



Biodiv Sci ›› 2026, Vol. 34 ›› Issue (2): 25373. DOI: 10.17520/biods.2025373 cstr: 32101.14.biods.2025373
• Original Papers: Plant Diversity • Previous Articles Next Articles
Jiajin Cheng1,2( ), Renyi Ma3,*(
), Renyi Ma3,*( )(
)( ), Kangshan Mao1,2,*(
), Kangshan Mao1,2,*( )(
)( )
)
Received:2025-09-18
Accepted:2026-01-26
Online:2026-02-20
Published:2026-03-23
Contact:
Renyi Ma, Kangshan Mao
Supported by:Jiajin Cheng, Renyi Ma, Kangshan Mao. Species delimitation of Juniperus recurva complex[J]. Biodiv Sci, 2026, 34(2): 25373.
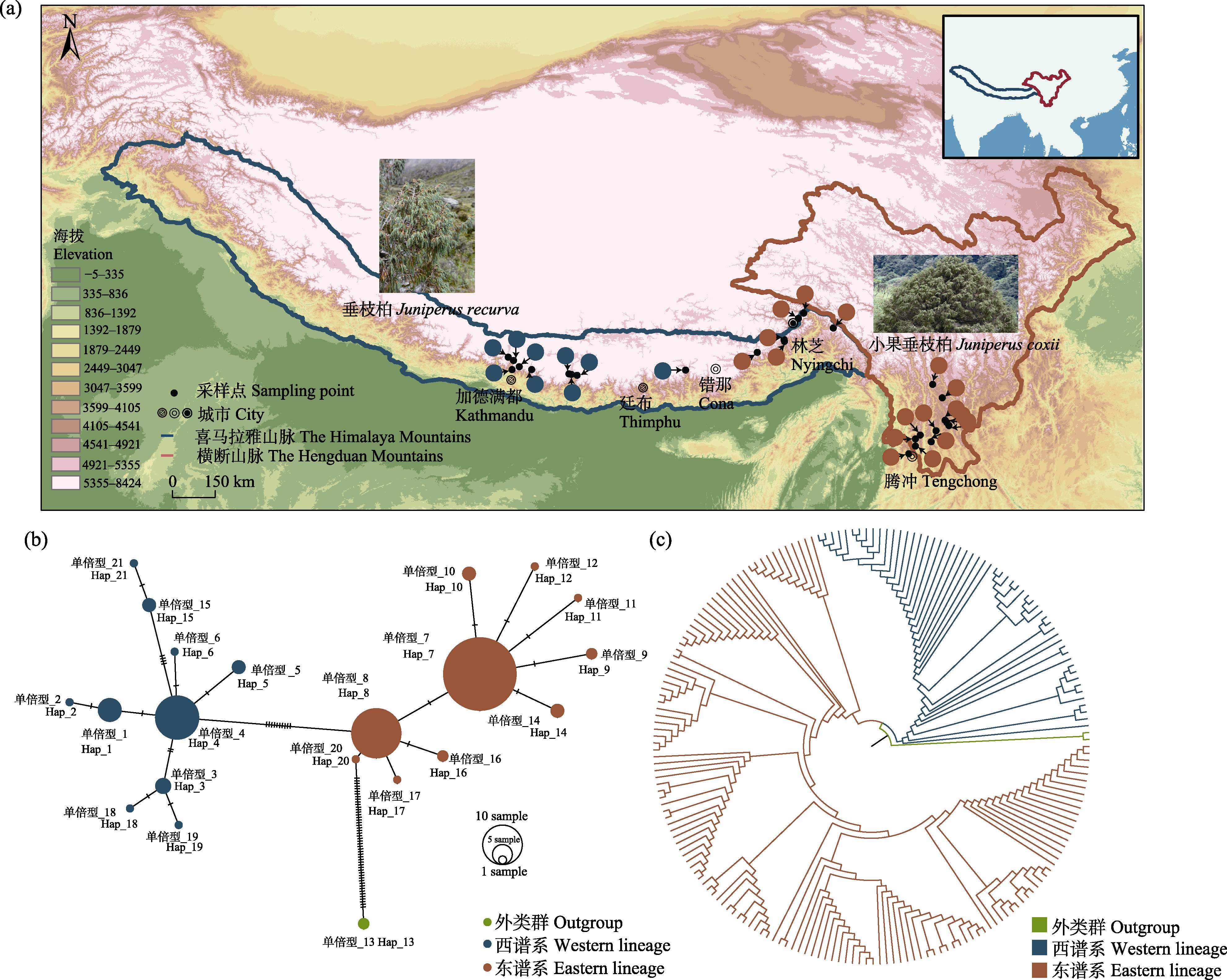
Fig. 1 Sampling distribution map and chloroplast SNP data analysis of the Juniperus recurva complex. (a) Sampling distribution map; (b) Chloroplast SNP haplotype network; (c) Phylogenetic tree of chloroplast SNPs. The geographical boundaries of the Himalaya-Hengduan Mountains region follow Liu et al. (2022).
