Xia MZ, Cai GZ, Yang P (2008a) The origin of cultivating buckwheat and the genetic analysis of the kindred species. Southwest China Journal of Agricultural Sciences, 21, 282-285.(in Chinese with English abstract) |
| [王安虎, 夏明忠, 蔡光泽, 杨坪 (2008a) 栽培苦荞麦的起源及其近缘种亲缘分析. 西南农业学报, 21, 282-285.] | |
| 20 | Wang AH, Xia MZ, Cai GZ, Ren YH (2008b) Studies on the characteristics and the geographical distribution diversity of wild buckwheat resources in Sichuan. Southwest China Journal of Agricultural Sciences, 21, 575-580.(in Chinese with English abstract) |
| [王安虎, 夏明忠, 蔡光泽, 任迎虹 (2008b) 四川野生荞麦资源的特征特性与地理分布多样性研究. 西南农业学报, 21, 575-580.] | |
| 21 | Wang LH, Yin FY, Liu JM, Ye CR (2004) Genetic diversity and relationships of the wild buckwheat resources from Yunnan Province by RAPD. Molecular Plant Breeding, 2, 807-815.(in Chinese with English abstract) |
| [王莉花, 殷富有, 刘继梅, 叶昌荣 (2004) 利用RAPD分析云南野生荞麦资源的多样性和亲缘关系. 分子植物育种, 2, 807-815.] | |
| 22 | White TJ, Bruns T, Lee S (1990) Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics. Academic Press, San Diego. |
| 23 | Xu LP (2014) Research progress of nutrient and chemical constituents in buckwheat. Sichuan Chemical Industry, 17(4), 4-8.(in Chinese with English abstract) |
| [徐珑珀 (2014) 荞麦营养与化学成份研究进展. 四川化工, 17(4), 4-8.] | |
| 24 | Yang YX (2008) Studies on Genetic Diversity of Buckwheat Germplasms. PhD dissertation, Sichuan Agricultural University, Chengdu.(in Chinese with English abstract) |
| [杨玉霞 (2008) 荞麦种质资源遗传多样性研究. 博士学位论文, 四川农业大学, 雅安] | |
| 25 | Yang XY, Chen H, Shao JR, Wu Q, Tang Y (2007) Preliminary study on genetic relationship among buckwheat species in north western Sichuan. Acta Botanica Boreali-Occidentalia Sinica, 27, 1752-1758.(in Chinese with English abstract) |
| [杨小艳, 陈惠, 邵继荣, 吴琦, 唐宇 (2007) 川西北荞麦种间亲缘关系初步研究. 西北植物学报, 27, 1752-1758.] | |
| 26 | Yu H, Mao J, Qu SP, Xiong L, Li YY, He R (2005) Study on the relationship of nine varieties of Asian Lilium based on nrDNA ITS. Southwest China Journal of Agricultural Sciences, 18, 387-391.(in Chinese with English abstract) |
| [虞泓, 毛钧, 瞿素萍, 熊丽, 李永谊, 和锐 (2005) 亚洲系百合九个品种的亲缘关系研究——来自nrDNA ITS证据. 西南农业学报, 18, 387-391.] | |
| 27 | Zhang L, Gao FH, Gao LJ, Yin XM, Diao Y (2011) Research progress of buckwheat nutrition, function and its products. South China Agriculture, 5(6), 74-77.(in Chinese) |
| [张玲, 高飞虎, 高伦江, 尹旭敏, 刁源 (2011) 荞麦营养功能及其利用研究进展. 南方农业, 5(6), 74-77.] |
| [1] | 陈静, 张丙昌, 刘燕晋, 武杰, 赵康, 明姣. 荒漠生物结皮细鞘丝藻类(Leptolyngbya-like)蓝藻多样性[J]. 生物多样性, 2024, 32(9): 24186-. |
| [2] | 陈旭, 王国严, 彭培好, 李景吉, 石松林, 张廷斌. 四川攀西地区云南松群落物种多样性和谱系多样性对紫茎泽兰入侵的影响[J]. 生物多样性, 2021, 29(7): 865-874. |
| [3] | 丁陆彬, 马楠, 王国萍, 何思源, 闵庆文. 生物多样性相关传统知识研究热点与前沿的可视化分析[J]. 生物多样性, 2019, 27(7): 716-727. |
| [4] | 吉乃提汗·马木提, 成小军, 谭敦炎. 荒漠短命植物异喙菊的小花异形性及繁殖特性[J]. 生物多样性, 2018, 26(5): 498-509. |
| [5] | 魏亚男, 王晓梅, 姚鹏程, 陈小勇, 李宏庆. 比较不同DNA条形码对中国海岸带耐盐植物的识别率[J]. 生物多样性, 2017, 25(10): 1095-1104. |
| [6] | 钱贞娜, 孟千万, 任明迅. 风筝果镜像花的雌雄异位变化及传粉生态型的形成[J]. 生物多样性, 2016, 24(12): 1364-1372. |
| [7] | 李长昊, 程涛, 董文攀, 周世良. “中国珍稀濒危植物DNA条形码鉴定平台”简介[J]. 生物多样性, 2014, 22(1): 110-110. |
| [8] | 刘文, 梁红, 杨碧忍, 林杰文, 胡延吉. 一种高等植物小片段RNA分子标记方法[J]. 生物多样性, 2012, 20(1): 86-93. |
| [9] | 郑钰, 高博, 孙立夫, 邴艳红, 裴克全. 银叶杜鹃和繁花杜鹃根部真菌的多样性[J]. 生物多样性, 2010, 18(1): 76-82. |
| [10] | 肖龙骞, 朱华. 苏铁nrDNA ITS区的序列多态性:不完全致同进化的证据[J]. 生物多样性, 2009, 17(5): 476-481. |
| [11] | 陈俊秋, 慈秀芹, 李巧明, 李捷. 樟科濒危植物思茅木姜子遗传多样性的ISSR分析[J]. 生物多样性, 2006, 14(5): 410-420. |
| [12] | 吴萍, 李奕仁, 王金玉, 陈宽维. 应用微卫星标记分析中国地方鸡种的遗传变异[J]. 生物多样性, 2003, 11(6): 461-466. |
| [13] | 李太武, 李成华, 宋林生, 苏秀榕. 5个泥蚶群体遗传多样性的RAPD分析[J]. 生物多样性, 2003, 11(2): 118-124. |
| [14] | 章初龙, 徐同. Trichoderma harzianum及其近缘种的分子系统学研究[J]. 生物多样性, 2003, 11(1): 10-19. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
备案号:京ICP备16067583号-7
Copyright © 2026 版权所有 《生物多样性》编辑部
地址: 北京香山南辛村20号, 邮编:100093
电话: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn
![]()