



Biodiv Sci ›› 2012, Vol. 20 ›› Issue (1): 86-93. DOI: 10.3724/SP.J.1003.2012.08123 cstr: 32101.14.SP.J.1003.2012.08123
• Methodologies • Previous Articles Next Articles
Wen Liu, Hong Liang, Biren Yang, Jiewen Lin, Yanji Hu( )
)
Received:2011-07-22
Accepted:2011-10-16
Online:2012-01-20
Published:2012-02-14
Contact:
Yanji Hu
Wen Liu, Hong Liang, Biren Yang, Jiewen Lin, Yanji Hu. Short RNAs, potential novel molecular markers for higher plants[J]. Biodiv Sci, 2012, 20(1): 86-93.
| 分类阶元 Taxonomic category | 中文名 Chinese name | 拉丁名 Scientific name | |
|---|---|---|---|
| 被子植物门 Angiospermae | |||
| 双子叶植物纲 Dicotyledononeae | 猕猴桃科 Actinidiaceae | 中华猕猴桃品种红阳♂ | A. chinensis ‘Hongyang’♂ |
| 中华猕猴桃品种红阳♀ | A. chinensis ‘Hongyang’♀ | ||
| 中华猕猴桃品种武雄♂ | A. chinensis ‘Wuxiong’♂ | ||
| 中华猕猴桃品种武植3号♀ | A. chinensis ‘Wuzhi No.3’♀ | ||
| 美味猕猴桃品种帮增1号♂ | A. chinensis var. deliciosa‘Bangzeng No.1’♂ | ||
| 美味猕猴桃品种和平1号♀ | A. chinensis var. deliciosa ‘Heping No.1’♀ | ||
| 美味猕猴桃品种米良1号♀ | A. chinensis var. deliciosa ‘Miliang No.1’ ♀ | ||
| 桑科 Moraceae | 大叶榕 | Ficus altissima Blume | |
| 水同木 | Ficus fistulosa Reinw. ex Blume | ||
| 垂叶榕 | Ficus benjamina L. | ||
| 柳叶榕 | Ficus variolosa Lindl. ex Benth. | ||
| 细叶榕 | Ficus microcarpa L. f. | ||
| 桂木 | Artocarpus nitidus subsp.lingnanensis(Merr.) F. M. Jarrett | ||
| 面包树 | Artocarpus communis J. R. Forst. et G. Forst. | ||
| 树菠萝 | Artocarpus heterophyllus Lam. | ||
| 豆科 Leguminosae | 红花羊蹄甲 | Bauhinia blakeanaDunn | |
| 酸豆 | Tamarindus indica L. | ||
| 紫花苜蓿 | Medicago sativa L. | ||
| 花生 | Arachis hypogaeaL. | ||
| 柱花草 | Stylosanthes scabra Vogel. | ||
| 黄槐 | Cassia surattensis Burm. f. | ||
| 红绒球 | Calliandra haematocephala Hassk. | ||
| 绿豆 | Vigna radiata (L.) R. Wilczek | ||
| 夹竹桃科 Apocynaceae | 夹竹桃 | Nerium oleander L. | |
| 鸡蛋花 | Plumeria rubraL. | ||
| 狗牙花 | Tabernaemontana divaricata (L.)R. Br. ex Roem. et Schult. | ||
| 长春花 | Catharanthus roseus (L.) G. Don | ||
| 沙漠玫瑰 | Adenium obesum (Forsk.) Roem. & Schult. | ||
| 木犀科 Oleaceae | 茉莉 | Jasminum sambac (L.) Aiton | |
| 樟科 Lauraceae | 樟树 | Cinnamomum camphora (L.) J. Presl. | |
| 单子叶植物纲Monocotyledoneae | 禾本科 Poaceae | 玉米 | Zea mays L. |
| 香根草 | Chrysopogon zizanioides (L.) Roberty | ||
| 狗尾草 | Setaria viridis (L.) P. Beauv. | ||
| 小麦 | Triticum aestivum L. | ||
| 水稻 | Oryza sativa L. | ||
| 佛肚竹 | Bambusa ventricosa McClure | ||
| 棕榈科 Arecaceae | 鱼尾葵 | Caryota ochlandra Hance | |
| 天南星科 Araceae | 合果芋 | Syngonium podophyllumSchott. | |
| 鸭跖草科 Commeliaceae | 鸭跖草 | Commelina communisL. | |
| 裸子植物门 Gymnospermae | 银杏科 Ginkgoaceae | 银杏 | Ginkgo bilobaL. |
| 苏铁科 Cycadaceae | 华南苏铁 | Cycas rumphiiMiq. | |
| 罗汉松科 Podocarpaceae | 罗汉松 | Podocarpus macrophyllus (Thunb.) Sweet | |
| 松科 Pinaceae | 马尾松 | Pinus massonianaD. Don | |
| 南洋杉科 Araucariaceae | 南洋杉 | Araucaria cunninghamii Aiton ex D. Don | |
| 杉科 Taxodiaceae | 落羽杉 | Taxodium distichum (L.) Rich. | |
| 柏科 Cupressaceae | 龙柏 | Sabina chinensis(L.) Antoine ‘Kaizuca’ | |
| 蕨类植物门 Pteridophyta | 海金沙科 Lygodiaceae | 海金沙 | Lygodium japonicum (Thunb.) Sw. |
| 凤尾蕨科 Pteridaceae | 井栏边草 | Pteris multifidaPoir. | |
| 铁角蕨科 Aspleniaceae | 鸟巢蕨 | Neottopteris nidus (L.) J. Sm. | |
| 鹿角蕨科 Polypodiaceae | 鹿角蕨 | Platycerium wallichiiHook. | |
| 肾蕨科 Davalliaceae | 波士顿肾蕨 | Nephrolepis exaltata (L.) Schott ‘Bostoniensis’ | |
Table 1 Plants used in the present study
| 分类阶元 Taxonomic category | 中文名 Chinese name | 拉丁名 Scientific name | |
|---|---|---|---|
| 被子植物门 Angiospermae | |||
| 双子叶植物纲 Dicotyledononeae | 猕猴桃科 Actinidiaceae | 中华猕猴桃品种红阳♂ | A. chinensis ‘Hongyang’♂ |
| 中华猕猴桃品种红阳♀ | A. chinensis ‘Hongyang’♀ | ||
| 中华猕猴桃品种武雄♂ | A. chinensis ‘Wuxiong’♂ | ||
| 中华猕猴桃品种武植3号♀ | A. chinensis ‘Wuzhi No.3’♀ | ||
| 美味猕猴桃品种帮增1号♂ | A. chinensis var. deliciosa‘Bangzeng No.1’♂ | ||
| 美味猕猴桃品种和平1号♀ | A. chinensis var. deliciosa ‘Heping No.1’♀ | ||
| 美味猕猴桃品种米良1号♀ | A. chinensis var. deliciosa ‘Miliang No.1’ ♀ | ||
| 桑科 Moraceae | 大叶榕 | Ficus altissima Blume | |
| 水同木 | Ficus fistulosa Reinw. ex Blume | ||
| 垂叶榕 | Ficus benjamina L. | ||
| 柳叶榕 | Ficus variolosa Lindl. ex Benth. | ||
| 细叶榕 | Ficus microcarpa L. f. | ||
| 桂木 | Artocarpus nitidus subsp.lingnanensis(Merr.) F. M. Jarrett | ||
| 面包树 | Artocarpus communis J. R. Forst. et G. Forst. | ||
| 树菠萝 | Artocarpus heterophyllus Lam. | ||
| 豆科 Leguminosae | 红花羊蹄甲 | Bauhinia blakeanaDunn | |
| 酸豆 | Tamarindus indica L. | ||
| 紫花苜蓿 | Medicago sativa L. | ||
| 花生 | Arachis hypogaeaL. | ||
| 柱花草 | Stylosanthes scabra Vogel. | ||
| 黄槐 | Cassia surattensis Burm. f. | ||
| 红绒球 | Calliandra haematocephala Hassk. | ||
| 绿豆 | Vigna radiata (L.) R. Wilczek | ||
| 夹竹桃科 Apocynaceae | 夹竹桃 | Nerium oleander L. | |
| 鸡蛋花 | Plumeria rubraL. | ||
| 狗牙花 | Tabernaemontana divaricata (L.)R. Br. ex Roem. et Schult. | ||
| 长春花 | Catharanthus roseus (L.) G. Don | ||
| 沙漠玫瑰 | Adenium obesum (Forsk.) Roem. & Schult. | ||
| 木犀科 Oleaceae | 茉莉 | Jasminum sambac (L.) Aiton | |
| 樟科 Lauraceae | 樟树 | Cinnamomum camphora (L.) J. Presl. | |
| 单子叶植物纲Monocotyledoneae | 禾本科 Poaceae | 玉米 | Zea mays L. |
| 香根草 | Chrysopogon zizanioides (L.) Roberty | ||
| 狗尾草 | Setaria viridis (L.) P. Beauv. | ||
| 小麦 | Triticum aestivum L. | ||
| 水稻 | Oryza sativa L. | ||
| 佛肚竹 | Bambusa ventricosa McClure | ||
| 棕榈科 Arecaceae | 鱼尾葵 | Caryota ochlandra Hance | |
| 天南星科 Araceae | 合果芋 | Syngonium podophyllumSchott. | |
| 鸭跖草科 Commeliaceae | 鸭跖草 | Commelina communisL. | |
| 裸子植物门 Gymnospermae | 银杏科 Ginkgoaceae | 银杏 | Ginkgo bilobaL. |
| 苏铁科 Cycadaceae | 华南苏铁 | Cycas rumphiiMiq. | |
| 罗汉松科 Podocarpaceae | 罗汉松 | Podocarpus macrophyllus (Thunb.) Sweet | |
| 松科 Pinaceae | 马尾松 | Pinus massonianaD. Don | |
| 南洋杉科 Araucariaceae | 南洋杉 | Araucaria cunninghamii Aiton ex D. Don | |
| 杉科 Taxodiaceae | 落羽杉 | Taxodium distichum (L.) Rich. | |
| 柏科 Cupressaceae | 龙柏 | Sabina chinensis(L.) Antoine ‘Kaizuca’ | |
| 蕨类植物门 Pteridophyta | 海金沙科 Lygodiaceae | 海金沙 | Lygodium japonicum (Thunb.) Sw. |
| 凤尾蕨科 Pteridaceae | 井栏边草 | Pteris multifidaPoir. | |
| 铁角蕨科 Aspleniaceae | 鸟巢蕨 | Neottopteris nidus (L.) J. Sm. | |
| 鹿角蕨科 Polypodiaceae | 鹿角蕨 | Platycerium wallichiiHook. | |
| 肾蕨科 Davalliaceae | 波士顿肾蕨 | Nephrolepis exaltata (L.) Schott ‘Bostoniensis’ | |
| 分类阶元 Taxonomic category | 材料数量 No. of species | 同源片段数目 No. of band analogs | 多态性位点数 No. of polymorphic loci | 总位点数 No. of total loci | 多态性百分比 PPB(%) | 多态性信息量 PIC |
|---|---|---|---|---|---|---|
| 禾本科 Poaceae | 6 | 5-6 | 9 | 10 | 90 | 0.859 |
| 桑科 Moraceae | 8 | 5 | 4 | 7 | 57.1 | 0.791 |
| 夹竹桃科 Apocynaceae | 5 | 5 | 8 | 9 | 88.9 | 0.840 |
| 豆科 Leguminosae | 7 | 5 | 17 | 17 | 100 | 0.926 |
| 蕨类植物门 Pteridophyta | 5 | 3-4 | 12 | 12 | 100 | 0.889 |
| 裸子植物门 Gymnospermae | 7 | 4-6 | 17 | 17 | 100 | 0.917 |
Table 2 The short RNA segments polymorphism analysis in higher plants
| 分类阶元 Taxonomic category | 材料数量 No. of species | 同源片段数目 No. of band analogs | 多态性位点数 No. of polymorphic loci | 总位点数 No. of total loci | 多态性百分比 PPB(%) | 多态性信息量 PIC |
|---|---|---|---|---|---|---|
| 禾本科 Poaceae | 6 | 5-6 | 9 | 10 | 90 | 0.859 |
| 桑科 Moraceae | 8 | 5 | 4 | 7 | 57.1 | 0.791 |
| 夹竹桃科 Apocynaceae | 5 | 5 | 8 | 9 | 88.9 | 0.840 |
| 豆科 Leguminosae | 7 | 5 | 17 | 17 | 100 | 0.926 |
| 蕨类植物门 Pteridophyta | 5 | 3-4 | 12 | 12 | 100 | 0.889 |
| 裸子植物门 Gymnospermae | 7 | 4-6 | 17 | 17 | 100 | 0.917 |
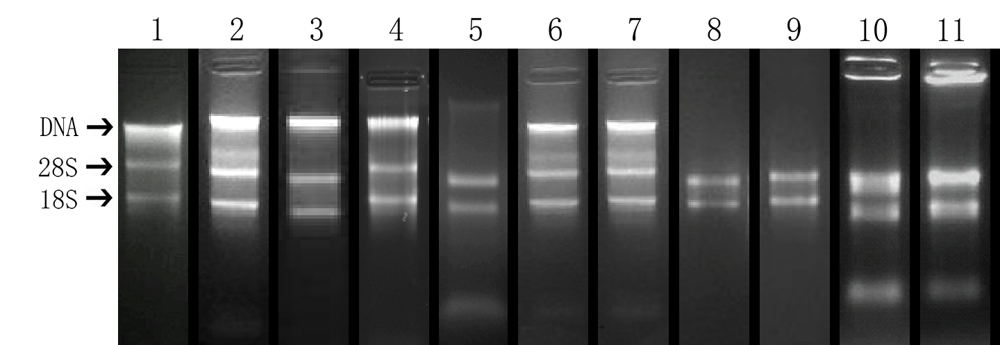
Fig. 1 The electrophoretogram of the total nucleic acid and RNA from certain plants. 1, Caryota ochlandra; 2, Ficus microcarpa; 3, F. benjamina; 4, Artocarpus lingnanensis; 5, Nerium oleander; 6, Total nucleic acid from Actinidia chinensis(♀, leaf); 7, Total nucleic acid from Actinidia chinensis(♂, leaf); 8, Total RNA from Actinidia chinensis(♀, bud); 9, Total RNA from Actinidia chinensis(♂, bud); 10, Total RNA from Ginkgo biloba; 11, Total RNA from Pteris multifida.
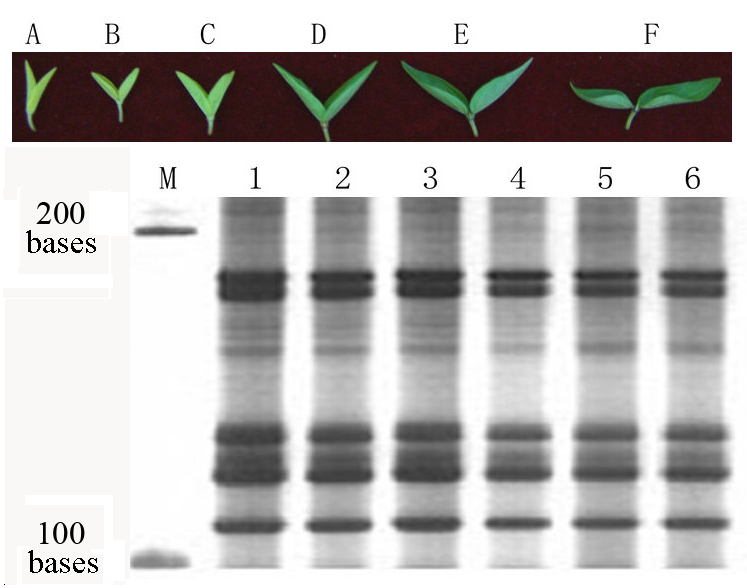
Fig. 2 The influence of illumination on the leaf of mung bean and short RNA segments. A to F show the leaves of mung bean after 0, 1, 2, 4, 6, 8 hours illumination each day, respectively (for seven days, 2,000 lx, 25℃); 1-6 show the short RNA segments after 0, 1, 2, 4, 6, 8 hours illumination each day for seven days. M is the single strand RNA marker.
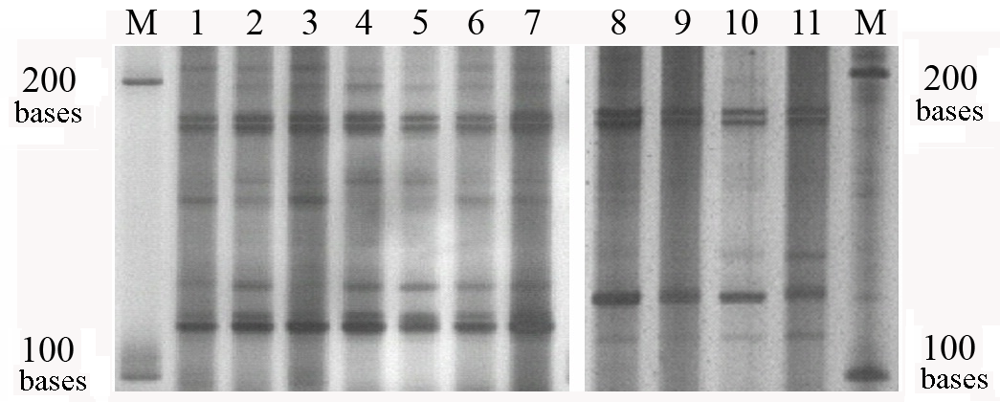
Fig. 3 The electrophoresis result of short RNA segments from different kiwifruit varieties and organs. 1-7 areActinidia chinensis ‘Hongyang’ (♂), A. chinensis ‘Wuxiong’ (♂), A. chinensis var. deliciosa ‘Bangzeng No.1’ (♂), A. chinensis ‘Hongyang’ (♀), A. chinensis ‘Wuzhi No. 3’♀, A. chinensisvar. deliciosa ‘Miliang No.1’ (♀), A. chinensisvar. deliciosa ‘Heping No.1’ (♀), respectively. The total RNA samples which had been hold in 70℃for 10 minutes. 8-11 are A. chinensis ‘Wuzhi No. 3’ (♀) bud, A. chinensis ‘Wuxiong’ (♂) bud, A. chinensis ‘Wuxiong’ (♂) leaf, A. chinensis ‘Wuzhi No. 3’ (♀) leaf, respectively. The total RNA samples which had 1-fold dilution and had not been hold in 70 ℃. M is the single strand RNA marker.
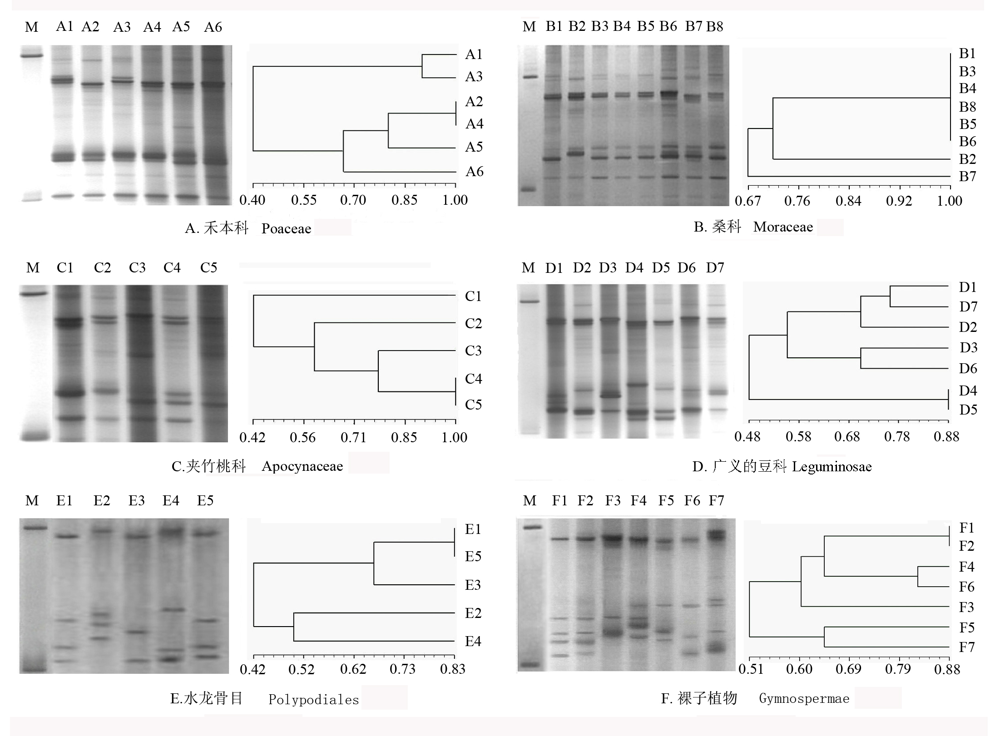
Fig. 4 The pattern and dendrogram of short RNA segments from higher plants. A1, Zea mays; A2, Setaria viridis; A3, Chrysopogon zizanioides; A4, Triticum aestivum; A5, Oryza sativa; A6, Bambusa ventricosa; B1, Ficus altissima; B2, Ficus harlandii; B3, Ficus benjamina; B4, Ficus variolosa; B5, Ficus microcarpa; B6, Artocarpus lingnanensis; B7, Artocarpus altilis; B8, Artocarpus heterophyllus, C1, Nerium oleander; C2, Plumeria rubra; C3, Ervatamia divaricata; C4, Catharanthus roseus; C5, Adenium obesum, D1, Bauhinia blakeana; D2, Tamarindus indica; D3, Medicago sativa; D4, Fabales, Arachis hypogaea; D5, Stylosanthes scabra; D6, Cassia surattensis; D7, Calliandra haematocephala; E1, Lygodium japonicum; E2, Platycerium wallichii; E3, Neottopteris nidus; E4, Nephrolepis exaltata‘Bostoniensis’; E5,Pteris multifida, F1, Sabina chinensis ‘Kaizuca’; F2, Pinus massomana; F3, Podocarpus macrophyllus; F4, Ginkgo biloba; F5, Cycas rumphii; F6, Taxodium distichum; F7, Araucaria cunninghamii. M is the single strand RNA marker; from up to down are 200 bases and 100 bases separately.
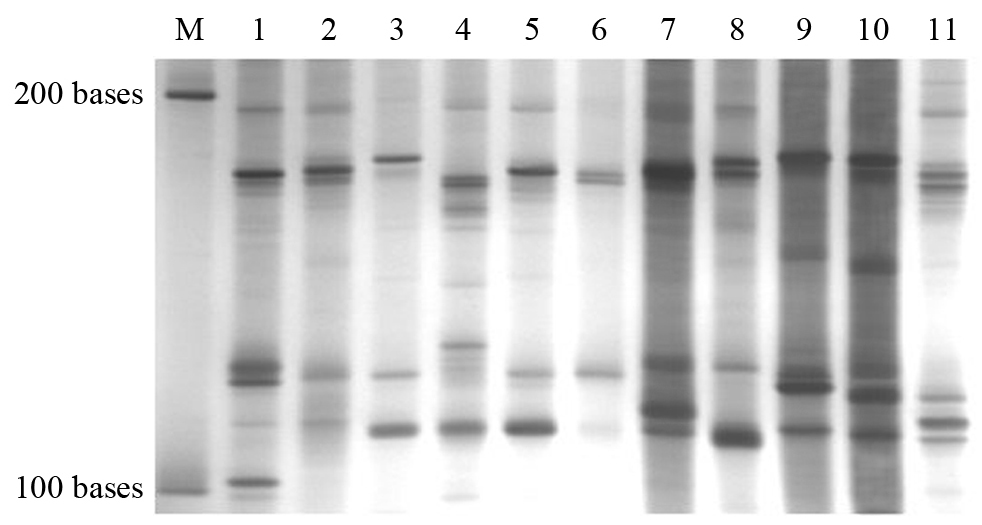
Fig. 5 The pattern of short RNA segments from 11 species from Angiosperm. 1-11 areSetaria viridis,Canna indica, Allium sativum, Caryota ochlandra, Syngonium podophyllum, Commelina communis, Jasminum sambac, Actinidia chinensis, Medicago sativa, Ervatamia divaricata, and Cinnamomum camphora, respectively. M is the single strand RNA marker.
| [1] | Ching RC (1954) A systematic arrangement of the Chinese fern families and genera with corresponding names in Chinese. Acta Phytotaxonomica Sinica (植物分类学报), 3,93-99. |
| [2] | Ching RC (1978) The Chinese fern families and genera: systematic arrangement and historical origin. Acta Phytotaxonomica Sinica (植物分类学报), 16,1-37. |
| [3] | Chun J (淳俊), Zheng YF (郑彦峰), Wang SH (王胜华), Chen F (陈放) (2008) A RNA isolation method suitable for a wide range of materials. Progress in Biochemistry and Biophysics (生物化学与生物物理进展), 35,591-598. (in Chinese with English abstract) |
| [4] | Fan LJ (樊龙江), Guo XY (郭兴益), Ma NX (马乃训) (2006) Comparative study on phylogenetics and sequences composition of bamboos and cereals. Forest Research (林业科学研究), 19,165-169. (in Chinese with English abstract) |
| [5] | Frankham R, Ballou JD, Briscoe A (translated by Huang HW (黄宏文), Kang M (康明)) (2005) Introduction to Conservation Genetics (保育遗传学导论). Science Press, Beijing. (in Chinese) |
| [6] | Guo BL (郭宝林), Hong DY (洪德元), Xiao PG (肖培根) (2008) Further research on chemotaxonomy of paeonol and analogs in Paeonia. Journal of Systematics and Evolution (植物分类学报), 46,724-729. (in Chinese with English abstract) |
| [7] | Guo XH (郭新弧) (2010) A new tentative idea on the classification system of Pteridophyta. Journal of Anhui Normal University(Natural Science) (安徽师范大学学报(自然科学版)), 33,52-56. (in Chinese with English abstract) |
| [8] | Huang HW (黄宏文) (2009) History of 100 years of domestication and improvement of kiwifruit and gene discovery from genetic introgressed populations in the wild. Chinese Bulletin of Botany (植物学报), 44,127-142. (in Chinese with English abstract) |
| [9] | Huang HW (黄宏文), Gong JJ (龚俊杰), Wang SM (王圣梅), He ZC (何子灿), Zhang ZH (张忠慧), Li JQ (李建强) (2000) Genetic diversity in the genus Actinidia. Biodiversity Science (生物多样性), (8)1-12. (in Chinese with English abstract) |
| [10] | Huang JX (黄久香), Zhang SZ (张寿洲), Zhuang XY (庄雪影), Li BT (李秉滔) (2007) RAPD analysis of twelve species of Apocynaceae. Journal of Fujian Forestry Science and Technology (福建林业科技), 34,14-18. (in Chinese with English abstract) |
| [11] | Jiang Y (蒋英) (1977) Apocynaceae. In: Flora Reipublicae Popularis Sinicae, Tomus 63 (中国植物志(第63卷)) (ed. DelectisFlorae Reipublicae Popularis Sinicae Agendae Academicae Sinicae Edita (中国科学院中国植物志编辑委员会), pp.63-246. Science Press, Beijing. (in Chinese) |
| [12] | Jiao MC (焦明超), Zhao DX (赵大显), Ouyang S (欧阳珊), Wu XP (吴小平) (2011) Application prospect of DNA barcode technology in taxonomy. Hubei Agricultural Sciences (湖北农业科学), 50,886-890. (in Chinese with English abstract) |
| [13] | Li H (李红), He P (何平) (1996) Numerical evaluation of morphology of intergeneric relationships of Apocynaceae in China. Journal of Southwest China Normal University (Natural Science) (西南师范大学学报(自然科学版)), 21,590-599. (in Chinese with English abstract) |
| [14] | Liu W (刘文), Ke HP (柯辉鹏), Liang H (梁红) (2008) A broad-suitable method for total nucleic acid extraction from plants and animals. Journal of Zhongkai University of Agriculture and Technology (仲恺农业技术学院学报), 21,17-21. (in Chinese with English abstract) |
| [15] | Mo BH (莫帮辉), Qu L (屈莉), Han S (韩松), He JW (何建伟), Zhao M (赵明), Zeng XM (曾晓茂) (2008) DNA Barcoding identification. I. Research progress and applied perspective of DNA barcoding. Sichuan Journal of Zoology (四川动物), 27,303-306. (in Chinese with English abstract) |
| [16] | Sen L (森林), Su YJ (苏应娟), Zhang B (张冰), Wang T (王艇) (2010) Adaptive evolution of the rbcl gene in Pteridaceous ferns. Journal of Tropical and Subtropical Botany (热带亚热带植物学报), 18,1-8. (in Chinese with English abstract) |
| [17] | Wang LX (王丽侠), Cheng XZ (程须珍), Wang SH (王素华) (2011) Genetic diversity in adzuki bean and its relatives based on SSR markers. Biodiversity Science (生物多样性), 19,17-23. (in Chinese with English abstract) |
| [18] | Wang RX (王任翔), Deng XC (邓晰朝), Li JR (李洁荣), Deng JZ (邓建珍), Lu SG (陆树刚) (2009) Leaf micromorphology of 12 species of Polypodiaceae from Guangxi of China and its taxonomic significance. Journal of Guangxi Normal University (Natural Science) (广西师范大学学报(自然科学版)), 27,104-108. (in Chinese with English abstract) |
| [19] | Wang RX (王任翔), Lu SG (陆树刚) (2010) Leaf epidermis micromorphology of subfam. Polypodioideae Nayar (Polypodiaceae) and its taxonomic significance. Journal of Wuhan Botanical Research (武汉植物学研究), 28,410-416. (in Chinese with English abstract) |
| [20] | Wei XQ (魏先芹), Li QQ (李琴琴), He XJ (何兴金), Gao YD (高云东), Zhao LH (赵丽华) (2011) A cytotaxonomic study on 21 populations of 13 Allium species. Plant Science Journal (植物科学学报), 29,18-30. (in Chinese with English abstract) |
| [21] | Zeng H (曾华), Li DW (李大卫), Huang HW (黄宏文) (2009) Distribution pattern of ploidy variation of Actinidia chinensis and A.deliciosa. Journal of Wuhan Botanical Research (武汉植物学研究), 27,312-317. (in Chinese with English abstract) |
| [22] | Zhang BB (张碧波), Wang RX (王任翔), Chang YF (常艳芬), Lu SG (陆树刚) (2006) Studies on the spore morphology of Polypodiodes Ching (Polypodiaceae) from southwest China. Journal of Wuhan Botanical Research (武汉植物学研究), 24,113-118. (in Chinese with English abstract) |
| [23] | Zhang HD (张宏达), Huang YH (黄云晖), Miao RH (缪汝槐), Ye CX (叶创兴), Liao WB (廖文波), Jin JH (金建华) (2006) Systematics of Spermatophyta (种子植物系统学). Science Press, Beijing. (in Chinese) |
| [24] | Zhang LJ (张丽君), Chen J (陈洁), Wang T (王艇) (2010) Adaptive evolution in the chloroplast gene rps4 in ferns. Bulletin of Botanical Research (植物研究), 30,42-50. (in Chinese with English abstract) |
| [25] | Zhong FL (钟凤林), Guo ZX (郭志雄), Ma CY (马传营), Pan DM (潘东明) (2010) The method of total RNA isolation from Pummelo juice Sac. Biotechnology Bulletin (生物技术通报), (02),109-112. (in Chinese with English abstract) |
| [26] | Zhou Y (周媛), Gao S (高磊), Wang ZW (汪志伟), Wang T (王艇) (2009) Application of molecular marker techniques in genetic diversity of Pteridophytes. Journal of Wuhan Botanical Research (武汉植物学研究), 27,667-673. (in Chinese with English abstract) |
| [27] | Zhu XY (朱相云), Du YF (杜玉芬) (2002) Exotic Legume species in China. Bulletin of Botanical Research (植物研究), 22,139-150. (in Chinese with English abstract) |
| [1] | Yuanyuan Li, Chaonan Liu, Rong Wang, Shuixing Luo, Shouqian Nong, Jingwen Wang, Xiaoyong Chen. Applications of molecular markers in conserving endangered species [J]. Biodiv Sci, 2020, 28(3): 367-375. |
| [2] | Shufang Liu, Jinxian Liu, Zhimeng Zhuang, Tianxiang Gao, Zhiqiang Han, Dagang Chen. Monophyletic origin and synonymic phenomena in the sub-family Cynoglossinae inferred from mitochondrial DNA sequences [J]. Biodiv Sci, 2010, 18(3): 275-282. |
| [3] | QIU Fang, FU Jian-Min, JIN De-Min, WANG Bin. The molecular detection of genetic diversity [J]. Biodiv Sci, 1998, 06(2): 143-150. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn