



Biodiv Sci ›› 2025, Vol. 33 ›› Issue (12): 25283. DOI: 10.17520/biods.2025283 cstr: 32101.14.biods.2025283
• Technology and Methodology • Previous Articles Next Articles
Jianing An1,2,3, Changchun Zhang1,2,3( ), Jiantao Wang5, Zhiyong Pei6, Dandan Bai7, Junguo Zhang1,2,4,*(
), Jiantao Wang5, Zhiyong Pei6, Dandan Bai7, Junguo Zhang1,2,4,*( )
)
Received:2025-07-20
Accepted:2025-08-28
Online:2025-12-20
Published:2026-01-09
Supported by:Jianing An, Changchun Zhang, Jiantao Wang, Zhiyong Pei, Dandan Bai, Junguo Zhang. An open-set domain adaptation method for wildlife image recognition via adversarial disentanglement and feature alignment[J]. Biodiv Sci, 2025, 33(12): 25283.
| 数据集 Datasets | 野猪 Wild boar | 中华斑羚 Chinese goral | 马鹿 Wapiti | 猞猁 Eurasian lynx | 貉 Raccoon dog | 狗獾 Eurasian badger | 东北兔 Manchurian hare | 狍 Eastern roe deer |
|---|---|---|---|---|---|---|---|---|
| D | 911 | 305 | 315 | 79 | 193 | 214 | 25 | 168 |
| N | 1,449 | 758 | 229 | 136 | 560 | 429 | 401 | 25 |
Table 1 Statistical information of the wildlife image dataset from the Ulaanba National Nature Reserve, Inner Mongolia
| 数据集 Datasets | 野猪 Wild boar | 中华斑羚 Chinese goral | 马鹿 Wapiti | 猞猁 Eurasian lynx | 貉 Raccoon dog | 狗獾 Eurasian badger | 东北兔 Manchurian hare | 狍 Eastern roe deer |
|---|---|---|---|---|---|---|---|---|
| D | 911 | 305 | 315 | 79 | 193 | 214 | 25 | 168 |
| N | 1,449 | 758 | 229 | 136 | 560 | 429 | 401 | 25 |
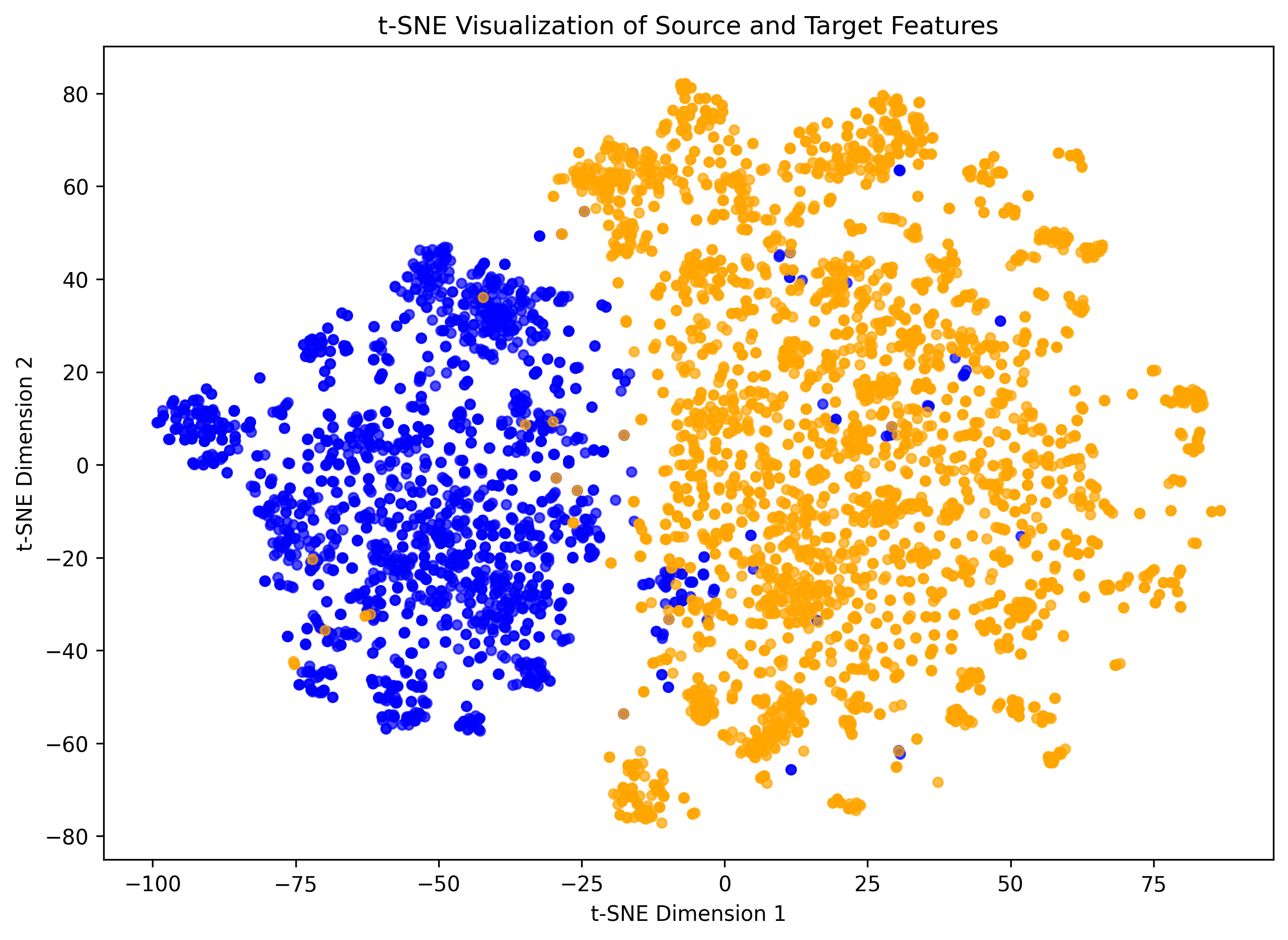
Fig. 1 t-SNE visualization of feature distributions for sub-datasets D and N in the DN Dataset. Blue dots represent source-domain samples, orange dots represent target-domain samples.
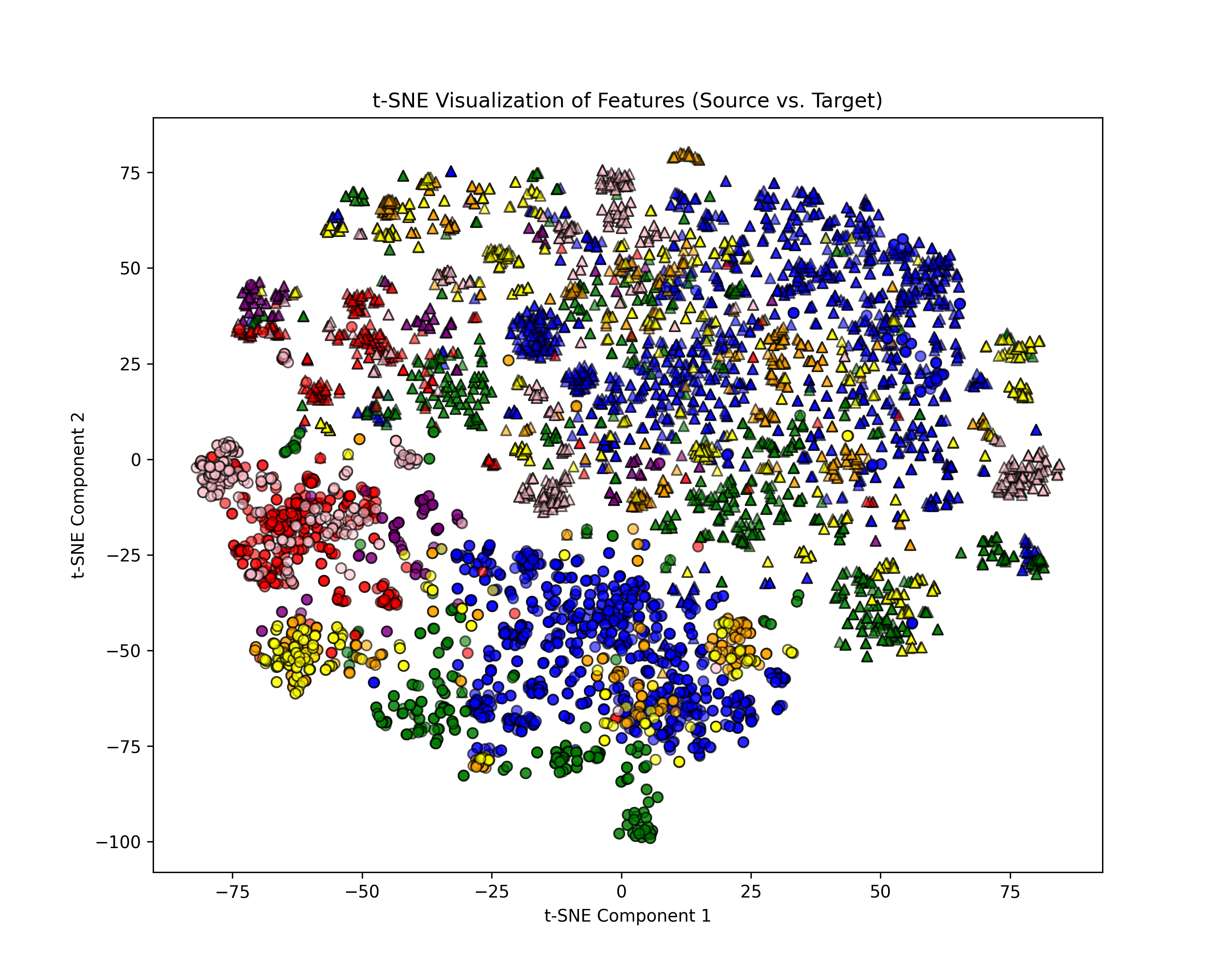
Fig. 2 Fine-grained class-conditional distribution comparison from sub-datasets D and N in the DN Dataset. Different colors indicate different classes: circles represent source-domain samples, and triangles represent target-domain samples.
| 模型 Model | 迁移任务 Transfer task D→N | 迁移任务 Transfer task N→D | Average-HOS (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| OS (%) | OS* (%) | UNK (%) | HOS (%) | OS (%) | OS* (%) | UNK (%) | HOS (%) | ||
| ResNet50 | 28.00 | 25.07 | 51.51 | 33.72 | 29.15 | 25.29 | 52.33 | 34.10 | 33.91 |
| DANN | 33.55 | 30.85 | 49.78 | 38.09 | 34.87 | 33.31 | 44.24 | 38.00 | 38.05 |
| ROS | 28.91 | 20.87 | 77.13 | 32.85 | 31.91 | 30.86 | 38.20 | 34.14 | 33.50 |
| ROS* | 24.36 | 20.58 | 47.05 | 28.63 | 30.25 | 28.89 | 38.42 | 32.98 | 30.81 |
| MTS | 31.78 | 37.08 | 25.78 | 30.41 | 22.15 | 25.84 | 39.26 | 31.17 | 30.79 |
| 本文方法 Ours | 43.21 | 41.21 | 55.16 | 47.18 | 44.32 | 40.30 | 68.39 | 50.72 | 48.95 |
Table 2 Comparison of experimental results of different models on the DN wildlife dataset
| 模型 Model | 迁移任务 Transfer task D→N | 迁移任务 Transfer task N→D | Average-HOS (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| OS (%) | OS* (%) | UNK (%) | HOS (%) | OS (%) | OS* (%) | UNK (%) | HOS (%) | ||
| ResNet50 | 28.00 | 25.07 | 51.51 | 33.72 | 29.15 | 25.29 | 52.33 | 34.10 | 33.91 |
| DANN | 33.55 | 30.85 | 49.78 | 38.09 | 34.87 | 33.31 | 44.24 | 38.00 | 38.05 |
| ROS | 28.91 | 20.87 | 77.13 | 32.85 | 31.91 | 30.86 | 38.20 | 34.14 | 33.50 |
| ROS* | 24.36 | 20.58 | 47.05 | 28.63 | 30.25 | 28.89 | 38.42 | 32.98 | 30.81 |
| MTS | 31.78 | 37.08 | 25.78 | 30.41 | 22.15 | 25.84 | 39.26 | 31.17 | 30.79 |
| 本文方法 Ours | 43.21 | 41.21 | 55.16 | 47.18 | 44.32 | 40.30 | 68.39 | 50.72 | 48.95 |
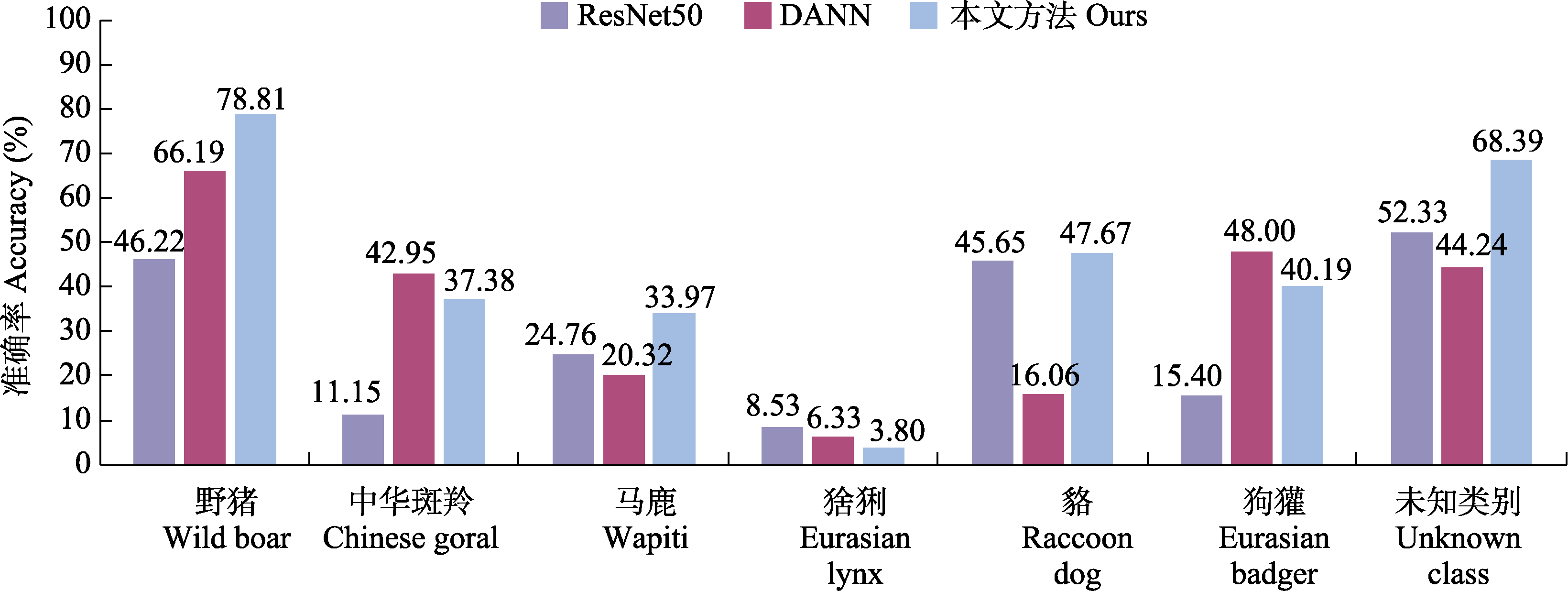
Fig. 6 Per-class recognition accuracy comparison under the N→D task. ResNet50 and DANN are comparative methods, while Ours denotes the proposed approach.
| 模型 Model | 迁移任务 Transfer task S1→S2 | 迁移任务 Transfer task S2→S1 | Average-HOS (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| OS (%) | OS* (%) | UNK (%) | HOS (%) | OS (%) | OS* (%) | UNK (%) | HOS (%) | ||
| ResNet50 | 28.00 | 25.07 | 51.51 | 33.72 | 37.91 | 37.25 | 43.15 | 39.99 | 36.85 |
| DANN | 32.58 | 30.22 | 51.40 | 38.06 | 39.52 | 37.92 | 52.27 | 43.96 | 41.01 |
| ROS | 23.58 | 18.39 | 65.10 | 28.68 | 23.94 | 19.37 | 60.51 | 29.34 | 29.01 |
| ROS* | 16.58 | 8.05 | 84.82 | 14.70 | 20.24 | 12.57 | 81.61 | 21.78 | 18.24 |
| MTS | 27.08 | 25.94 | 36.20 | 30.22 | 35.83 | 34.58 | 33.42 | 33.99 | 32.11 |
| 本文方法 Ours | 38.05 | 35.04 | 62.10 | 44.80 | 41.80 | 39.34 | 61.43 | 47.96 | 46.38 |
Table 3 Comparison of experimental results of different models on the S1S2 wildlife dataset
| 模型 Model | 迁移任务 Transfer task S1→S2 | 迁移任务 Transfer task S2→S1 | Average-HOS (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| OS (%) | OS* (%) | UNK (%) | HOS (%) | OS (%) | OS* (%) | UNK (%) | HOS (%) | ||
| ResNet50 | 28.00 | 25.07 | 51.51 | 33.72 | 37.91 | 37.25 | 43.15 | 39.99 | 36.85 |
| DANN | 32.58 | 30.22 | 51.40 | 38.06 | 39.52 | 37.92 | 52.27 | 43.96 | 41.01 |
| ROS | 23.58 | 18.39 | 65.10 | 28.68 | 23.94 | 19.37 | 60.51 | 29.34 | 29.01 |
| ROS* | 16.58 | 8.05 | 84.82 | 14.70 | 20.24 | 12.57 | 81.61 | 21.78 | 18.24 |
| MTS | 27.08 | 25.94 | 36.20 | 30.22 | 35.83 | 34.58 | 33.42 | 33.99 | 32.11 |
| 本文方法 Ours | 38.05 | 35.04 | 62.10 | 44.80 | 41.80 | 39.34 | 61.43 | 47.96 | 46.38 |
| 网络 Network | 迁移任务 Transfer task D→N | 迁移任务Transfer task N→D | Average-HOS (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| OS (%) | OS* (%) | UNK (%) | HOS (%) | OS (%) | OS* (%) | UNK (%) | HOS (%) | ||
| ResNet18 | 29.88 | 32.04 | 16.90 | 22.13 | 36.20 | 37.14 | 30.57 | 33.54 | 27.83 |
| ResNet34 | 36.19 | 32.05 | 61.03 | 42.03 | 37.41 | 38.46 | 31.09 | 34.38 | 38.21 |
| ResNet50 | 43.21 | 41.21 | 55.16 | 47.18 | 44.32 | 40.30 | 68.39 | 50.72 | 48.95 |
| ResNet101 | 35.55 | 32.40 | 54.46 | 40.63 | 27.16 | 28.32 | 20.21 | 23.59 | 32.11 |
| ResNet152 | 33.65 | 35.04 | 25.35 | 29.42 | 42.26 | 38.94 | 62.18 | 47.89 | 38.65 |
| Wide_ResNet50_2 | 33.14 | 29.16 | 57.04 | 38.59 | 44.62 | 48.51 | 21.24 | 29.55 | 34.07 |
| ResNext50_32x4d | 34.95 | 31.23 | 57.28 | 40.42 | 26.61 | 27.50 | 21.24 | 23.97 | 32.19 |
Table 4 Comparison of performance of feature extraction network
| 网络 Network | 迁移任务 Transfer task D→N | 迁移任务Transfer task N→D | Average-HOS (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|
| OS (%) | OS* (%) | UNK (%) | HOS (%) | OS (%) | OS* (%) | UNK (%) | HOS (%) | ||
| ResNet18 | 29.88 | 32.04 | 16.90 | 22.13 | 36.20 | 37.14 | 30.57 | 33.54 | 27.83 |
| ResNet34 | 36.19 | 32.05 | 61.03 | 42.03 | 37.41 | 38.46 | 31.09 | 34.38 | 38.21 |
| ResNet50 | 43.21 | 41.21 | 55.16 | 47.18 | 44.32 | 40.30 | 68.39 | 50.72 | 48.95 |
| ResNet101 | 35.55 | 32.40 | 54.46 | 40.63 | 27.16 | 28.32 | 20.21 | 23.59 | 32.11 |
| ResNet152 | 33.65 | 35.04 | 25.35 | 29.42 | 42.26 | 38.94 | 62.18 | 47.89 | 38.65 |
| Wide_ResNet50_2 | 33.14 | 29.16 | 57.04 | 38.59 | 44.62 | 48.51 | 21.24 | 29.55 | 34.07 |
| ResNext50_32x4d | 34.95 | 31.23 | 57.28 | 40.42 | 26.61 | 27.50 | 21.24 | 23.97 | 32.19 |
| 对抗学习 Adversarial training | 正交投影损失Orthogonal projection loss | 中心损失 Center loss | 迁移任务 Transfer task D→N | 迁移任务 Transfer task N→D | Average-HOS (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| OS (%) | OS* (%) | UNK (%) | HOS (%) | OS (%) | OS* (%) | UNK (%) | HOS (%) | ||||
| - | - | - | 28.00 | 25.07 | 51.51 | 33.72 | 29.15 | 25.29 | 52.33 | 34.10 | 33.91 |
| - | - | √ | 30.67 | 28.00 | 46.70 | 35.01 | 32.11 | 29.12 | 50.00 | 36.80 | 35.91 |
| - | √ | - | 31.00 | 28.27 | 47.36 | 35.40 | 26.53 | 20.61 | 62.04 | 30.94 | 33.17 |
| √ | - | - | 37.85 | 35.91 | 49.53 | 41.63 | 41.99 | 38.54 | 62.69 | 47.73 | 44.68 |
| - | √ | √ | 30.77 | 28.12 | 46.70 | 35.10 | 32.72 | 29.57 | 51.57 | 37.59 | 36.35 |
| √ | √ | - | 33.18 | 30.29 | 50.47 | 37.86 | 40.56 | 35.14 | 73.06 | 47.46 | 42.66 |
| √ | - | √ | 46.78 | 50.08 | 27.00 | 35.08 | 43.48 | 38.64 | 72.54 | 50.42 | 42.75 |
| √ | √ | √ | 43.21 | 41.21 | 55.16 | 47.18 | 44.32 | 40.30 | 68.39 | 50.72 | 48.95 |
Table 5 Ablation experimental results of model
| 对抗学习 Adversarial training | 正交投影损失Orthogonal projection loss | 中心损失 Center loss | 迁移任务 Transfer task D→N | 迁移任务 Transfer task N→D | Average-HOS (%) | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| OS (%) | OS* (%) | UNK (%) | HOS (%) | OS (%) | OS* (%) | UNK (%) | HOS (%) | ||||
| - | - | - | 28.00 | 25.07 | 51.51 | 33.72 | 29.15 | 25.29 | 52.33 | 34.10 | 33.91 |
| - | - | √ | 30.67 | 28.00 | 46.70 | 35.01 | 32.11 | 29.12 | 50.00 | 36.80 | 35.91 |
| - | √ | - | 31.00 | 28.27 | 47.36 | 35.40 | 26.53 | 20.61 | 62.04 | 30.94 | 33.17 |
| √ | - | - | 37.85 | 35.91 | 49.53 | 41.63 | 41.99 | 38.54 | 62.69 | 47.73 | 44.68 |
| - | √ | √ | 30.77 | 28.12 | 46.70 | 35.10 | 32.72 | 29.57 | 51.57 | 37.59 | 36.35 |
| √ | √ | - | 33.18 | 30.29 | 50.47 | 37.86 | 40.56 | 35.14 | 73.06 | 47.46 | 42.66 |
| √ | - | √ | 46.78 | 50.08 | 27.00 | 35.08 | 43.48 | 38.64 | 72.54 | 50.42 | 42.75 |
| √ | √ | √ | 43.21 | 41.21 | 55.16 | 47.18 | 44.32 | 40.30 | 68.39 | 50.72 | 48.95 |
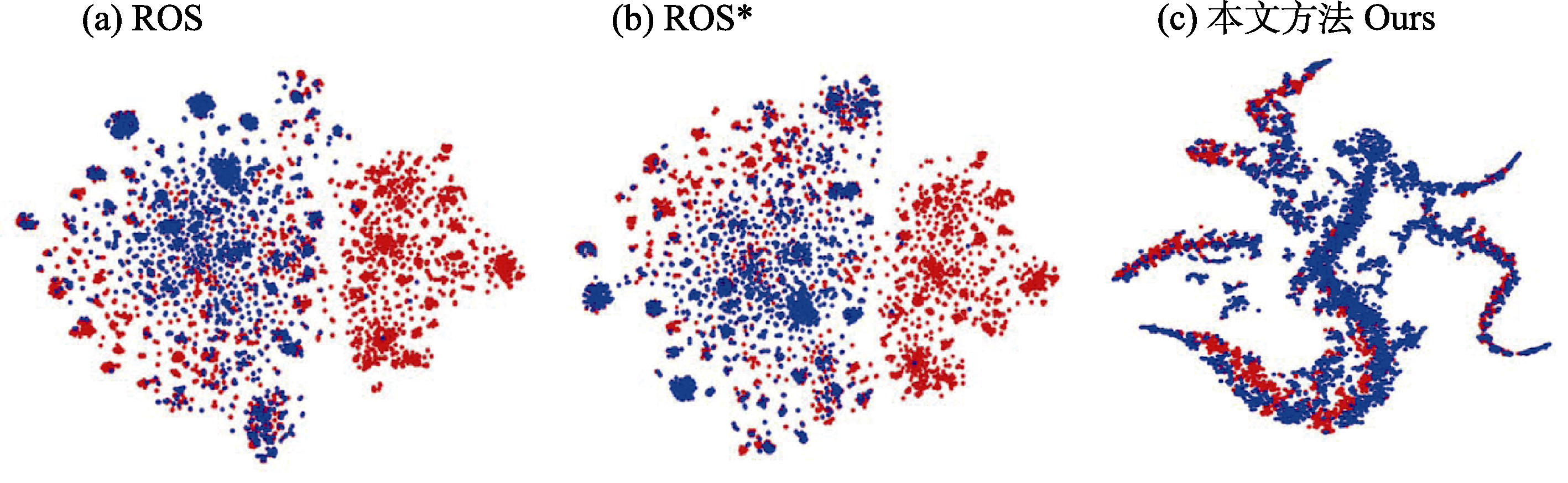
Fig. 8 t-SNE embedded visualization of feature distributions. ROS and ROS* denote the comparative methods used in this study, where ROS* represents the weighted version of ROS. Red dots represent source-domain features, and blue dots represent target-domain features.
| [1] | Bai SH, Zhang M, Zhou WQ, Huang ST, Luan ZR, Wang DL, Chen BD (2024) Prompt-based distribution alignment for unsupervised domain adaptation. In: 2024 AAAI Conference on Artificial Intelligence, pp.729-737. AAAI Press, Vancouver. |
| [2] | Bucci S, Loghmani MR, Tommasi T (2020) On the effectiveness of image rotation for open set domain adaptation. In: 2020 European Conference on Computer Vision (ECCV), pp.422-438. Springer International Publishing, Glasgow. |
| [3] | Busto PP, Gall J (2017) Open set domain adaptation. In: 2017 IEEE International Conference on Computer Vision (ICCV), pp. 754-763. IEEE, Venice. |
| [4] |
Chang DL, Sain A, Ma ZY, Song YZ, Wang RP, Guo J (2024) Mind the gap: Open set domain adaptation via mutual-to-separate framework. IEEE Transactions on Circuits and Systems for Video Technology, 34, 4159-4174.
DOI URL |
| [5] | Ganin Y, Lempitsky V (2015) Unsupervised domain adaptation by backpropagation.In: 2015 IEEE International Conference on Machine Learning (ICML), pp. 1180-1189. IEEE, Lille. |
| [6] |
Gao F, Pi DC, Chen JF (2024) Balanced and robust unsupervised open set domain adaptation via joint adversarial alignment and unknown class isolation. Expert Systems with Applications, 238, 122127.
DOI URL |
| [7] | He KM, Zhang XY, Ren SQ, Sun J (2016) Deep residual learning for image recognition. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), pp. 770-778. IEEE, Las Vegas. |
| [8] | He ZW, Zhang ZL, Zhang L (2024) Survey on cross-domain object detection in open environment. Journal of Computer-Aided Design & Computer Graphics, 36, 485-502. (in Chinese with English abstract) |
| [何贞苇, 张治龙, 张磊 (2024) 开放环境下的跨域物体检测综述. 计算机辅助设计与图形学学报, 36, 485-502.] | |
| [9] | Jing TT, Liu HF, Ding ZM (2021) Towards novel target discovery through open-set domain adaptation. In: 2021 IEEE International Conference on Computer Vision (ICCV), pp. 9302-9311. IEEE, Montreal. |
| [10] | Liu H, Cao ZJ, Long MH, Wang JM, Yang Q (2019) Separate to adapt: Open set domain adaptation via progressive separation. In: 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 2922-2931. IEEE, Long Beach. |
| [11] |
Loghmani MR, Vincze M, Tommasi T (2020) Positive-unlabeled learning for open set domain adaptation. Pattern Recognition Letters, 136, 198-204.
DOI URL |
| [12] |
Long SF, Wang SS, Zhao X, Fu ZH, Wang BL (2023) Sample separation and domain alignment complementary learning mechanism for open set domain adaptation. Applied Intelligence, 53, 18790-18805.
DOI |
| [13] |
Lu JY, Shi XY, Duo LA, Wang TM, Li ZL (2024) Circadian rhythms of urban terrestrial mammals in Tianjin based on camera trapping method. Biodiversity Science, 32, 23369. (in Chinese with English abstract)
DOI |
|
[卢佳玉, 石小亿, 多立安, 王天明, 李治霖 (2024) 基于红外相机技术的天津城市地栖哺乳动物昼夜活动节律评价. 生物多样性, 32, 23369.]
DOI |
|
| [14] |
Ma ZB, Dong YQ, Xia Y, Xu DL, Xu F, Chen FX (2024) Wildlife real-time detection in complex forest scenes based on YOLOv5s deep learning network. Remote Sensing, 16, 1350.
DOI URL |
| [15] | Qu SQ, Zou TP, He LH, Röhrbein F, Knoll A, Chen G, Jiang CJ (2024) LEAD: Learning decomposition for source-free universal domain adaptation.In: 2024 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 23334-23343. IEEE, Seattle. |
| [16] | Ranasinghe K, Naseer M, Hayat M, Khan S, Khan FS (2021) Orthogonal projection loss.In: 2021 IEEE/CVF International Conference on Computer Vision (ICCV), pp. 12313-12323. IEEE, Montreal. |
| [17] |
Roy AM, Bhaduri J, Kumar T, Raj K (2023) WilDect-YOLO: An efficient and robust computer vision-based accurate object localization model for automated endangered wildlife detection. Ecological Informatics, 75, 101919.
DOI URL |
| [18] | Saito K, Yamamoto S, Ushiku Y, Harada T (2018) Open set domain adaptation by backpropagation. In: 2018 European Conference on Computer Vision (ECCV), pp.156-171. Springer International Publishing, Munich. |
| [19] | Tabak MA, Norouzzadeh MS, Wolfson DW, Sweeney SJ, Vercauteren KC, Snow NP, Halseth JM, Salvo PA, Lewis JS, White MD, Teton B, Beasley JC, Schlichting PE, Boughton RK, Wight B, Newkirk ES, Ivan JS, Odell EA, Brook RK, Lukacs PM, Moeller AK, Mandeville EG, Clune J, Miller RS (2019) Machine learning to classify animal species in camera trap images: Applications in Ecology.Methods in Ecology and Evolution, 10, 585-590. |
| [20] | Tan SH, Jiao JN, Zheng WS (2019) Weakly supervised open-set domain adaptation by dual-domain collaboration.In: 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), pp. 5389-5398. IEEE, Long Beach. |
| [21] |
Wang DW, Cheng S, Feng JW, Wang TM (2025) The wildlife camera-trapping dataset of Zhangguangcai Mountains in Northeast China (2015-2020). Biodiversity Science, 33, 24384. (in Chinese with English abstract)
DOI |
|
[王大伟, 程帅, 冯佳伟, 王天明 (2025) 东北地区张广才岭2015-2020年野生动物红外相机监测数据集. 生物多样性, 33, 24384.]
DOI |
|
| [22] | Wen YD, Zhang KP, Li ZF, Qiao Y (2016) A discriminative feature learning approach for deep face recognition. In: 2016 European Conference on Computer Vision (ECCV), pp. 499-515. Springer International Publishing, Amsterdam. |
| [23] |
Xiao ZS, Xiao WH, Wang TM, Li S, Lian XM, Song DZ, Deng XQ, Zhou QH (2022) Wildlife monitoring and research using camera-trapping technology across China: The current status and future issues. Biodiversity Science, 30, 22451. (in Chinese with English abstract)
DOI |
|
[肖治术, 肖文宏, 王天明, 李晟, 连新明, 宋大昭, 邓雪琴, 周岐海 (2022) 中国野生动物红外相机监测与研究: 现状及未来. 生物多样性, 30, 22451.]
DOI |
|
| [24] | Yousif H, Kays R, He Z (2019) Dynamic programming selection of object proposals for sequence-level animal species classification in the wild. IEEE Transactions on Circuits and Systems for Video Technology, 2, 28-29. |
| [25] |
Zhang CC, Zhang JG (2023) DJAN: Deep joint adaptation network for wildlife image recognition. Animals, 13, 3333.
DOI URL |
| [26] | Zhao ET, Zhang CC, Zhao HT, Zhang JG (2025) A recognition method of domain adaptation for wildlife images based on adversarial learning. Scientia Silvae Sinicae, 61(4), 1-8. (in Chinese with English abstract) |
| [赵恩庭, 张长春, 赵海涛, 张军国 (2025) 基于对抗学习的野生动物图像域适应识别方法. 林业科学, 61(4), 1-8.] | |
| [27] | Zheng F, Lü LX, Lü HX, Shi J, Li R, Yang SL, Liu BW (2025) The research on the mammal diversity and its dynamic based on camera-trapping in Heilongjiang Liangshui National Nature Reserve. Acta Ecologica Sinica, 45, 5289-5296. (in Chinese with English abstract) |
| [郑霏, 吕来新, 吕泓学, 师杰, 李瑞, 杨世丽, 刘丙万 (2025) 基于红外相机监测黑龙江凉水国家级自然保护区兽类多样性及其变化. 生态学报, 45, 5289-5296.] | |
| [28] |
Zheng JP, Wen YB, Chen MX, Yuan S, Li WJ, Zhao Y, Wu WZ, Zhang LX, Dong RM, Fu HH (2024) Open-set domain adaptation for scene classification using multi-adversarial learning. ISPRS Journal of Photogrammetry and Remote Sensing, 208, 245-260.
DOI URL |
| [29] |
Zhong L, Fang Z, Liu F, Yuan B, Zhang GQ, Lu J (2023) Bridging the theoretical bound and deep algorithms for open set domain adaptation. IEEE Transactions on Neural Networks and Learning Systems, 34, 3859-3873.
DOI URL |
| [1] | Lin Ji, Chenxun Deng, Lifeng Wang, Degang Wang, Jiantao Wang, Yongyong Yu, Junguo Zhang. A wildlife recognition method for skewed distributions in the Ulanba Nature Reserve [J]. Biodiv Sci, 2026, 34(2): 25256-. |
| [2] | Xiaoyu Sun. Legal approaches to resolving human–wildlife conflicts in national parks of China [J]. Biodiv Sci, 2026, 34(1): 25040-. |
| [3] | Luyao Tian, Hao Yin. Research status and strategies for China’s ecological railway development based on biodiversity conservation [J]. Biodiv Sci, 2025, 33(8): 24495-. |
| [4] | Junni Zheng, Yuanlingbo Shang, Yao Luo, Ying Wei, Zhiwei Gao, Zongze Zhou, Lingjuan Liao, Daode Yang. Refining the adjustment method for local key protected wildlife lists: A case study of terrestrial vertebrates in Hunan Province, China [J]. Biodiv Sci, 2025, 33(8): 25055-. |
| [5] | Jing Mao, Jing Wang, Jie Huang, Shuhong Xiong, Ziliang Zhang, Youxiang Zhang, Tao Wu. Bird and mammal diversity monitoring dataset in Gaowangjie National Nature Reserve (2021-2023) [J]. Biodiv Sci, 2025, 33(6): 24489-. |
| [6] | Fu Mengdi, Zhu Yanpeng, Ren Yueheng, Li Shuang, Qin Le, Xie Zhengjun, Wang Qingchun, Zhang Libo. Research on the optimization of wildlife passage spatial layout in Xinjiang [J]. Biodiv Sci, 2025, 33(3): 24346-. |
| [7] | Wang Dawei, Cheng Shuai, Feng Jiawei, Wang Tianming. The wildlife camera-trapping dataset of Zhangguangcai Mountains in Northeast China (2015-2020) [J]. Biodiv Sci, 2025, 33(2): 24384-. |
| [8] | Sicheng Han, Daowei Lu, Yuchen Han, Ruohan Li, Jing Yang, Ge Sun, Lu Yang, Junwei Qian, Xiang Fang, Shu-Jin Luo. Distribution of leopard cats in the nearest mountains to urban Beijing and its affecting environmental factors [J]. Biodiv Sci, 2024, 32(8): 24138-. |
| [9] | Suyan Ba, Chunyan Zhao, Yuan Liu, Qiang Fang. Constructing a pollination network by identifying pollen on insect bodies: Consistency between human recognition and an AI model [J]. Biodiv Sci, 2024, 32(6): 24088-. |
| [10] | Baican Li, Junguo Zhang, Changchun Zhang, Lifeng Wang, Jiliang Xu, Li Liu. Rare bird recognition method in Beijing based on TC-YOLO model [J]. Biodiv Sci, 2024, 32(5): 24056-. |
| [11] | Binyue Lu, Kun Li, Chenxi Wang, Sheng Li. The application and outlook of wildlife tracking using sensor-based tags in China [J]. Biodiv Sci, 2024, 32(5): 23497-. |
| [12] | Jinfeng Chen, Xinjing Wu, Hai Lin, Guofa Cui. A comparative analysis of the List of State Key Protected Wild Animals and other wildlife protection lists [J]. Biodiv Sci, 2023, 31(6): 22639-. |
| [13] | Haigang Ma, Penglai Fan. Application, progress, and future perspective of passive acoustic monitoring in terrestrial mammal research [J]. Biodiv Sci, 2023, 31(1): 22374-. |
| [14] | Yiqing Wang, Ziyu Ma, Gang Wang, Yanlin Liu, Dazhao Song, Beibei Liu, Lu Li, Xinguo Fan, Qiaowen Huang, Sheng Li. Spatiotemporal patterns of cattle depredation by the North Chinese leopard in Taihang Mountains and its management strategy: A case study in Heshun, Shanxi Province [J]. Biodiv Sci, 2022, 30(9): 21510-. |
| [15] | Rui Song, Jing Deng, Tao Qin. Development dilemma and optimization path of public liability insurance for wildlife accidents [J]. Biodiv Sci, 2022, 30(7): 22291-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn