



Biodiv Sci ›› 2013, Vol. 21 ›› Issue (1): 28-37. DOI: 10.3724/SP.J.1003.2013.10097 cstr: 32101.14.SP.J.1003.2013.10097
• Orginal Article • Previous Articles Next Articles
Jianyu He, Xuezhu Liu, Rongtao Zhao, Fangwei Wu, Jianxin Wang*( )
)
Received:2012-04-11
Accepted:2012-08-10
Online:2013-01-20
Published:2013-02-04
Contact:
Wang Jianxin
Jianyu He,Xuezhu Liu,Rongtao Zhao,Fangwei Wu,Jianxin Wang. Diversity of cultured and uncultured bacteria in surface layer sediment from the East China Sea[J]. Biodiv Sci, 2013, 21(1): 28-37.
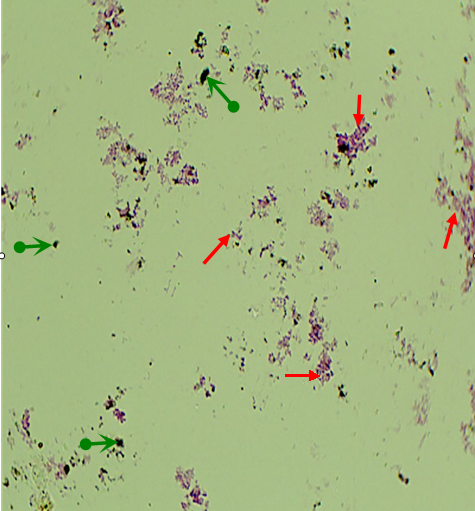
Fig. 1 Direct observation of the sediment from East China Sea surface by the method of buried piece to find the distribution of microorganisms and organics in situ. Red arrows indicate microorganisms, and green arrows indicate organics.
| 培养基 Medium | 稀释倍数 Dilution multiple | 平板菌落数 No. of plate culture count | 菌落单元数量 Numbers of colony- forming units(CFU/g) |
|---|---|---|---|
| M1 | 1,000 | 73 | (7.30±.0.14)×104 |
| Zobell 2216 | 1,000 | 71 | (7.10±0.08)×104 |
| MR2A | 1,000 | 59 | (5.90±0.09)×104 |
| RO | 10,000 | 49 | (4.90±0.17)×105 |
Table 1 Statistics of colony-forming units in four kinds of medias by the dilution method of plate counting
| 培养基 Medium | 稀释倍数 Dilution multiple | 平板菌落数 No. of plate culture count | 菌落单元数量 Numbers of colony- forming units(CFU/g) |
|---|---|---|---|
| M1 | 1,000 | 73 | (7.30±.0.14)×104 |
| Zobell 2216 | 1,000 | 71 | (7.10±0.08)×104 |
| MR2A | 1,000 | 59 | (5.90±0.09)×104 |
| RO | 10,000 | 49 | (4.90±0.17)×105 |
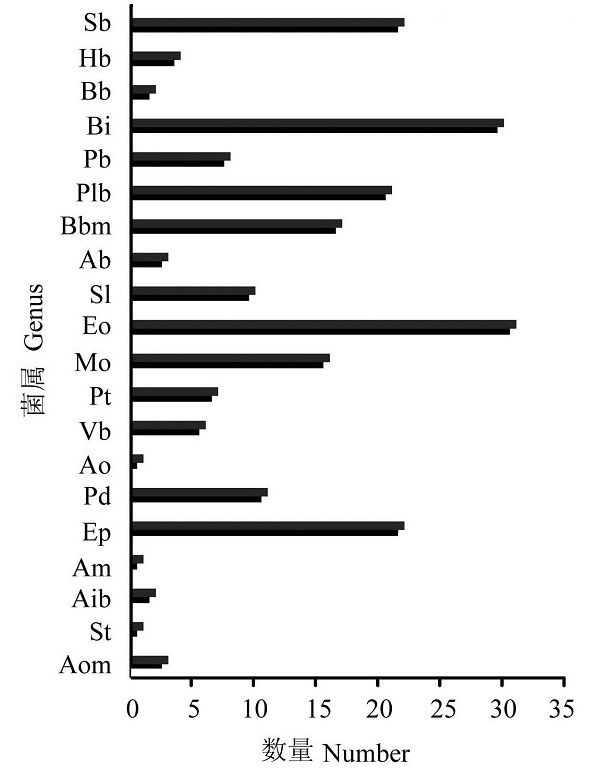
Fig. 2 Distribution of the different genus. Sb, Sporolac- tobacillus; Hb,Halobacillus; Bb, Brevibacillus; Bi, Bacillus; Pb, Paenibacillus; Plb, Pimelobacter; Bbm, Brachybacterium; Ab, Arthrobactor; Sl, Staphylococcus; Eo, Enterococcus; Mo, Marinococcus; Pt, Pasteurella; Vb, Vibrio; Ao, Alteromonas; Pd, Pseudomonas; Ep, Erysipelothrix; Am, Agromyces; Aib, Aneurinibacillus; St, Streptococcus; Aom, Actinomyces.
| 代表菌株 Strain no. | 登录号 Accession no. | 相似菌株(登录号) Closest species (Accession no.) | 相似性 Homology (%) |
|---|---|---|---|
| 芽孢杆菌纲 Bacilli | |||
| DHC01 | JQ904712 | Bacillus pumilus strain PR13 (JQ435673) | 99 |
| DHC02 | JQ904713 | Bacterium YCG4 (JF775413) | 97 |
| DHC03 | JQ904714 | Bacillus tequilensis strain KM35 (JF411314) | 99 |
| DHC04 | JQ904715 | Bacillus sp. M71_S53 (FM992830) | 100 |
| DHC06 | JQ904717 | Bacillus cereus strain CB-10 (JF496530) | 96 |
| DHC07 | JQ904718 | Bacillus subtilis strain B7 (JQ086378) | 100 |
| DHC08 | JQ904719 | Bacillus horikoshii strain NBSL26 (FJ984529) | 99 |
| DHC10 | JQ904720 | Bacillus sp. RHS-str.320 (HE586887) | 97 |
| DHC11 | JQ904721 | Bacillus infantis strain AL10 (JF411305) | 99 |
| DHC12 | JQ904722 | Bacillus aryabhattai isolate PSB59 (HQ242772) | 100 |
| DHC13 | JQ904723 | Bacillus barbaricus strain P2 (GQ200827) | 99 |
| DHC14 | JQ904724 | Bacillus gibsonii strain S-2 (AY737309) | 99 |
| DHC15 | JQ904726 | Paenibacillus sp. GP26-03 (AM162316) | 99 |
| 黄杆菌纲 Flavobacteria | |||
| DHC05 | JQ904716 | Cloacibacterium normanense strain tu33 (FJ544402) | 100 |
| 放线菌纲 Actinobacteria | |||
| DHC09 | JQ904725 | Brevibacterium sp. 210_14 (GQ199716) | 99 |
| γ-变形菌纲 Gamma-proteobacteria | |||
| DHC16 | JQ904727 | Alteromonas sp. SCSWC10 (FJ461444) | 99 |
| DHC17 | JQ904728 | Pseudoalteromonas sp. B296 (FN295786) | 98 |
| DHC18 | JQ904729 | Pseudoalteromonas sp. VS-2 (FJ497709) | 97 |
| DHC19 | JQ904730 | Vibrio algoinfesta strain NBRC 14665 (AB680645) | 99 |
| DHC20 | JQ904731 | Vibrio harveyi EHP8 (FJ227114) | 99 |
| DHC21 | JQ904732 | Vibrio azureus strain CAIM 1457 (JN603238) | 99 |
| DHC22 | JQ904733 | Vibrio parahaemolyticus strain CM12 (EU660326) | 99 |
Table 2 Identification of pure-cultured bacteria isolated from East China Sea surface sediment by bacterial 16S rDNA gene
| 代表菌株 Strain no. | 登录号 Accession no. | 相似菌株(登录号) Closest species (Accession no.) | 相似性 Homology (%) |
|---|---|---|---|
| 芽孢杆菌纲 Bacilli | |||
| DHC01 | JQ904712 | Bacillus pumilus strain PR13 (JQ435673) | 99 |
| DHC02 | JQ904713 | Bacterium YCG4 (JF775413) | 97 |
| DHC03 | JQ904714 | Bacillus tequilensis strain KM35 (JF411314) | 99 |
| DHC04 | JQ904715 | Bacillus sp. M71_S53 (FM992830) | 100 |
| DHC06 | JQ904717 | Bacillus cereus strain CB-10 (JF496530) | 96 |
| DHC07 | JQ904718 | Bacillus subtilis strain B7 (JQ086378) | 100 |
| DHC08 | JQ904719 | Bacillus horikoshii strain NBSL26 (FJ984529) | 99 |
| DHC10 | JQ904720 | Bacillus sp. RHS-str.320 (HE586887) | 97 |
| DHC11 | JQ904721 | Bacillus infantis strain AL10 (JF411305) | 99 |
| DHC12 | JQ904722 | Bacillus aryabhattai isolate PSB59 (HQ242772) | 100 |
| DHC13 | JQ904723 | Bacillus barbaricus strain P2 (GQ200827) | 99 |
| DHC14 | JQ904724 | Bacillus gibsonii strain S-2 (AY737309) | 99 |
| DHC15 | JQ904726 | Paenibacillus sp. GP26-03 (AM162316) | 99 |
| 黄杆菌纲 Flavobacteria | |||
| DHC05 | JQ904716 | Cloacibacterium normanense strain tu33 (FJ544402) | 100 |
| 放线菌纲 Actinobacteria | |||
| DHC09 | JQ904725 | Brevibacterium sp. 210_14 (GQ199716) | 99 |
| γ-变形菌纲 Gamma-proteobacteria | |||
| DHC16 | JQ904727 | Alteromonas sp. SCSWC10 (FJ461444) | 99 |
| DHC17 | JQ904728 | Pseudoalteromonas sp. B296 (FN295786) | 98 |
| DHC18 | JQ904729 | Pseudoalteromonas sp. VS-2 (FJ497709) | 97 |
| DHC19 | JQ904730 | Vibrio algoinfesta strain NBRC 14665 (AB680645) | 99 |
| DHC20 | JQ904731 | Vibrio harveyi EHP8 (FJ227114) | 99 |
| DHC21 | JQ904732 | Vibrio azureus strain CAIM 1457 (JN603238) | 99 |
| DHC22 | JQ904733 | Vibrio parahaemolyticus strain CM12 (EU660326) | 99 |
| 代表克隆子 Clones no. | 登录号 Accession no. | 相似菌株(登录号) Closest species (Accession no.) | 相似菌株分离源 Closest species origin | 相似性 Homology (%) |
|---|---|---|---|---|
| 芽孢杆菌纲 Bacilli | ||||
| DHUC01 | JQ904734 | Bacillus cereus strain E22(GU826149 ) | 巴西土壤 Brazilian soils | 99 |
| DHUC02 | JQ904735 | Bacillus sp. C1(JN571045) | 人造草坪土壤 Chinampa soils | 98 |
| DHUC03 | JQ904736 | Bacillus aquimaris strain T172 (HQ647277) | 东乡野生水稻 Dongxiang wild rice | 99 |
| DHUC04 | JQ904737 | Bacillus subtilis strain MSI-9 (JN993708) | 人参土壤 Ginseng soil | 98 |
| DHUC05 | JQ904738 | Bacillus megaterium strain XAS5-1 (JF496312) | 生物土壤结皮 Biological soil crusts | 100 |
| DHUC06 | JQ904739 | Bacillus subterraneus strain DSM13966T (FR733689) | 普通土壤 Normal soil | 97 |
| DHUC07 | JQ904746 | Bacillus barbaricus strain P2 (GQ200827) | 盐田 Solar salterns | 94 |
| DHUC08 | JQ904740 | Uncultured Bacillus sp. (HQ397100) | 普通土壤 Normal soil | 100 |
| DHUC09 | JQ904741 | Geobacillus stearothermophilus strain DoB50 (JQ359102) | 铁皮石斛 Dendrobium officinale | 99 |
| DHUC10 | JQ904747 | Brevibacillus sp. 100-5 (FJ959373) | 麦田土壤 Wheat field soil | 94 |
| α-变形菌纲 Alpha-proteobacteria | ||||
| DHUC11 | JQ904742 | Uncultured alpha-proteobacterium (FJ205181) | 不活动的热液区沉积物 Inactive hydrothermal field sediments | 95 |
| γ-变形菌纲 Gamma-proteobacteria | ||||
| DHUC12 | JQ904743 | Uncultured Alteromonas sp. (HQ012278) | 海洋 Marine | 100 |
| DHUC13 | JQ904744 | Vibrio harveyi strain EHP8(FJ227114) | 枪乌贼发光器官 Loliginid squid light organs | 100 |
| DHUC14 | JQ904745 | Vibrio hepatarius (NR025491) | 水生动物和海洋环境 Aquatic animals and the marine environment | 98 |
Table 3 Taxonomic affiliation of clones recovered in the bacterial clone libraries from East China Sea surface sediment and analysis of closest species origin
| 代表克隆子 Clones no. | 登录号 Accession no. | 相似菌株(登录号) Closest species (Accession no.) | 相似菌株分离源 Closest species origin | 相似性 Homology (%) |
|---|---|---|---|---|
| 芽孢杆菌纲 Bacilli | ||||
| DHUC01 | JQ904734 | Bacillus cereus strain E22(GU826149 ) | 巴西土壤 Brazilian soils | 99 |
| DHUC02 | JQ904735 | Bacillus sp. C1(JN571045) | 人造草坪土壤 Chinampa soils | 98 |
| DHUC03 | JQ904736 | Bacillus aquimaris strain T172 (HQ647277) | 东乡野生水稻 Dongxiang wild rice | 99 |
| DHUC04 | JQ904737 | Bacillus subtilis strain MSI-9 (JN993708) | 人参土壤 Ginseng soil | 98 |
| DHUC05 | JQ904738 | Bacillus megaterium strain XAS5-1 (JF496312) | 生物土壤结皮 Biological soil crusts | 100 |
| DHUC06 | JQ904739 | Bacillus subterraneus strain DSM13966T (FR733689) | 普通土壤 Normal soil | 97 |
| DHUC07 | JQ904746 | Bacillus barbaricus strain P2 (GQ200827) | 盐田 Solar salterns | 94 |
| DHUC08 | JQ904740 | Uncultured Bacillus sp. (HQ397100) | 普通土壤 Normal soil | 100 |
| DHUC09 | JQ904741 | Geobacillus stearothermophilus strain DoB50 (JQ359102) | 铁皮石斛 Dendrobium officinale | 99 |
| DHUC10 | JQ904747 | Brevibacillus sp. 100-5 (FJ959373) | 麦田土壤 Wheat field soil | 94 |
| α-变形菌纲 Alpha-proteobacteria | ||||
| DHUC11 | JQ904742 | Uncultured alpha-proteobacterium (FJ205181) | 不活动的热液区沉积物 Inactive hydrothermal field sediments | 95 |
| γ-变形菌纲 Gamma-proteobacteria | ||||
| DHUC12 | JQ904743 | Uncultured Alteromonas sp. (HQ012278) | 海洋 Marine | 100 |
| DHUC13 | JQ904744 | Vibrio harveyi strain EHP8(FJ227114) | 枪乌贼发光器官 Loliginid squid light organs | 100 |
| DHUC14 | JQ904745 | Vibrio hepatarius (NR025491) | 水生动物和海洋环境 Aquatic animals and the marine environment | 98 |
| 1 | Bowman JP, McCuaig RD (2003) Biodiversity, community structural shifts, and biogeography of prokaryotes within Antarctic continental shelf sediment. Applied and Environmental Microbiology, 69, 2463-2483. |
| 2 | Chen C (陈琛), Huang S (黄珊), Wu QH (吴群河), Li RY (李锐仪), Zhang RD (张仁铎) (2010) Factors affecting the DAPI fluorescence direct count in the tidal river sediment. Environmental Science(环境科学), 31, 1918-1925. (in Chinese with English abstract) |
| 3 | Dong HL, Zhang GX, Jiang HC, Yu BS, Chapman LR, Lucas CR, Fields MW (2006) Microbial diversity in sediments of saline Qinghai Lake, China: linking geochemical controls to microbial ecology. Microbial Ecology, 51, 65-82. |
| 4 | Dong XZ (东秀珠), Cai MY (蔡妙英) (2001) Manual for Identification of Common Bacterial System (常见细菌系统鉴定手册). Science Press, Beijing. (in Chinese) |
| 5 | Feng BW, Li XR, Wang JH, Hu ZY, Meng H, Xiang LY, Quan ZX (2009) Bacterial diversity of water and sediment in the Changjiang estuary and coastal area of the East China Sea. FEMS Microbiology Ecology, 70, 236-248. |
| 6 | Gontang EA, Fenical W, Jensen PR (2007) Phylogenetic diversity of gram-positive bacteria cultured from marine sediments. Applied and Environmental Microbiology, 73, 3272-3282. |
| 7 | Gray JP, Herwig RP (1996) Phylogenetic analysis of the bacterial communities in marine sediments. Applied and Environmental Microbiology, 62, 4049-4059. |
| 8 | Guo F (郭非), Geng XY (耿绪云), Li X (李翔), Wei JL (魏俊利), Sun JS (孙金生) (2009) Preliminary investigation on the culturable bacteria diversity in Tianjin Coastal Waters. Marine Sciences(海洋科学), 33, 35-40. (in Chinese with English abstract) |
| 9 | Haglund AL, Lantz P, Törnblom E, Tranvik L (2003) Depth distribution of active bacteria and bacterial activity in lake sediment. FEMS Microbiology Ecology, 46, 31-38. |
| 10 | Hill TCJ, Walsh KA, Harris JA, Moffett BF (2003) Using ecological diversity measures with bacterial communities. FEMS Microbiology Ecology, 43, 1-11. |
| 11 | Huo YY (霍颖异), Xu XW (许学伟), Wang CS (王春生), Yang JY (杨俊毅), Wu M (吴敏) (2008) Bacterial diversity of the sediment from Cangnan Large Fishing Bay. Acta Ecologica Sinica(生态学报), 28, 5166-5172. (in Chinese with English abstract) |
| 12 | Inagaki F, Suzuki M, Takai K, Oida H, Sakamoto T, Aoki K, Nealson KH, Horikoshi K (2003) Microbial communities associated with geological horizons in coastal subseafloor sediments from the Sea of Okhotsk. Applied and Environmental Microbiology, 69, 7224-7235. |
| 13 | Jiao NZ (焦念志) (2006) Marine Microbial Ecology (海洋微型生物生态学). Science Press, Beijing. (in Chinese) |
| 14 | Kepner RL Jr, Pratt JR (1994) Use of fluorochromes for direct enumeration of total bacteria in environmental samples: past and present. Microbiological Reviews, 58, 603-615. |
| 15 | Li JB (李家彪) (2008) Donghai Regional Geology (东海区域地质). China Ocean Press, Beijing. (in Chinese) |
| 16 | Li JL, Bai J, Gao HW, Liu GX (2011) Distribution of ammonia-oxidizing Beta-proteobacteria community in surface sediment off the Changjiang River Estuary in summer. Acta Oceanologica Sinica, 30, 92-99. |
| 17 | Li L, Kato C, Horikoshi K (1999) Microbial diversity in sediments collected from the deepest cold-seep area, the Japan Trench. Marine Biotechnology, 1, 391-400. |
| 18 | Li T (李涛), Wang P (王鹏), Wang PX (汪品先) (2008) Microbial diversity in surface sediments of the Xisha Trough, the South China Sea. Acta Ecologica Sinica(生态学报), 28, 1166-1173. (in Chinese with English abstract) |
| 19 | Li ZG (李振高), Luo YM (骆永明), Teng Y (滕应) (2008) Research Methods for Soil and Environmental Microbiology (土壤与环境微生物研究法). Science Press, Beijing. (in Chinese) |
| 20 | Ludwig JA, Reynolds JF (1988) Statistical Ecology. John Wiley & Sons, New York. |
| 21 | Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Briefings in Bioinformatics, 5, 150-163. |
| 22 | Köster M, Wardenga R, Blume M (2008) Microscale investigations of microbial communities in coastal surficial sediments. Marine Ecology, 29, 89-105. |
| 23 | Neto M, Ohannessian A, Delolme C, Bedell JP (2007) Towards an optimized protocol for measuring global dehydrogenase activity in storm-water sediments. Journal of Soils and Sediments, 7, 101-110. |
| 24 | Park SJ, Park BJ, Pham VH, Yoon DN, Kim SK, Rhee SK (2008) Microeukaryotic diversity in marine environments, an analysis of surface layer sediments from the East Sea. The Journal of Microbiology, 46, 244-249. |
| 25 | Polymenakou PN, Fragkioudaki G, Tselepides A (2007) Bacterial and organic matter distribution in the sediments of the Thracian Sea (NE Aegean Sea). Continental Shelf Research, 27, 2187-2197. |
| 26 | Ramalho R, Cunha J, Teixeira P, Gibbs PA (2001) Improved methods for the enumeration of heterotrophic bacteria in bottled mineral waters. Journal of Microbiological Methods, 44, 97-103. |
| 27 | Ravenschlag K, Sahm K, Amann R (2001) Quantitative molecular analysis of the microbial community in marine Arctic sediments (Svalbard). Applied and Environmental Microbiology, 67, 387-395. |
| 28 | Saha P, Chakrabarti T (2006) Emticicia oligotrophica gen. nov., sp. nov., a new member of the family 'Flexibac- teraceae', phylum Bacteroidetes. International Journal of Systematic and Evolutionary Microbiology, 56, 991-995. |
| 29 | Schloss PD, Handelsman J (2005) Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Applied and Environmental Microbiology, 71, 1501-1506. |
| 30 | Song JM (宋金明) (2000) Roles of marine sediments diversity in biogenic element. Marine Sciences(海洋科学), 24, 22-26. (in Chinese with English abstract) |
| 31 | Song ZG (宋志刚), Xu QZ (许强芝), Lu XA (鲁心安), Jiao BH (焦炳华), Ai F (艾峰), Yang Y (杨妤), Liu XY (刘小宇), Shi XQ (施晓琼) (2006) A primary study on population biodiversity of marine microorganisms from East China Sea. Microbiology(微生物学通报), 33, 63-67. (in Chinese with English abstract) |
| 32 | Sun FQ (孙风芹), Wang BJ (汪保江), Li GY (李光玉), Liu XP (刘秀片), Du YP (杜雅萍), Lai QL (赖其良), Shao ZZ (邵宗泽) (2008) Diversity of bacteria isolated from the South China Sea sediments. Acta Microbiologica Sinica(微生物学报), 48, 1578-1587. (in Chinese with English abstract) |
| 33 | Sun HM (孙慧敏), Dai SK (戴世鲲), Wang GH (王广华), Xie LW (谢练武), Li X (李翔) (2010) Phylogenetic diversity analysis of bacteria in the deep-sea sediments from the Bashi Channel by 16S rDNA BLAST. Journal of Tropical Oceanography(热带海洋学报), 29, 41-46. (in Chinese with English abstract) |
| 34 | Urakawa H, Kita-Tsukamota K, Ohwada K (1999) Microbial diversity in marine sediments from Sagami Bay and Tokyo Bay, Japan, as determined by 16S rRNA gene analysis. Microbiology, 145, 3305-3315. |
| 35 | Wang JX (王健鑫), Liu XZ (刘雪珠), Li P (李鹏), Chen XL (陈小龙) (2008) Microbes isolated from the intertidal zone of Zhoushan sea area and their antimicrobial activity. Journal of Shanghai Fisheries University(上海水产大学学报), 17, 507-511. (in Chinese with English abstract) |
| 36 | Wang JX (王健鑫), Xu XE (许贤恩), Zhou LL (周链链), Tao S (陶诗), Liu XZ (刘雪珠), Shi G (石戈) (2012) A preliminary study of microbial diversity of the surface layer sediments from the East China Sea. Oceanologia et Limnologia Sinica(海洋与湖沼), 43, 805-813. (in Chinese with English abstract) |
| 37 | Wang SJ (王书锦), Hu JC (胡江春), Xue DL (薛德林), Ma CX (马成新), Xie QH (谢秋宏), Liu QY (刘全永) (2001) Study of marine microbial resources in Huanghai, Bohai and Liaoning offshore of China. Journal of Jinzhou Normal College (Natural Sciences Edition) (锦州师范学院学报(自然科学版)), 22, 1-5. (in Chinese with English abstract) |
| 38 | Xu F (许飞), Dai X (戴欣), Chen YQ (陈月琴), Zhou H (周惠), Cai JH (蔡剑华), Qu LH (屈良鹄) (2004) Phylogenetic diversity of bacteria and archaea in the Nansha marine sediments, as determined by 16S rDNA analysis. Oceanologia et Limnologia Sinica(海洋与湖沼), 35, 89-94. (in Chinese with English abstract) |
| 39 | Xu MX, Wang P, Wang FP, Xiao X (2005) Microbial diversity at a deep-sea station of the Pacific Nodule Province. Biodiversity and Conservation, 14, 3363-3380. |
| 40 | Zeng YH (曾永辉) (2008) Microbial Diversity and Environ- mental Adaptation Mechanisms in Typical Marine Envir- onments (典型海洋环境中浮游细菌多样性及环境适应机制的研究). PhD dissertation, Xiamen University, Xiamen. (in Chinese with English abstract) |
| 41 | Zhang LB (张林宝), Li TG (李铁刚), Dang HY (党宏月), Sharen GW (萨仁高娃), Zhang W (张伟) (2010) Vertical distribution of Archaea in mud wedge sediments located at inner continental shelf of the East China Sea. Earth Science (Journal of China University of Geosciences) (地球科学(中国地质大学学报)), 35, 254-260. (in Chinese with English abstract) |
| 42 | Zheng ZX (郑兆祥), Yuan Y (袁扬), Chen MM (陈蒙蒙), Du C (杜翠), Liu XZ (刘雪珠) (2011) Isolation of marine bacteria from marine mud of the sea floor in Zhoushan Sea area and screening of marine bacteria against Vibrio spp., the pathogens in aquacultue. Journal of Zhejiang Ocean University (Natural Science) (浙江海洋学院学报(自然科学版)), 30, 51-55, 90. (in Chinese with English abstract) |
| 43 | ZoBell CE (1941) Studies on marine bacteria. I. The cultural requirements of heterotrophic aerobes. Journal of Marine Research, 4, 42-75. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2022 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn ![]()