



Biodiv Sci ›› 2014, Vol. 22 ›› Issue (1): 97-108. DOI: 10.3724/SP.J.1003.2014.13135 cstr: 32101.14.SP.J.1003.2014.13135
Special Issue: 基因组和生物多样性; 传粉生物学
• Orginal Article • Previous Articles Next Articles
Received:2013-06-06
Accepted:2013-10-09
Online:2014-01-20
Published:2014-02-10
Contact:
Wang Hong
Wei Zhou,Hong Wang. Pollen dispersal analysis using DNA markers[J]. Biodiv Sci, 2014, 22(1): 97-108.
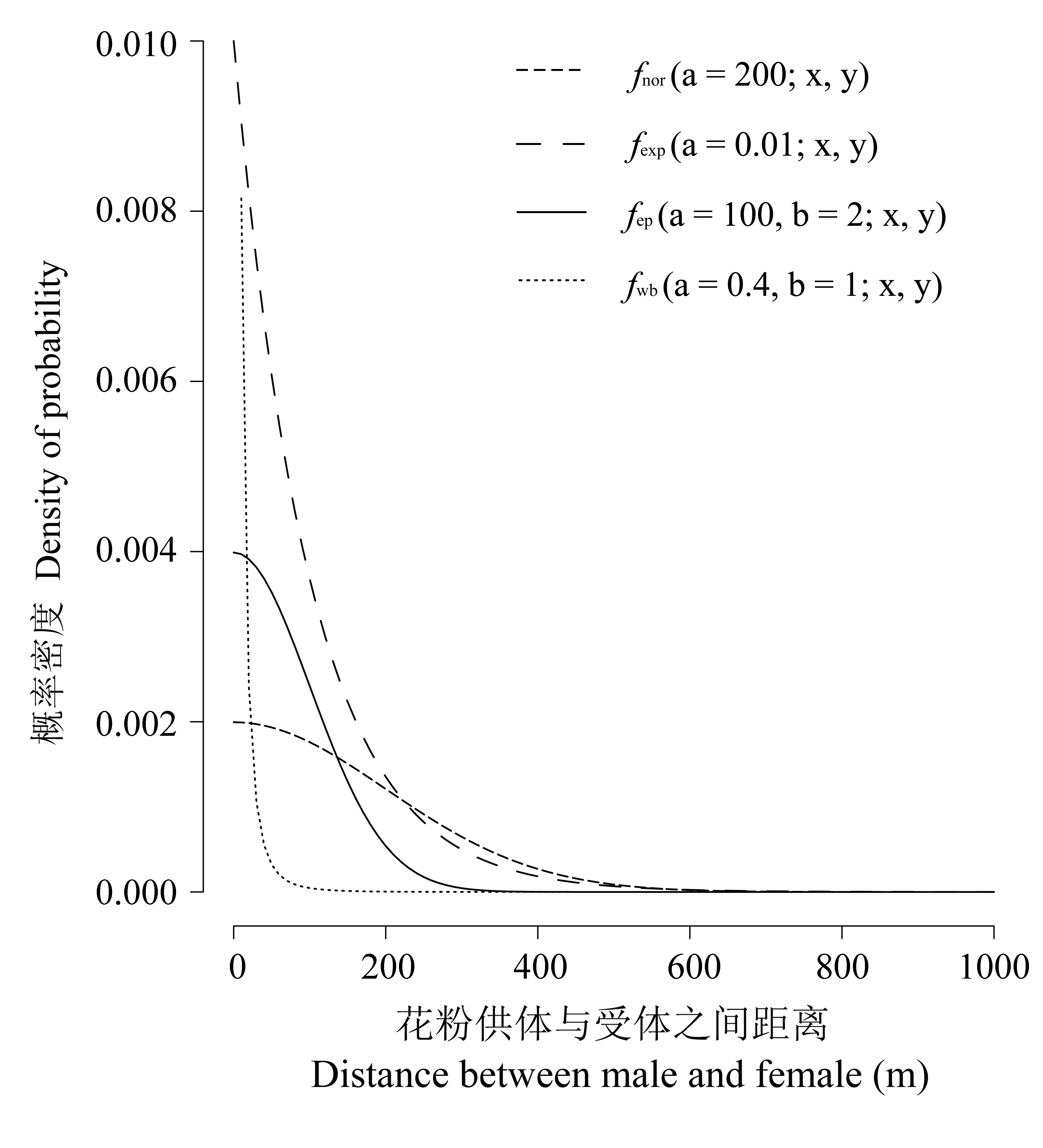
Fig. 2 Example of curves and the characters of tail for the different pollen dispersal models. Single parameter: normal distribution model, exponential distribution model; Two parameters: exponential power distribution model and Weibull distribution model.
| [1] | Abraham ST, Zaya DN, Koenig WD, Ashley MV (2011) Interspecific and intraspecific pollination patterns of valley oak, Quercus lobata, in a mixed stand in coastal central California.International Journal of Plant Sciences, 172, 691-699. |
| [2] | Abramowitz M, Stegun IA (1964) Handbook of Mathematical Functions with Formulas, Graphs, and Mathematical Tables. U.S. Government Printing Office, Washington D.C. |
| [3] | Adams WT, Griffin AR, Moran GF (1992) Using paternity analysis to measure effective pollen dispersal in plant populations.The American Naturalist, 140, 762-780. |
| [4] | Adler LS, Irwin RE (2006) Comparison of pollen transfer dynamics by multiple floral visitors: experiments with pollen and fluorescent dye.Annals of Botany, 97, 141-150. |
| [5] | Ahmed S, Compton SG, Butlin RK, Gilmartin PM (2009) Wind-borne insects mediate directional pollen transfer between desert fig trees 160 kilometers apart. Proceedings of the National Academy of Sciences,USA, 106, 20342-20347. |
| [6] | Anderson EC, Garza JC (2006) The power of single-nucleotide polymorphisms for large-scale parentage inference.Genetics, 172, 2567-2582. |
| [7] | Ashley MV (2010) Plant parentage, pollination, and dispersal: how DNA microsatellites have altered the landscape.Critical Reviews in Plant Sciences, 29, 148-161. |
| [8] | Ashley MV, Wilk JA, Styan SMN, Craft KJ, Jones KL, Feldheim KA, Lewers KS, Ashman TL (2003) High variability and disomic segregation of microsatellites in the octoploid Fragaria virginiana Mill. (Rosaceae).Theoretical and Applied Genetics, 107, 1201-1207. |
| [9] | Austerlitz F, Dick CW, Dutech C, Klein EK, Oddou-Muratorio S, Smouse PE, Sork VL (2004) Using genetic markers to estimate the pollen dispersal curve.Molecular Ecology, 13, 937-954. |
| [10] | Austerlitz F, Gleiser G, Teixeira S, Bernasconi G (2012) The effects of inbreeding, genetic dissimilarity and phenotype on male reproductive success in a dioecious plant.Proceedings of the Royal Society B: Biological Sciences, 279, 91-100. |
| [11] | Bacles CFE, Ennos RA (2008) Paternity analysis of pollen-mediated gene flow for Fraxinus excelsior L. in a chronically fragmented landscape.Heredity, 101, 368-380. |
| [12] | Baena-Díaz F, Fornoni J, Sosenski P, Molina-Freaner FE, Weller SG, Pérez-Ishiwara R, Domínguez CA (2012) Changes in reciprocal herkogamy during the tristyly-distyly transition in Oxalis alpina increase efficiency in pollen transfer.Journal of Evolutionary Biology, 25, 574-583. |
| [13] | Bai WN, Zeng YF, Zhang DY (2007) Mating patterns and pollen dispersal in a heterodichogamous tree, Juglans mandshurica (Juglandaceae).New Phytologist, 176, 699-707. |
| [14] | Baird NA, Etter PD, Atwood TS, Currey MC, Shiver AL, Lewis ZA, Selker EU, Cresko WA, Johnson EA (2008) Rapid SNP discovery and genetic mapping using sequenced RAD markers.PLoS ONE, 3, e3376, doi: 10.1371/journal. pone.0003376. |
| [15] | Barrett SCH (2010) Darwin’s legacy: the forms, function and sexual diversity of flowers.Philosophical Transactions of the Royal Society B: Biological Sciences, 365, 351-368. |
| [16] | Burgarella C, Lorenzo Z, Jabbour-Zahab R, Lumaret R, Guichoux E, Petit RJ, Soto A, Gil L (2009) Detection of hybrids in nature: application to oaks (Quercus suber and Q. ilex).Heredity, 102, 442-452. |
| [17] | Byrne M, Elliott CP, Yates CJ, Coates DJ (2008) Maintenance of high pollen dispersal in Eucalyptus wandoo, a dominant tree of the fragmented agricultural region in Western Australia.Conservation Genetics, 9, 97-105. |
| [18] | Campbell DR (1991) Comparing pollen dispersal and gene flow in a natural population.Evolution, 45, 1965-1968. |
| [19] | Castillo RA, Cordero C, Domínguez CA (2002) Are reward polymorphisms subject to frequency- and density-dependent selection? Evidence from a monoecious species pollinated by deceit.Journal of Evolutionary Biology, 15, 544-552. |
| [20] | Cavagnaro PF, Senalik DA, Yang LM, Simon PW, Harkins TT, Kodira CD, Huang SW, Weng YQ (2010) Genome-wide characterization of simple sequence repeats in cucumber (Cucumis sativus L.).BMC Genomics, 11, 569. |
| [21] | Chakrabo R, Shaw M, Schull W (1974) Exclusion of paternity- current state of art.American Journal of Human Genetics, 26, 477-488. |
| [22] | Christie MR, Tennessen JA, Blouin MS (2013) Bayesian paren-tage analysis with systematic accountability of genotyping error, missing data and false matching.Bioinfor- matics, 29, 725-732. |
| [23] | Clark JS (1998) Why trees migrate so fast: confronting theory with dispersal biology and the paleorecord.The American Naturalist, 152, 204-224. |
| [24] | Craft KJ, Ashley MV, Koenig WD (2002) Limited hybridization between Quercus lobata and Quercus douglasii (Fagaceae) in a mixed stand in central coastal California.American Journal of Botany, 89, 1792-1798. |
| [25] | Craft KJ, Ashley MV (2010) Pollen-mediated gene flow in isolated and continuous stands of bur oak, Quercus macrocarpa (Fagaceae).American Journal of Botany, 97, 1999-2006. |
| [26] | Curtu AL, Gailing O, Finkeldey R (2007) Evidence for hybridization and introgression within a species-rich oak (Quercus spp.) community.BMC Evolutionary Biology, 7, 218. |
| [27] | Curtu AL, Gailing O, Finkeldey R (2009) Patterns of contemporary hybridization inferred from paternity analysis in a four-oak-species forest.BMC Evolutionary Biology, 9, 284. |
| [28] | Dakin EE, Avise JC (2004) Microsatellite null alleles in parentage analysis.Heredity, 93, 504-509. |
| [29] | Davey JW, Hohenlohe PA, Etter PD, Boone JQ, Catchen JM, Blaxter ML (2011) Genome-wide genetic marker discovery and genotyping using next-generation sequencing.Nature Reviews Genetics, 12, 499-510. |
| [30] | Dick CW, Etchelecu G, Austerlitz F (2003) Pollen dispersal of tropical trees (Dinizia excelsa: Fabaceae) by native insects and African honeybees in pristine and fragmented Amazonian rainforest.Molecular Ecology, 12, 753-764. |
| [31] | DiFazio SP, Leonardi S, Slavov GT, Garman SL, Adams WT, Strauss SH (2012) Gene flow and simulation of transgene dispersal from hybrid poplar plantations.New Phytologist, 193, 903-915. |
| [32] | Dunphy BK, Hamrick JL (2005) Gene flow among established Puerto Rican populations of the exotic tree species, Albizia lebbeck.Heredity, 94, 418-425. |
| [33] | Dunphy BK, Hamrick JL (2007) Estimation of gene flow into fragmented populations of Bursera simaruba (Burseraceae) in the dry-forest life zone of Puerto Rico.American Journal of Botany, 94, 1786-1794. |
| [34] | Eckert KA, Hile SE (2009) Every microsatellite is different: intrinsic DNA features dictate mutagenesis of common microsatellites present in the human genome.Molecular Carcinogenesis, 48, 379-388. |
| [35] | Ellegren H (2004) Microsatellites: simple sequences with complex evolution.Nature Reviews Genetics, 5, 435-445. |
| [36] | Field DL, Barrett SCH (2012) Disassortative mating and the maintenance of sexual polymorphism in painted maple.Molecular Ecology, 21, 3640-3643. |
| [37] | Fuchs EJ, Hamrick JL (2011) Mating system and pollen flow between remnant populations of the endangered tropical tree, Guaiacum sanctum (Zygophyllaceae).Conservation Genetics, 12, 175-185. |
| [38] | Geng QF, Lian CL, Goto S, Tao JM, Kimura M, Islam MDS, Hogetsu T (2008) Mating system, pollen and propagule dispersal, and spatial genetic structure in a high-density population of the mangrove tree Kandelia candel.Molecular Ecology, 17, 4724-4739. |
| [39] | Gerber S, Mariette S, Streiff R, Bodenes C, Kremer A (2000) Comparison of microsatellites and amplified fragment length polymorphism markers for parentage analysis.Molecular Ecology, 9, 1037-1048. |
| [40] | Gleiser G, Segarra-Moragues JG, Pannell JR, Verdu M (2008a) Siring success and paternal effects in heterodichogamous Acer opalus.Annals of Botany, 101, 1017-1026. |
| [41] | Gleiser G, Verdu M, Segarra-Moragues JG, Gonzalez-Martinez SC, Pannell JR (2008b) Disassortative mating, sexual specialization, and the evolution of gender dimorphism in heterodichogamous Acer opalus.Evolution, 62, 1676-1688. |
| [42] | Goto S, Shimatani K, Yoshimaru H, Takahashi Y (2006) Fat-tailed gene flow in the dioecious canopy tree species Fraxinus mandshurica var. japonica revealed by microsatellites.Molecular Ecology, 15, 2985-2996. |
| [43] | Hadfield JD, Richardson DS, Burke T (2006) Towards unbiased parentage assignment: combining genetic, behavioural and spatial data in a Bayesian framework.Molecular Ecology, 15, 3715-3730. |
| [44] | Hadley AS, Betts MG (2009) Tropical deforestation alters hummingbird movement patterns.Biology Letters, 5, 207-210. |
| [45] | Hasegawa Y, Suyama Y, Seiwa K (2009) Pollen donor composition during the early phases of reproduction revealed by DNA genotyping of pollen grains and seeds of Castanea crenata.New Phytologist, 182, 994-1002. |
| [46] | Hauser L, Baird M, Hilborn R, Seeb LW, Seeb JE (2011) An empirical comparison of SNPs and microsatellites for parentage and kinship assignment in a wild sockeye salmon (Oncorhynchus nerka) population.Molecular Ecology Resources, 11, 150-161. |
| [47] | He TH (何田华), Ge S (葛颂) (2001) Mating system, paternity analysis and gene flow in plant populations.Acta Phytoecologica Sinica(植物生态学报), 25, 144-154. (in Chinese with English abstract) |
| [48] | Hodgins KA, Barrett SCH (2008) Asymmetrical mating patterns and the evolution of biased style-morph ratios in a tristylous daffodil.Genetics Research, 90, 3-15. |
| [49] | Hoffman JI, Amos W (2005) Does kin selection influence fostering behaviour in Antarctic fur seals (Arctocephalus gazella)?Proceedings of the Royal Society B: Biological Sciences, 272, 2017-2022. |
| [50] | Huang SQ, Guo YH (2000) New advances in pollination biology and the studies in China.Chinese Science Bulletin, 45, 1441-1447. |
| [51] | Ishihama F, Ueno S, Tsumura Y, Washitani I (2006) Effects of density and floral morph on pollen flow and seed reproduction of an endangered heterostylous herb. Primula sieboldii.Journal of Ecology, 94, 846-855. |
| [52] | Jiang ZX, Xia HB, Basso B, Lu BR (2012) Introgression from cultivated rice influences genetic differentiation of weedy rice populations at a local spatial scale.Theoretical and Applied Genetics, 124, 309-322. |
| [53] | Kamm U, Gugerli F, Rotach P, Edwards P, Holderegger R (2010) Open areas in a landscape enhance pollen-mediated gene flow of a tree species: evidence from northern Switzerland.Landscape Ecology, 25, 903-911. |
| [54] | Karron JD, Ivey CT, Mitchell RJ, Whitehead MR, Peakall R, Case AL (2012) New perspectives on the evolution of plant mating systems.Annals of Botany, 109, 493-503. |
| [55] | Kelkar YD, Tyekucheva S, Chiaromonte F, Makova KD (2008) The genome-wide determinants of human and chimpanzee microsatellite evolution.Genome Research, 18, 30-38. |
| [56] | Kikuchi S, Shibata M, Tanaka H, Yoshimaru H, Niiyama K (2009) Analysis of the disassortative mating pattern in a heterodichogamous plant, Acer mono Maxim. using microsatellite markers.Plant Ecology, 204, 43-54. |
| [57] | Kimura MK, Goto S, Suyama Y, Matsui M, Woeste K, Seiwa K (2012) Morph-specific mating patterns in a low-density population of a heterodichogamous tree, Juglans ailantifolia.Plant Ecology, 213, 1477-1487. |
| [58] | Kitamoto N, Ueno S, Takenaka A, Tsumura Y, Washitani I, Ohsawa R (2006) Effect of flowering phenology on pollen flow distance and the consequences for spatial genetic structure within a population of Primula sieboldii (Primulaceae).American Journal of Botany, 93, 226-233. |
| [59] | Kitamoto N, Ueno S, Tsumura Y, Washitani I, Ohsawa R (2008) Effect of population density of compatible neighbours on inbreeding level within a Primula sieboldii population.Ecological Research, 23, 307-315. |
| [60] | Lander TA, Boshier DH, Harris SA (2010) Fragmented but not isolated: contribution of single trees, small patches and long-distance pollen flow to genetic connectivity for Gomortega keule, an endangered Chilean tree.Biological Conservation, 143, 2583-2590. |
| [61] | Leonarduzzi C, Leonardi S, Menozzi P, Piotti A (2012) Towards an optimal sampling effort for paternity analysis in forest trees: what do the raw numbers tell us?iForest, 5, 18-25. |
| [62] | Lepais O, Gerber S (2011) Reproductive patterns shape introgression dynamics and species succession within the European white oak species complex.Evolution, 65, 156-170. |
| [63] | Levin DA, Kerster HW, Niedzlek M (1971) Pollinator flight directionality and its effect on pollen flow.Evolution, 25, 113-118. |
| [64] | Lewis PO, Snow AA (1992) Deterministic paternity exclusion using RAPD markers.Molecular Ecology, 1, 155-160. |
| [65] | Lexer C, Heinze B, Gerber S, Macalka-Kampfer S, Steinkeller H, Kremer A, Glossl J (2000) Microsatellite analysis of maternal half-sib families of Quercus robur, pedunculate oak: II. inferring the number of pollen donors from the offspring.Theoretical and Applied Genetics, 100, 858-865. |
| [66] | Li JM, Koski MH, Ashman TL (2012) Functional characteriza- tion of gynodioecy in Fragaria vesca ssp. bracteata (Rosaceae).Annals of Botany, 109, 545-552. |
| [67] | Li QJ, Xu ZF, Kress WJ, Xia YM, Zhang L, Deng XB, Gao JY, Bai ZL (2001) Pollination: flexible style that encourages outcrossing.Nature, 410, 432. |
| [68] | Li YC, Korol AB, Fahima T, Beiles A, Nevo E (2002) Microsatellites: genomic distribution, putative functions and mutational mechanisms: a review.Molecular Ecology, 11, 2453-2465. |
| [69] | Marshall TC, Slate J, Kruuk LEB, Pemberton JM (1998) Statistical confidence for likelihood-based paternity inference in natural populations.Molecular Ecology, 7, 639-655. |
| [70] | Meagher TR, Thompson E (1986) The relationship between single parent and parent pair genetic likelihoods in genealogy reconstruction.Theoretical Population Biology, 29, 87-106. |
| [71] | Neff BD, Repka J, Gross MR (2001) A Bayesian framework for parentage analysis: the value of genetic and other biological data.Theoretical Population Biology, 59, 315-331. |
| [72] | Nielsen R, Mattila DK, Clapham PJ, Palsboll PJ (2001) Statistical approaches to paternity analysis in natural populations and applications to the North Atlantic humpback whale.Genetics, 157, 1673-1682. |
| [73] | Niggemann M, Wiegand T, Robledo-Arnuncio JJ, Bialozyt R (2012) Marked point pattern analysis on genetic paternity data for uncertainty assessment of pollen dispersal kernels.Journal of Ecology, 100, 264-276. |
| [74] | Nilsson LA, Rabakonandrianina E, Pettersson B (1992) Exact tracking of pollen transfer and mating in plants.Nature, 360, 666-668. |
| [75] | O’Connell LM, Mosseler A, Rajora OP (2006) Impacts of forest fragmentation on the mating system and genetic diversity of white spruce (Picea glauca) at the landscape level.Heredity, 97, 418-426. |
| [76] | Oddou-Muratorio S, Klein EK, Austerlitz F (2005) Pollen flow in the wildservice tree, Sorbus torminalis (L.) Crantz. II. Pollen dispersal and heterogeneity in mating success inferred from parent-offspring analysis.Molecular Ecology, 14, 4441-4452. |
| [77] | Ottewell K, Grey E, Castillo F, Karubian J (2012) The pollen dispersal kernel and mating system of an insect-pollinated tropical palm, Oenocarpus bataua.Heredity, 109, 332-339. |
| [78] | Parra V, Vargas CF, Eguiarte LE (1993) Reproductive-biology, pollen and seed dispersal, and neighborhood size in the hu- mmingbird-pollinated Echeveria gibbiflora (Crassulaceae).American Journal of Botany, 80, 153-159. |
| [79] | Pasquet RS, Peltier A, Hufford MB, Oudin E, Saulnier J, Paul L, Knudsen JT, Herren HR, Gepts P (2008) Long-distance pollen flow assessment through evaluation of pollinator foraging range suggests transgene escape distances. Proceedings of the National Academy of Sciences,USA, 105, 13456-13461. |
| [80] | Penaloza-Ramirez JM, Gonzalez-Rodriguez A, Mendoza- Cuenca L, Caron H, Kremer A, Oyama K (2010) Interspecific gene flow in a multispecies oak hybrid zone in the Sierra Tarahumara of Mexico.Annals of Botany, 105, 389-399. |
| [81] | Pons E, Navarro A, Ollitrault P, Peña L (2011) Pollen competition as a reproductive isolation barrier represses transgene flow between compatible and co-flowering citrus genotypes.PLoS ONE, 6, e25810, doi: 10.1371/journal. pone.0025810. |
| [82] | Pospiskova M, Salkova I (2006) Population structure and parentage analysis of black poplar along the Morava River.Canadian Journal of Forest Research, 36, 1067-1076. |
| [83] | Rahmé J, Widmer A, Karrenberg S (2009) Pollen competition as an asymmetric reproductive barrier between two closely related Silene species.Journal of Evolutionary Biology, 22, 1937-1943. |
| [84] | Robledo-Arnuncio JJ (2011) Wind pollination over mesoscale distances: an investigation with Scots pine.New Phytologist, 190, 222-233. |
| [85] | Robledo-Arnuncio JJ, Austerlitz F, Smouse PE (2006) A new method of estimating the pollen dispersal curve inde- pendently of effective density.Genetics, 173, 1033-1045. |
| [86] | Robledo-Arnuncio JJ, Austerlitz F, Smouse PE (2007) POLD- ISP: a software package for indirect estimation of contem- porary pollen dispersal.Molecular Ecology Notes, 7, 763-766. |
| [87] | Robledo-Arnuncio JJ, Gil L (2005) Patterns of pollen dispersal in a small population of Pinus sylvestris L. revealed by total-exclusion paternity analysis.Heredity, 94, 13-22. |
| [88] | Rymer PD, Johnson SD, Savolainen V (2010) Pollinator behaviour and plant speciation: Can assortative mating and disruptive selection maintain distinct floral morphs in sympatry?New Phytologist, 188, 426-436. |
| [89] | Salvini D, Bruschi P, Fineschi S, Grossoni P, Kjaer ED, Vendramin GG (2009) Natural hybridization between Quercus petraea (Matt.) Liebl. and Quercus pubescens Willd. within an Italian stand as revealed by microsatellite fingerprinting.Plant Biology, 11, 758-765. |
| [90] | Seeb JE, Carvalho G, Hauser L, Naish K, Roberts S, Seeb LW (2011) Single-nucleotide polymorphism (SNP) discovery and applications of SNP genotyping in nonmodel organisms.Molecular Ecology Resources, 11, 1-8. |
| [91] | Shang H, Luo YB, Bai WN (2012) Influence of asymmetrical mating patterns and male reproductive success on the maintenance of sexual polymorphism in Acer pictum subsp. mono (Aceraceae).Molecular Ecology, 21, 3869-3878. |
| [92] | Slavov GT, Leonardi S, Burczyk J, Adams WT, Strauss SH, Difazio SP (2009) Extensive pollen flow in two ecologically contrasting populations of Populus trichocarpa.Molecular Ecology, 18, 357-373. |
| [93] | Spigler RB, Ashman TL (2012) Gynodioecy to dioecy: Are we there yet?Annals of Botany, 109, 531-543. |
| [94] | Thomson JD, Price MV, Waser NM, Stratton DA (1986) Comparative studies of pollen and fluorescent dye transport by bumble bees visiting Erythronium grandiflorum.Oecologia, 69, 561-566. |
| [95] | Tokarska M, Marshall T, Kowalczyk R, Wójcik JM, Pertoldi C, Kristensen TN, Loeschcke V, Gregersen VR, Bendixen C (2009) Effectiveness of microsatellite and SNP markers for parentage and identity analysis in species with low genetic diversity: the case of European bison.Heredity, 103, 326-332. |
| [96] | Townsend PA, Levey DJ (2005) An experimental test of whether habitat corridors affect pollen transfer.Ecology, 86, 466-475. |
| [97] | Valbuena-Carabaña M, González-Martínez SC, Sork VL, Collada C, Soto A, Goicoechea PG, Gil L (2005) Gene flow and hybridisation in a mixed oak forest (Quercus pyrenaica Willd. and Quercus petraea (Matts.) Liebl.) in central Spain.Heredity, 95, 457-465. |
| [98] | van Rossum F (2009) Pollen dispersal and genetic variation in an early-successional forest herb in a peri-urban forest.Plant Biology, 11, 725-737. |
| [99] | van Rossum F, Stiers I, van Geert A, Triest L, Hardy OJ (2011) Fluorescent dye particles as pollen analogues for measuring pollen dispersal in an insect-pollinated forest herb.Oecologia, 165, 663-674. |
| [100] | Walther-Hellwig K, Frankl R (2000) Foraging distances of Bombus muscorum, Bombus lapidarius, and Bombus terrestris (Hymenoptera, Apidae).Journal of Insect Beh- avior, 13, 239-246. |
| [101] | Wang HF, Sork VL, Wu JG, Ge JP (2010) Effect of patch size and isolation on mating patterns and seed production in an urban population of Chinese pine (Pinus tabulaeformis Carr.).Forest Ecology and Management, 260, 965-974. |
| [102] | Wang KS (2004) Gene flow in European beech (Fagus sylvatica L.).Genetica, 122, 105-113. |
| [103] | Wikelski M, Moxley J, Eaton-Mordas A, López-Uribe MM, Holland R, Moskowitz D, Roubik DW, Kays R (2010) Large-range movements of neotropical orchid bees observed via radio telemetry.PLoS ONE, 5, e10738, doi: 10.1371/journal.pone.0010738. |
| [104] | Zalapa JE, Cuevas H, Zhu HY, Steffan S, Senalik D, Zeldin E, McCown B, Harbut R, Simon P (2012) Using next-generation sequencing approaches to isolate simple sequence repeat (ssr) loci in the plant sciences.American Journal of Botany, 99, 193-208. |
| [105] | Zhu H, Senalik D, McCown BH, Zeldin EL, Speers J, Hyman J, Bassil N, Hummer K, Simon PW, Zalapa JE (2012) Mining and validation of pyrosequenced simple sequence repeats (SSRs) from American cranberry (Vaccinium macrocarpon Ait.).Theoretical and Applied Genetics, 124, 87-96. |
| [1] | Yanwen Lv, Ziyun Wang, Yu Xiao, Zihan He, Chao Wu, Xinsheng Hu. Advances in lineage sorting theories and their detection methods [J]. Biodiv Sci, 2024, 32(4): 23400-. |
| [2] | Chen Feng, Jie Zhang, Hongwen Huang. Parallel situ conservation: A new plant conservation strategy to integrate in situ and ex situ conservation of plants [J]. Biodiv Sci, 2023, 31(9): 23184-. |
| [3] | Qiong Sun, Rong Wang, Xiaoyong Chen. Genomic island of divergence during speciation and its underlying mechanisms [J]. Biodiv Sci, 2022, 30(3): 21383-. |
| [4] | Xiangxiang Chen, Zhongshuai Gai, Juntuan Zhai, Jindong Xu, Peipei Jiao, Zhihua Wu, Zhijun Li. Genetic diversity and construction of core conservation units of the natural populations of Populus euphratica in Northwest China [J]. Biodiv Sci, 2021, 29(12): 1638-1649. |
| [5] | Chao Li,Jinjin Jin,Jinzhen Luo,Chunhui Wang,Junjie Wang,Jun Zhao. Genetic relationships of hatchery populations and wild populations of Tanichthys albonubes near Guangzhou [J]. Biodiv Sci, 2020, 28(4): 474-484. |
| [6] | Yuanyuan Li, Chaonan Liu, Rong Wang, Shuixing Luo, Shouqian Nong, Jingwen Wang, Xiaoyong Chen. Applications of molecular markers in conserving endangered species [J]. Biodiv Sci, 2020, 28(3): 367-375. |
| [7] | Li Meng, Wei Tingting, Shi Boyang, Hao Xiyang, Xu Haigen, Sun Hongying. Biodiversity monitoring of freshwater benthic macroinvertebrates using environmental DNA [J]. Biodiv Sci, 2019, 27(5): 480-490. |
| [8] | Yahong Zhang, Huixia Jia, Zhibin Wang, Pei Sun, Demei Cao, Jianjun Hu. Genetic diversity and population structure of Populus yunnanensis [J]. Biodiv Sci, 2019, 27(4): 355-365. |
| [9] | Jian-Feng Mao, Yongpeng Ma, Renchao Zhou. Approaches used to detect and test hybridization: combining phylogenetic and population genetic analyses [J]. Biodiv Sci, 2017, 25(6): 577-599. |
| [10] | Jie Yang, Jia He, Danbi Wang, En Shi, Wenyu Yang, Qifang Geng, Zhongsheng Wang*. Progress in research and application of InDel markers [J]. Biodiv Sci, 2016, 24(2): 237-243. |
| [11] | Haomin Lyu, Renchao Zhou, Suhua Shi. Recent advances in the study of ecological speciation [J]. Biodiv Sci, 2015, 23(3): 398-407. |
| [12] | Dangni Zhang, Lianming Zheng, Jinru He, Wenjing Zhang, Yuanshao Lin, Yang Li. DNA barcoding of hydromedusae in northern Beibu Gulf for species identification [J]. Biodiv Sci, 2015, 23(1): 50-60. |
| [13] | Zhonghu Li,Zhanlin Liu,Mali Wang,Zengqiang Qian,Peng Zhao,Juan Zhu,Yixin Yang,Xiaohao Yan,Yinjun Li,Guifang Zhao. A review on studies of speciation in the presence of gene flow: evolution of reproductive isolation [J]. Biodiv Sci, 2014, 22(1): 88-96. |
| [14] | Wen Liu, Hong Liang, Biren Yang, Jiewen Lin, Yanji Hu. Short RNAs, potential novel molecular markers for higher plants [J]. Biodiv Sci, 2012, 20(1): 86-93. |
| [15] | Zhaoxia Cui, Huan Zhang, Linsheng Song, Feng You. Genetic diversity of marine animals in China: a summary and prospec- tiveness [J]. Biodiv Sci, 2011, 19(6): 815-833. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2022 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn ![]()