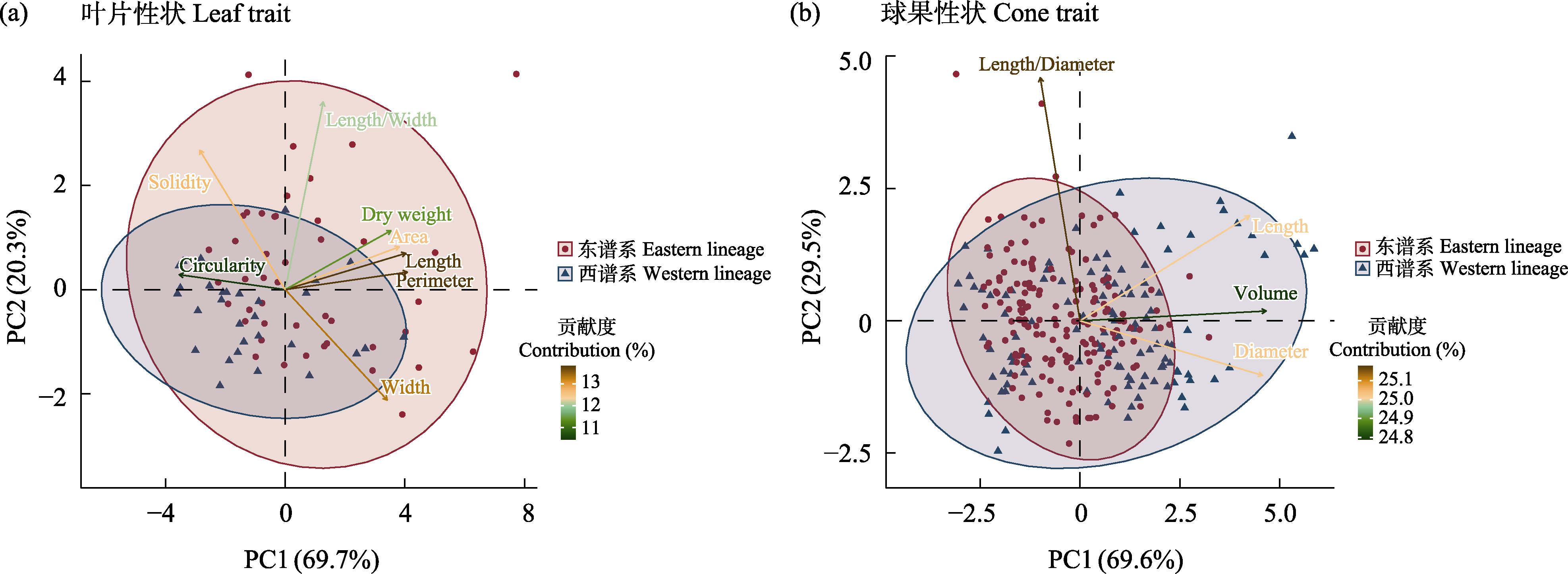
Fig. 2 Morphological trait differentiation of the Juniperus recurva complex. (a) Leaf trait; (b) Cone trait. The variables in the figure correspond to the leaf and cone traits measured in this study.
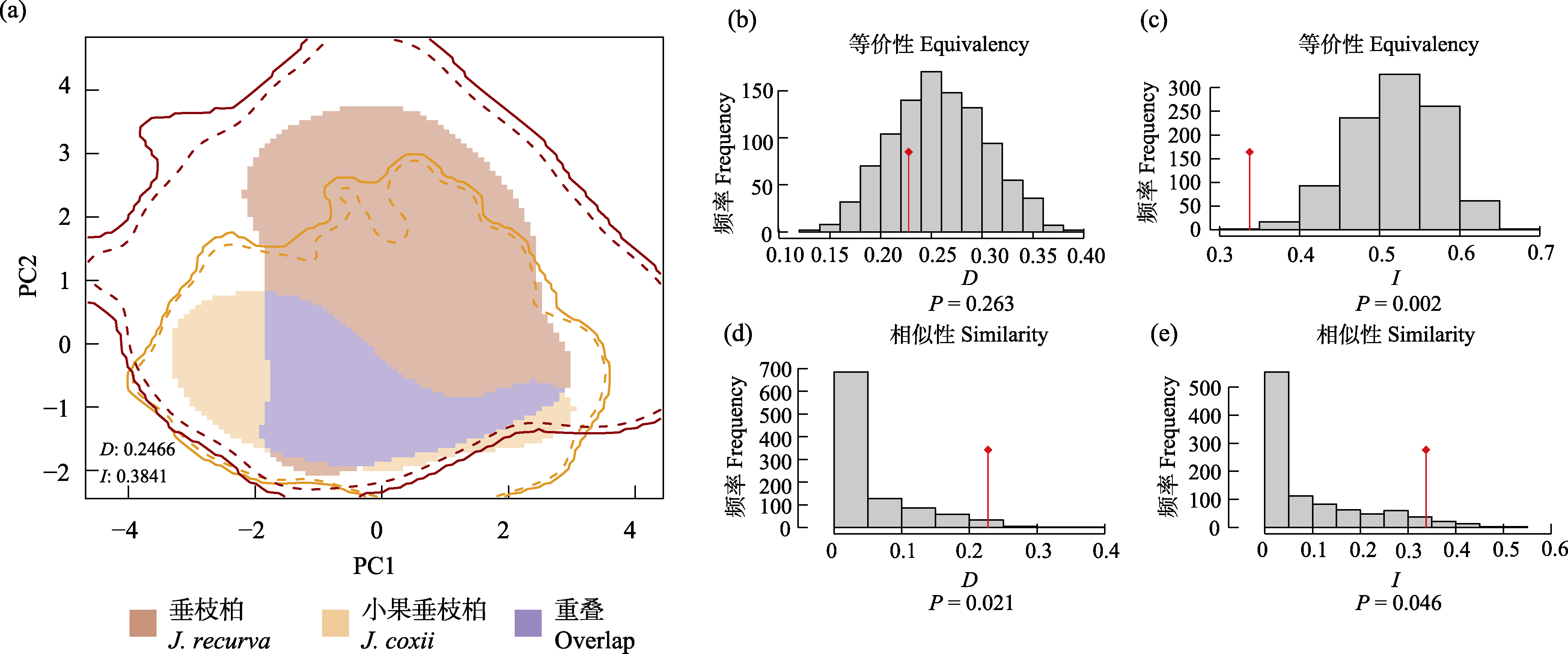
Fig. 4 Niche overlap and tests of equivalency and similarity between Juniperus recurva and J. coxii. (a) Niche overlap analysis. (b-c) Niche equivalency test for D and I. (d-e) Niche similarity test for D and I. D and I represent Schoener’s D and Hellinger-based I indices, respectively, with values ranging from 0 (no overlap) to 1 (complete overlap). The grey bars indicate the null distribution of D and I values, and the red lines indicate the observed values of D and I.
| 变量 Variables | 贡献度 Contribution (PC1) | 贡献度 Contribution (PC2) |
|---|---|---|
| 年平均气温 Annual mean temperature (bio_1) | 10.99% | 15.40% |
| 平均昼夜温差 Mean diurnal range (bio_2) | 8.18% | 13.78% |
| 最湿季降水量 Precipitation of wettest quarter (bio_16) | 0.98% | 26.29% |
| 最冷季降水量 Precipitation of coldest quarter (bio_19) | 1.09% | 26.19% |
| 气候湿度指标 Climatic moisture index (climaticmoistureindex) | 22.83% | 2.08% |
| 气温大陆度指数 Continentality (continentality) | 24.11% | 0.25% |
| 潜在蒸散量的月度变化 Potential Evapotranspiration seasonality (PETseasonality) | 24.00% | 1.96% |
| SAGA-GIS地形湿润指数 SAGA-GIS based topographic wetness index (topo Wet) | 1.91% | 2.35% |
| 年平均太阳辐射 Annual mean solar radiation (sard) | 5.92% | 11.71% |
Table 1 Variance contribution of the nine environmental variables to PC1 and PC2 in ecological niche differentiation analysis
| 变量 Variables | 贡献度 Contribution (PC1) | 贡献度 Contribution (PC2) |
|---|---|---|
| 年平均气温 Annual mean temperature (bio_1) | 10.99% | 15.40% |
| 平均昼夜温差 Mean diurnal range (bio_2) | 8.18% | 13.78% |
| 最湿季降水量 Precipitation of wettest quarter (bio_16) | 0.98% | 26.29% |
| 最冷季降水量 Precipitation of coldest quarter (bio_19) | 1.09% | 26.19% |
| 气候湿度指标 Climatic moisture index (climaticmoistureindex) | 22.83% | 2.08% |
| 气温大陆度指数 Continentality (continentality) | 24.11% | 0.25% |
| 潜在蒸散量的月度变化 Potential Evapotranspiration seasonality (PETseasonality) | 24.00% | 1.96% |
| SAGA-GIS地形湿润指数 SAGA-GIS based topographic wetness index (topo Wet) | 1.91% | 2.35% |
| 年平均太阳辐射 Annual mean solar radiation (sard) | 5.92% | 11.71% |
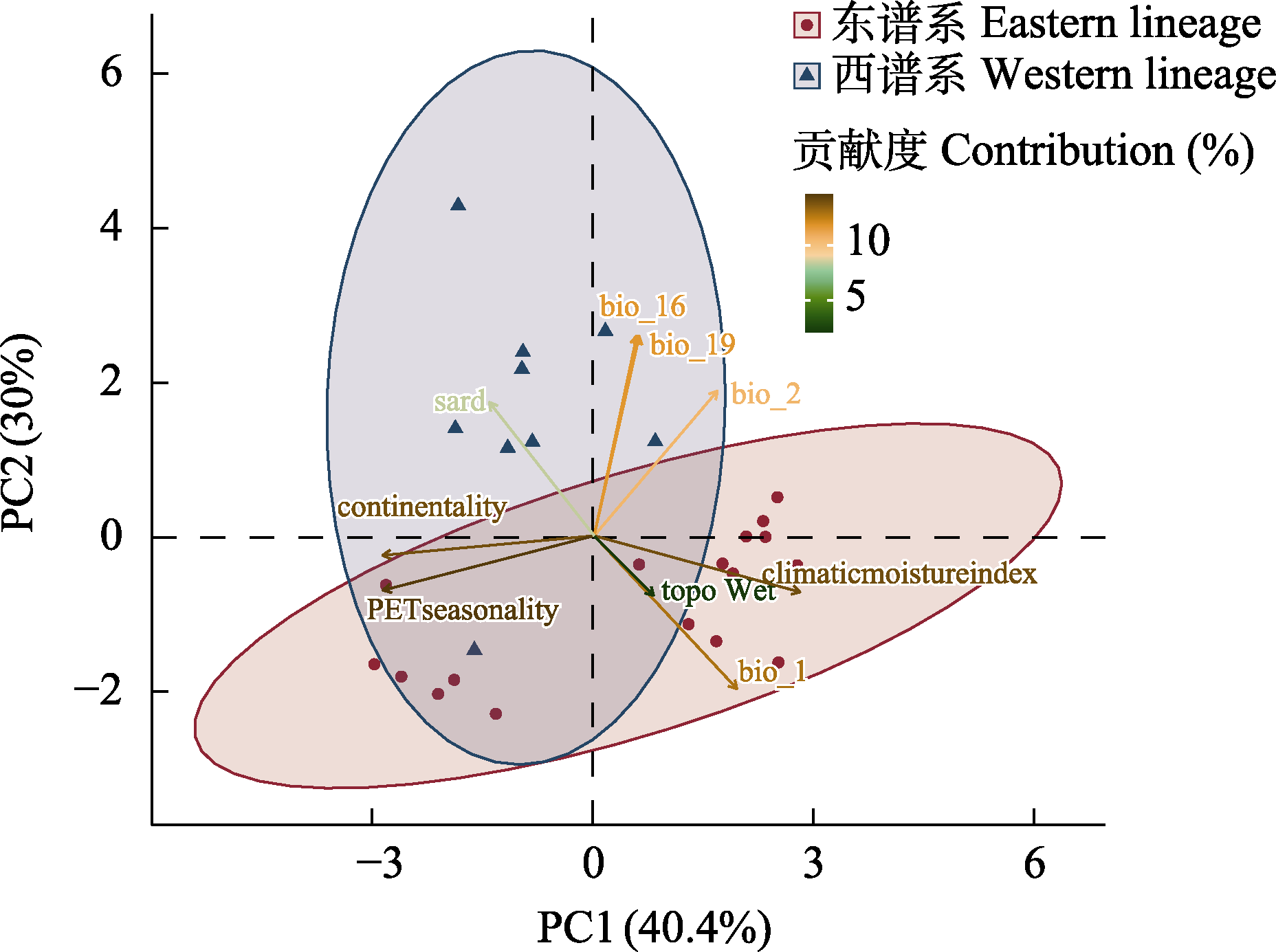
Fig. 5 Niche differentiation analysis in the Juniperus recurva complex. Red circles represent eastern lineages; blue triangles indicate western lineages. The environmental variables shown are as defined in Table 1.
| [1] |
Adams KL, Palmer JD (2003) Evolution of mitochondrial gene content: Gene loss and transfer to the nucleus. Molecular Phylogenetics and Evolution, 29, 380-395.
DOI PMID |
| [2] |
Adams RP (2000) Systematics of the one seeded Juniperus of the eastern hemisphere based on leaf essential oils and random amplified polymorphic DNAs (RAPDs). Biochemical Systematics and Ecology, 28, 529-543.
DOI URL |
| [3] |
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics, 30, 2114-2120.
DOI PMID |
| [4] |
Brown JL, Bennett JR, French CM (2017) SDMtoolbox 2.0: The next generation Python-based GIS toolkit for landscape genetic, biogeographic and species distribution model analyses. PeerJ, 5, e4095.
DOI URL |
| [5] |
Brown JL, Hill DJ, Dolan AM, Carnaval AC, Haywood AM (2018) PaleoClim, high spatial resolution paleoclimate surfaces for global land areas. Scientific Data, 5, 180254.
DOI URL |
| [6] |
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: A tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics, 25, 1972-1973.
DOI PMID |
| [7] | Claridge MF, Dawah AH, Wilson MR (1997) Species: The Units of Biodiversity. Springer, Berlin. |
| [8] |
Clark MJ, Chen R, Lam HYK, Karczewski KJ, Chen R, Euskirchen G, Butte AJ, Snyder M (2011) Performance comparison of exome DNA sequencing technologies. Nature Biotechnology, 29, 908-914.
DOI PMID |
| [9] | Coyne JA, Coyne HA, Orr HA (2004) Speciation. Oxford University Press, Oxford. |
| [10] | Cracraft J (1983) Species concepts and speciation analysis. In: CurrentOrnithology (ed. Johnston RF), pp.159-187. Springer, New York. |
| [11] |
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST, McVean G, Durbin R (2011) The variant call format and VCFtools. Bioinformatics, 27, 2156-2158.
DOI PMID |
| [12] |
Darling AE, Mau B, Perna NT (2010) progressiveMauve: Multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE, 5, e11147.
DOI URL |
| [13] | Darwin C (1859) On the Origin of Species by Means of Natural Selection, or, The Preservation of Favoured Races in the Struggle for Life. John Murray, London. |
| [14] |
Dayrat B (2005) Towards integrative taxonomy. Biological Journal of the Linnean Society, 85, 407-415.
DOI URL |
| [15] |
DePristo MA, Banks E, Poplin R, Garimella KV, Maguire JR, Hartl C, Philippakis AA, del Angel G, Rivas MA, Hanna M, McKenna A, Fennell TJ, Kernytsky AM, Sivachenko AY, Cibulskis K, Gabriel SB, Altshuler D, Daly MJ (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genetics, 43, 491-498.
DOI PMID |
| [16] |
Di Cola V, Broennimann O, Petitpierre B, Breiner FT, D’Amen M, Randin C, Engler R, Pottier J, Pio D, Dubuis A, Pellissier L, Mateo RG, Hordijk W, Salamin N, Guisan A (2017) ecospat: An R package to support spatial analyses and modeling of species niches and distributions. Ecography, 40, 774-787.
DOI URL |
| [17] |
Duan RY, Kong XQ, Huang MY, Fan WY, Wang ZG (2014) The predictive performance and stability of six species distribution models. PLoS ONE, 9, e112764.
DOI URL |
| [18] |
Duminil J, Di Michele M (2009) Plant species delimitation: A comparison of morphological and molecular markers. Plant Biosystems, 143, 528-542.
DOI URL |
| [19] |
Elith J, Leathwick JR (2009) Species distribution models: Ecological explanation and prediction across space and time. Annual Review of Ecology, Evolution, and Systematics, 40, 677-697.
DOI URL |
| [20] |
Excoffier L, Lischer HEL (2010) Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources, 10, 564-567.
DOI PMID |
| [21] | Farjon A (2000) A Monograph of Cupressaceae and Sciadopitys. Royal Botanic Gardens, Kew, London. |
| [22] |
Fennessy J, Bidon T, Reuss F, Kumar V, Elkan P, Nilsson MA, Vamberger M, Fritz U, Janke A (2016) Multi-locus analyses reveal four giraffe species instead of one. Current Biology, 26, 2543-2549.
DOI PMID |
| [23] |
Fick SE, Hijmans RJ (2017) WorldClim 2: New 1-km spatial resolution climate surfaces for global land areas. International Journal of Climatology, 37, 4302-4315.
DOI URL |
| [24] | Franco-Estrada D, Ortiz E, Villaseñor JL, Arias S (2022) Species distribution modelling and predictor variables for species distribution and niche preferences Pilosocereus leucocephalus group s. s. (Cactaceae). Systematics and Biodiversity, 20, 1-17. |
| [25] |
Frankham R (2003) Genetics and conservation biology. Comptes Rendus Biologies, 326, 22-29.
DOI URL |
| [26] |
Fu CN, Wu CS, Ye LJ, Mo ZQ, Liu J, Chang YW, Li DZ, Chaw SM, Gao LM (2019) Prevalence of isomeric plastomes and effectiveness of plastome super-barcodes in yews (Taxus) worldwide. Scientific Reports, 9, 2773.
DOI |
| [27] | Fu LG, Yu YF, Farjon A (1999) “Juniperus Linnaeus, Sp. Pl. 2: 1038. 1753.” Flora of China, 4, 69-77. Science Press, Beijing. |
| [28] |
Hong DY (2016a) Biodiversity pursuits need a scientific and operative species concept. Biodiversity Science, 24, 979-999. (in Chinese with English abstract)
DOI URL |
| [洪德元 (2016a) 生物多样性事业需要科学、可操作的物种概念. 生物多样性, 24, 979-999.] | |
| [29] |
Hong DY (2016b) Opinion of raising rationality in species delimitation. Biodiversity Science, 24, 360-361. (in Chinese with English abstract)
DOI URL |
| [洪德元 (2016b) 关于提高物种划分合理性的意见. 生物多样性, 24, 360-361.] | |
| [30] |
Hong DY (2020) Gen-morph species concept—A new and integrative species concept for outbreeding organisms. Journal of Systematics and Evolution, 58, 725-742.
DOI URL |
| [31] |
Jiang YZ, Yang JB, Folk RA, Zhao JL, Liu J, He ZS, Peng H, Yang SX, Xiang CL, Yu XQ (2024) Species delimitation of tea plants (Camellia sect. Thea) based on super-barcodes. BMC Plant Biology, 24, 181.
DOI |
| [32] |
Jin JJ, Yu WB, Yang JB, Song Y, de Pamphilis CW, Yi TS, Li DZ (2020) GetOrganelle: A fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biology, 21, 241.
DOI |
| [33] |
Kass JM, Muscarella R, Galante PJ, Bohl CL, Pinilla-Buitrago GE, Boria RA, Soley-Guardia M, Anderson RP (2021) ENMeval 2.0: Redesigned for customizable and reproducible modeling of species’ niches and distributions. Methods in Ecology and Evolution, 12, 1602-1608.
DOI URL |
| [34] |
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Molecular Biology and Evolution, 30, 772-780.
DOI PMID |
| [35] | Lê S, Josse J, Husson F (2008) FactoMineR: An R package for multivariate analysis. Journal of Statistical Software, 25, 1-18. |
| [36] |
Leigh JW, Bryant D (2015) Popart: Full-feature software for haplotype network construction. Methods in Ecology and Evolution, 6, 1110-1116.
DOI URL |
| [37] | Li GY, Lin L, Yi SH (2024) Predicting the potential suitable growth area of Juniperus recurva in Xizang based on MaxEnt model. Journal of Plateau Agriculture, 8, 302-309, 341. (in Chinese with English abstract) |
| [李国营, 林玲, 易胜寒 (2024) 基于MaxEnt模型预测垂枝柏在西藏的潜在适生区. 高原农业, 8, 302-309, 341.] | |
| [38] |
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics, 25, 1754-1760.
DOI PMID |
| [39] |
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics, 25, 2078-2079.
DOI PMID |
| [40] |
Li JL, Zhang YJ, Ruhsam M, Milne RI, Wang Y, Wu DY, Jia SY, Tao TZ, Mao KS (2022) Seeing through the hedge: Phylogenomics of Thuja (Cupressaceae) reveals prominent incomplete lineage sorting and ancient introgression for Tertiary relict flora. Cladistics, 38, 187-203.
DOI URL |
| [41] |
Li YC, Wen J, Ren Y, Zhang JQ (2019) From seven to three: Integrative species delimitation supports major reduction in species number in Rhodiola section Trifida (Crassulaceae) on the Qinghai-Tibetan Plateau. Taxon, 68, 268-279.
DOI URL |
| [42] |
Liu CR, White M, Newell G (2013) Selecting thresholds for the prediction of species occurrence with presence-only data. Journal of Biogeography, 40, 778-789.
DOI URL |
| [43] | Liu HT, Wang WJ, Song G, Qu YH, Li SH, Fjeldså J, Lei FM (2012) Interpreting the process behind endemism in China by integrating the phylogeography and ecological niche models of the Stachyridopsis ruficeps. PLoS ONE, 7, e46761. |
| [44] |
Liu J, Milne RI, Zhu GF, Spicer RA, Wambulwa MC, Wu ZY, Boufford DE, Luo YH, Provan J, Yi TS, Cai J, Wang H, Gao LM, Li DZ (2022) Name and scale matter: Clarifying the geography of Tibetan Plateau and adjacent mountain regions. Global and Planetary Change, 215, 103893.
DOI URL |
| [45] |
Liu J, Möller M, Provan J, Gao LM, Poudel RC, Li DZ (2013) Geological and ecological factors drive cryptic speciation of yews in a biodiversity hotspot. New Phytologist, 199, 1093-1108.
DOI PMID |
| [46] |
Liu JQ (2016) “The integrative species concept” and “species on the speciation way”. Biodiversity Science, 24, 1004-1008. (in Chinese with English abstract)
DOI URL |
|
[刘建全 (2016) “整合物种概念”和“分化路上的物种”. 生物多样性, 24, 1004-1008.]
DOI |
|
| [47] | Luo D, Xu B, Li ZM, Sun H (2017) The ‘Ward Line-Mekong-Salween Divide’ is an important floristic boundary between the eastern Himalaya and Hengduan Mountains: Evidence from the phylogeographical structure of subnival herbs Marmoritis complanatum (Lamiaceae). Botanical Journal of the Linnean Society, 185, 482-496. |
| [48] |
Lyu LK, Wang DL, Li L, Zhu YY, Jiang DC, Liu JQ, Xu XT (2021) Polyphyly and species delimitation of Picea brachytyla (Pinaceae) based on population genetic data. Journal of Systematics and Evolution, 59, 515-523.
DOI URL |
| [49] | Mayr E (1949) Systematics and the Origin of Species. Columbia University Press, New York. |
| [50] |
Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, Lanfear R (2020) IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Molecular Biology and Evolution, 37, 1530-1534.
DOI PMID |
| [51] |
Montès N, Tosini L, Laffont-Schwob I, Le Bagousse-Pinguet Y, Folzer H (2024) FAMeLeS: A multispecies and fully automated method to measure morphological leaf traits. Methods in Ecology and Evolution, 15, 484-492.
DOI URL |
| [52] |
Mu QY, Yu CC, Wang Y, Han TS, Wang H, Ding WN, Zhang QY, Low SL, Zheng QJ, Peng C, Hu ZY, Xing YW (2022) Comparative phylogeography of Acanthocalyx (Caprifoliaceae) reveals distinct genetic structures in the Himalaya-Hengduan Mountains. Alpine Botany, 132, 153-168.
DOI |
| [53] |
Ott T, Schall M, Vogt R, Oberprieler C (2022) The warps and wefts of a polyploidy complex: Integrative species delimitation of the diploid Leucanthemum (Compositae, Anthemideae) representatives. Plants, 11, 1878.
DOI URL |
| [54] |
Padial JM, De La Riva I (2010) A response to recent proposals for integrative taxonomy. Biological Journal of the Linnean Society, 101, 747-756.
DOI URL |
| [55] |
Peng YX, Li Y, Cao GL, Li HL, Shin Y, Piao ZJ, Perez F, Zhu WH, Borzée A (2023) Estimation of habitat suitability and landscape connectivity for Liaoning and Jilin clawed salamanders (Hynobiidae: Onychodactylus) in the transboundary region between the People’s Republic of China and the Democratic People’s Republic of Korea. Global Ecology and Conservation, 48, e02694.
DOI URL |
| [56] |
Phillips SJ, Anderson RP, Schapire RE (2006) Maximum entropy modeling of species geographic distributions. Ecological Modelling, 190, 231-259.
DOI URL |
| [57] |
Prata EMB, Sass C, Rodrigues DP, Domingos FMCB, Specht CD, Damasco G, Ribas CC, Fine PVA, Vicentini A (2018) Towards integrative taxonomy in Neotropical botany: Disentangling the Pagamea guianensis species complex (Rubiaceae). Botanical Journal of the Linnean Society, 188, 213-231.
DOI URL |
| [58] |
Rissler LJ, Apodaca JJ (2007) Adding more ecology into species delimitation: Ecological niche models and phylogeography help define cryptic species in the black salamander (Aneides flavipunctatus). Systematic Biology, 56, 924-942.
DOI PMID |
| [59] | Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6:DNA sequence polymorphism analysis of large data sets. Molecular Biology and Evolution, 34, 3299-3302. |
| [60] |
Ruiz-Sanchez E, Sosa V (2010) Delimiting species boundaries within the Neotropical bamboo Otatea (Poaceae: Bambusoideae) using molecular, morphological and ecological data. Molecular Phylogenetics and Evolution, 54, 344-356.
DOI URL |
| [61] |
Smith MA, Poyarkov NA JR, Hebert PDN (2008) CO 1 DNA barcoding amphibians: Take the chance, meet the challenge. Molecular Ecology Resources, 8, 235-246.
DOI PMID |
| [62] |
Sobel JM, Chen GF, Watt LR, Schemske DW (2010) The biology of speciation. Evolution, 64, 295-315.
DOI PMID |
| [63] |
Song BH, Wang XQ, Wang XR, Ding KY, Hong DY (2003) Cytoplasmic composition in Pinus densata and population establishment of the diploid hybrid pine. Molecular Ecology, 12, 2995-3001.
DOI URL |
| [64] | Tang JM, Yin XJ, Gao WJ, Liu YF, Li ZK (2024) Distribution of potential suitable areas of rare and endangered Cupressaceae species in China under climate change. Journal of Central South University of Forestry & Technology, 44(8), 49-61. (in Chinese with English abstract) |
| [唐继敏, 殷晓洁, 高伟杰, 刘一飞, 李子康 (2024) 气候变化下中国珍稀濒危柏科树种潜在适生区分布. 中南林业科技大学学报, 44(8), 49-61.] | |
| [65] |
Tarasov A, Vilella AJ, Cuppen E, Nijman IJ, Prins P (2015) Sambamba: Fast processing of NGS alignment formats. Bioinformatics, 31, 2032-2034.
DOI PMID |
| [66] |
Title PO, Bemmels JB (2018) ENVIREM: An expanded set of bioclimatic and topographic variables increases flexibility and improves performance of ecological niche modeling. Ecography, 41, 291-307.
DOI URL |
| [67] |
Vignali S, Barras AG, Arlettaz R, Braunisch V (2020) SDMtune: An R package to tune and evaluate species distribution models. Ecology and Evolution, 10, 11488-11506.
DOI PMID |
| [68] |
Wagner DB (1992) Nuclear, chloroplast, and mitochondrial DNA polymorphisms as biochemical markers in population genetic analyses of forest trees. New Forests, 6, 373-390.
DOI URL |
| [69] | Wang CQ, Qin H, Zhao C, Yang L, Yu TT, Zhang YX, Luo X, Qin QB, Liu SJ (2021) Whole-genome re-sequencing and transcriptome reveal oogenesis-related genes in autotetraploid Carassius auratus. Marine Biotechnology, 23, 233-241. |
| [70] |
Wang IJ, Bradburd GS (2014) Isolation by environment. Molecular Ecology, 23, 5649-5662.
DOI PMID |
| [71] |
Warren DL, Glor RE, Turelli M (2008) Environmental niche equivalency versus conservatism: Quantitative approaches to niche evolution. Evolution, 62, 2868-2883.
DOI PMID |
| [72] |
Warren DL, Glor RE, Turelli M (2010) ENMTools: A toolbox for comparative studies of environmental niche models. Ecography, 33, 607-611.
DOI URL |
| [73] | Wilkins J (2009) Species:A History of the Idea. University of California Press, Berkeley. |
| [74] | Yang GS (2017) Studies on the Conservation Biology and Chloroplast Genomes of the Endangered Resource Plant Yunnanopilia longistaminea and Related Species. PhD dissertation, Yunnan University, Kunming. (in Chinese with English abstract) |
| [杨冠松 (2017) 濒危资源植物甜菜树保护生物学及其近缘种叶绿体基因组研究.博士学位论文, 云南大学, 昆明.] | |
| [75] | Yang L, Wei FW, Zhan XJ, Fan HZ, Zhao PP, Huang GP, Chang J, Lei YH, Hu YB (2022) Evolutionary conservation genomics reveals recent speciation and local adaptation in threatened takins. Molecular Biology and Evolution, 39, msac111. |
| [76] | Zhou J, Jiao ZY, Guo JH, Wang BS, Zheng JW (2021) Complete chloroplast genome sequencing of five Salix species and its application in the phylogeny and taxonomy of the genus. Mitochondrial DNA Part B: Resources, 6, 2348-2352. |
| [77] |
Zhou YF, Abbott RJ, Jiang ZY, Du FK, Milne RI, Liu JQ (2010) Gene flow and species delimitation: A case study of two pine species with overlapping distributions in Southeast China. Evolution, 64, 2342-2352.
DOI PMID |
| [1] | Hong Deyuan. A brief discussion on methodology in taxonomy [J]. Biodiv Sci, 2025, 33(2): 24541-. |
| [2] | Zhizhong Li, Shuai Peng, Qingfeng Wang, Wei Li, Shichu Liang, Jinming Chen. Cryptic diversity of the genus Ottelia in China [J]. Biodiv Sci, 2023, 31(2): 22394-. |
| [3] | Yanyan Liu, Chang Liu, Xiaoxin Wei. Current status of taxonomy, systematics and conservation of the white pines in China and adjacent regions [J]. Biodiv Sci, 2022, 30(2): 21344-. |
| [4] | De-Yuan Hong, Shiliang Zhou, Xingjin He, Junhui Yuan, Yanlong Zhang, Fangyun Cheng, Xiuli Zeng, Yan Wang, Xiuxin Zhang. Current status of wild tree peony species with special reference to conservation [J]. Biodiv Sci, 2017, 25(7): 781-793. |
| [5] | Qiner Yang. Comments on species-level taxonomy of plants in China [J]. Biodiv Sci, 2016, 24(9): 1024-1030. |
| [6] | Jianquan Liu. “The integrative species concept” and “species on the speciation way” [J]. Biodiv Sci, 2016, 24(9): 1004-1008. |
| [7] | De-Yuan Hong. Biodiversity pursuits need a scientific and operative species concept [J]. Biodiv Sci, 2016, 24(9): 979-999. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn