



Biodiv Sci ›› 2022, Vol. 30 ›› Issue (6): 21545. DOI: 10.17520/biods.2021545 cstr: 32101.14.biods.2021545
• Original Papers: Genetic Diversity • Previous Articles Next Articles
Yongguang Li1,2, Hui Ren1, Yingjie Zhang1, Ruining Li2, Hao Ai2, Xianzhong Huang2,3,*( )
)
Received:2021-12-30
Accepted:2022-03-10
Online:2022-06-20
Published:2022-03-11
Contact:
Xianzhong Huang
Yongguang Li, Hui Ren, Yingjie Zhang, Ruining Li, Hao Ai, Xianzhong Huang. Analysis of the molecular evolution of the PEBP gene family in cruciferous plants[J]. Biodiv Sci, 2022, 30(6): 21545.
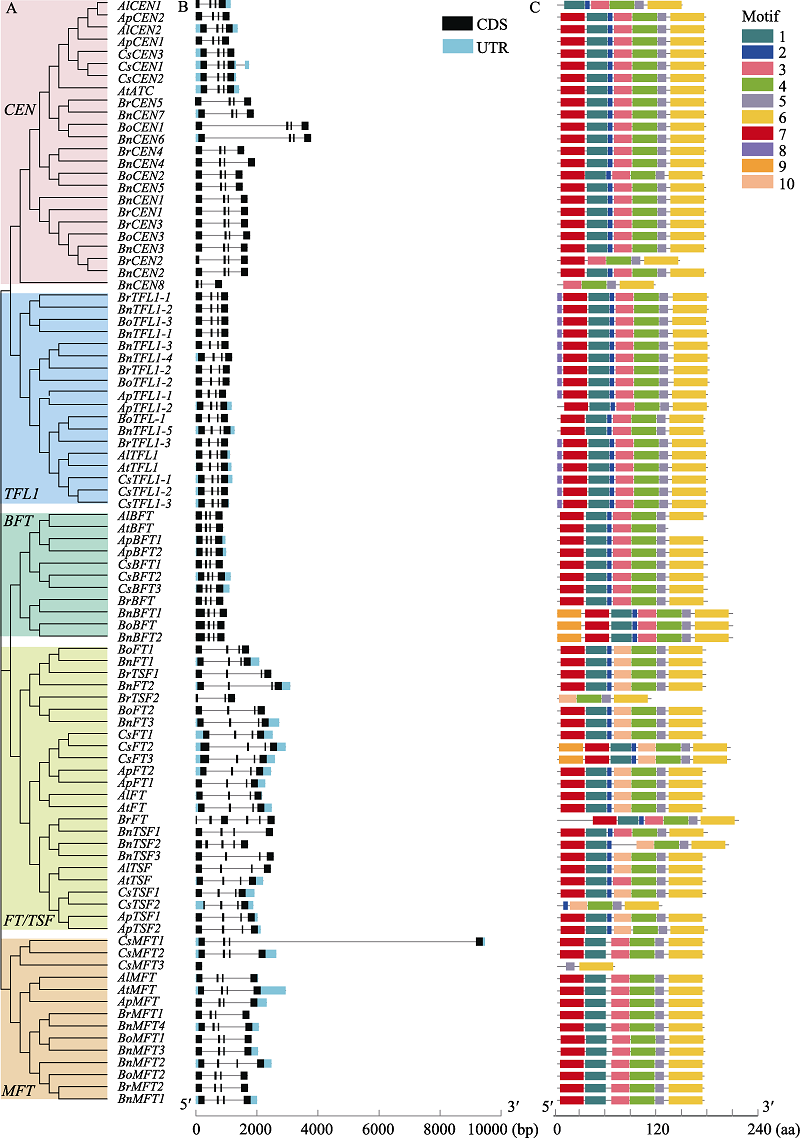
Fig. 1 Motif distributions and exon-intron structures of the PEBP family members in Arabidopsis thaliana, A. lyrata, A. pumila, Camelina sativa, Brassica oleracea, B. rapa, and B. napus. A, PEBP phylogenetic tree of seven species; B, Exon-intron distribution of PEBP genes; C, Distribution characteristics of the conserved motifs of PEBP proteins; CDS, Coding sequence; UTR, Untranslated region; aa, Amino acid.
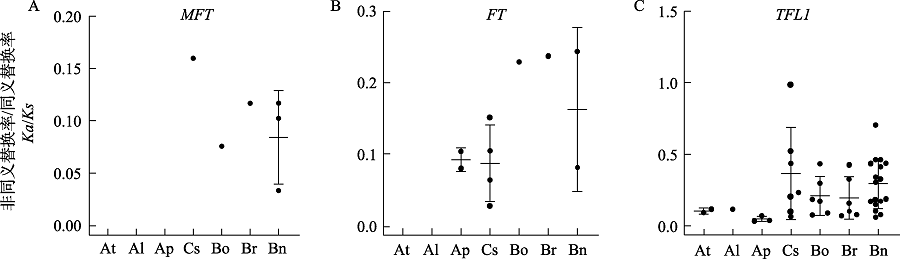
Fig. 2 Statistics of selection pressures on the PEBP collinearity genes within species. At, Arabidopsis thaliana; Al, A. lyrata; Ap, A. pumila; Cs, Camelina sativa; Bo, Brassica oleracea; Br, B. rapa; Bn, B. napus.
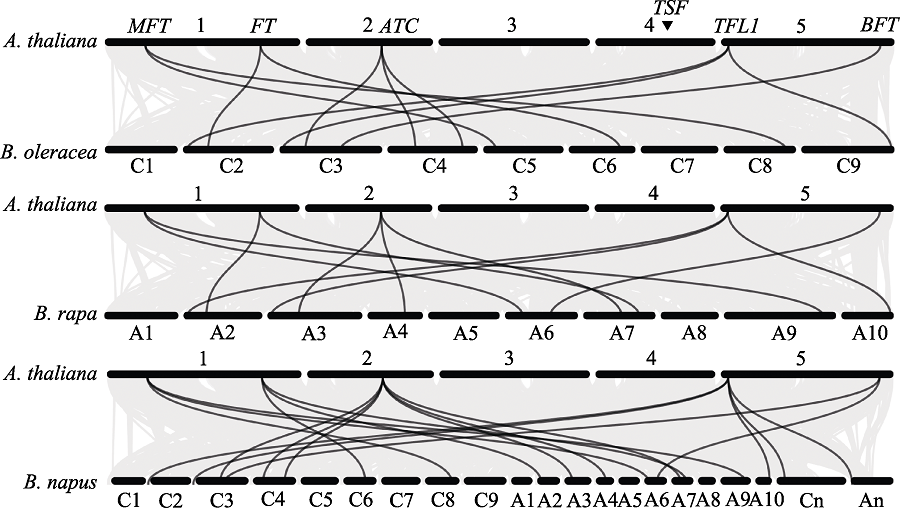
Fig. 3 Syntenic relationships of PEBP genes in Arabidopsis thaliana, Brassica oleracea, B. rapa, and B. napus. The cylinder represents 5, 9, 10 and 21 chromosomes of A. thaliana, B. oleracea, B. rapa, and B. napus, respectively. The black line represents a common linear relationship of the PEBP gene, and the gray line represents a genomic linear relationship. The triangle represents the location of the TSF gene.
| 物种 Species | PEBP基因数量 No. of PEBP gene | 不同复制类型PEBP基因数量(百分比) No. of PEBP genes from different origins (percentage) | ||||
|---|---|---|---|---|---|---|
| 单例型 Singleton | 分散型 Dispersed | 近端型 Proximal | 串联重复 Tandem | 全基因组复制或片段复制 WGD/S | ||
| 拟南芥 Arabidopsis thaliana | 6 | 0 | 1 (16.6) | 0 | 0 | 5 (83.3) |
| 琴叶拟南芥 Arabidopsis lyrata | 7 | 0 | 7 (100) | 0 | 0 | 0 |
| 小鼠耳芥 Arabidopsis pumila | 11 | 0 | 1 (9.1) | 0 | 0 | 10 (90.9) |
| 亚麻芥 Camelina sativa | 17 | 0 | 1 (5.9) | 0 | 0 | 16 (94.1) |
| 甘蓝 Brassica oleracea | 11 | 0 | 1 (9.1) | 0 | 0 | 10 (90.9) |
| 白菜 Brassica rapa | 14 | 0 | 3 (21.4) | 1 (7.1) | 0 | 10 (71.4) |
| 油菜 Brassica napus | 25 | 0 | 6 (24) | 0 | 0 | 19 (76) |
Table 1 Types of replication of PEBP gene family in Cruciferae plants
| 物种 Species | PEBP基因数量 No. of PEBP gene | 不同复制类型PEBP基因数量(百分比) No. of PEBP genes from different origins (percentage) | ||||
|---|---|---|---|---|---|---|
| 单例型 Singleton | 分散型 Dispersed | 近端型 Proximal | 串联重复 Tandem | 全基因组复制或片段复制 WGD/S | ||
| 拟南芥 Arabidopsis thaliana | 6 | 0 | 1 (16.6) | 0 | 0 | 5 (83.3) |
| 琴叶拟南芥 Arabidopsis lyrata | 7 | 0 | 7 (100) | 0 | 0 | 0 |
| 小鼠耳芥 Arabidopsis pumila | 11 | 0 | 1 (9.1) | 0 | 0 | 10 (90.9) |
| 亚麻芥 Camelina sativa | 17 | 0 | 1 (5.9) | 0 | 0 | 16 (94.1) |
| 甘蓝 Brassica oleracea | 11 | 0 | 1 (9.1) | 0 | 0 | 10 (90.9) |
| 白菜 Brassica rapa | 14 | 0 | 3 (21.4) | 1 (7.1) | 0 | 10 (71.4) |
| 油菜 Brassica napus | 25 | 0 | 6 (24) | 0 | 0 | 19 (76) |
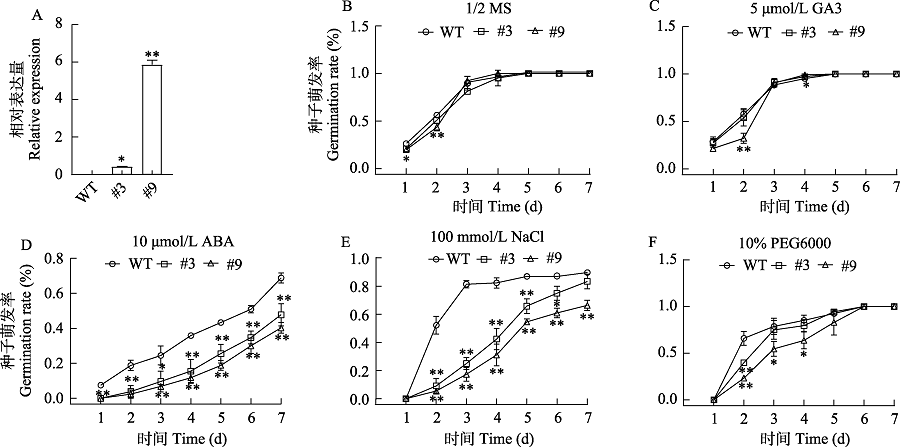
Fig. 4 The germination rate of 35S:GhMFT1 transgenic Arabidopsis thaliana seeds treated with GA, ABA, NaCl and PEG6000. WT, Wild type; #3, #9: 35S:ApMFT transgenic lines, * Significant differences at P < 0.05; ** Significant differences at P < 0.01.
| [1] |
Abe M, Kobayashi Y, Yamamoto S, Daimon Y, Yamaguchi A, Ikeda Y, Ichinoki H, Notaguchi M, Goto K, Araki T (2005) FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science, 309, 1052-1056.
DOI URL |
| [2] |
Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature, 408, 796-815.
DOI URL |
| [3] |
Banfield MJ, Barker JJ, Perry AC, Brady RL (1998) Function from structure? The crystal structure of human phosphatidylethanolamine-binding protein suggests a role in membrane signal transduction. Structure, 6, 1245-1254.
PMID |
| [4] |
Bi ZH, Li X, Huang HS, Hua YW (2016) Identification, functional study, and promoter analysis of HbMFT1, a homolog of MFT from rubber tree (Hevea brasiliensis). International Journal of Molecular Sciences, 17, 247.
DOI URL |
| [5] |
Blanc G, Hokamp K, Wolfe KH (2003) A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome. Genome Research, 13, 137-144.
DOI URL |
| [6] |
Bradley D, Carpenter R, Copsey L, Vincent C, Rothstein S, Coen E (1996) Control of inflorescence architecture in Antirrhinum. Nature, 379, 791-797.
DOI URL |
| [7] |
Chalhoub B, Denoeud F, Liu SY, Parkin IAP, Tang HB, Wang XY, Chiquet J, Belcram H, Tong CB, Samans B, Corréa M, Da Silva C, Just J, Falentin C, Koh CS, Le Clainche I, Bernard M, Bento P, Noel B, Labadie K, Alberti A, Charles M, Arnaud D, Guo H, Daviaud C, Alamery S, Jabbari K, Zhao M, Edger PP, Chelaifa H, Tack D, Lassalle G, Mestiri I, Schnel N, Le Paslier MC, Fan GY, Renault V, Bayer PE, Golicz AA, Manoli S, Lee TH, Thi VHD, Chalabi S, Hu Q, Fan CC, Tollenaere R, Lu YH, Battail C, Shen JX, Sidebottom CHD, Wang XF, Canaguier A, Chauveau A, Bérard A, Deniot G, Guan M, Liu ZS, Sun FM, Lim YP, Lyons E, Town CD, Bancroft I, Wang XW, Meng JL, Ma JX, Pires JC, King GJ, Brunel D, Delourme R, Renard M, Aury JM, Adams KL, Batley J, Snowdon RJ, Tost J, Edwards D, Zhou YM, Hua W, Sharpe AG, Paterson AH, Guan CY, Wincker P (2014) Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science, 345, 950-953.
DOI PMID |
| [8] |
Chen CJ, Chen H, Zhang Y, Thomas HR, Frank MH, He YH, Xia R (2020) TBtools: An integrative toolkit developed for interactive analyses of big biological data. Molecular Plant, 13, 1194-1202.
DOI URL |
| [9] |
Cheng SF, van den Bergh E, Zeng P, Zhong X, Xu JJ, Liu X, Hofberger J, de Bruijn S, Bhide AS, Kuelahoglu C, Bian C, Chen J, Fan GY, Kaufmann K, Hall JC, Becker A, Bräutigam A, Weber APM, Shi CC, Zheng ZJ, Li WJ, Lv MJ, Tao YM, Wang JY, Zou HF, Quan ZW, Hibberd JM, Zhang GY, Zhu XG, Xu X, Schranz ME (2013) The Tarenaya hassleriana genome provides insight into reproductive trait and genome evolution of crucifers. Plant Cell, 25, 2813-2830.
DOI URL |
| [10] |
Clough SJ, Bent AF (1998) Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. The Plant Journal, 16, 735-743.
DOI URL |
| [11] |
Conti L, Bradley D (2007) TERMINAL FLOWER1 is a mobile signal controlling Arabidopsis architecture. Plant Cell, 19, 767-778.
DOI URL |
| [12] |
Edger PP, Pires JC (2009) Gene and genome duplications: The impact of dosage-sensitivity on the fate of nuclear genes. Chromosome Research, 17, 699-717.
DOI URL |
| [13] |
Eshed Y, Lippman ZB (2019) Revolutions in agriculture chart a course for targeted breeding of old and new crops. Science, 366, eaax0025.
DOI URL |
| [14] |
Flagel LE, Wendel JF (2009) Gene duplication and evolutionary novelty in plants. New Phytologist, 183, 557-564.
DOI PMID |
| [15] |
Hedman H, Källman T, Lagercrantz U (2009) Early evolution of the MFT-like gene family in plants. Plant Molecular Biology, 70, 359-369.
DOI URL |
| [16] |
Hu TT, Pattyn P, Bakker EG, Cao J, Cheng JF, Clark RM, Fahlgren N, Fawcett JA, Grimwood J, Gundlach H, Haberer G, Hollister JD, Ossowski S, Ottilar RP, Salamov AA, Schneeberger K, Spannagl M, Wang X, Yang L, Nasrallah ME, Bergelson J, Carrington JC, Gaut BS, Schmutz J, Mayer KFX, van de Peer Y, Grigoriev IV, Nordborg M, Weigel D, Guo YL (2011) The Arabidopsis lyrata genome sequence and the basis of rapid genome size change. Nature Genetics, 43, 476-481.
DOI URL |
| [17] |
Huang XZ, Yang LF, Jin YH, Lin J, Liu F (2017) Generation, annotation, and analysis of a large-scale expressed sequence tag library from Arabidopsis pumila to explore salt-responsive genes. Frontiers in Plant Science, 8, 955.
DOI URL |
| [18] | Jin S, Nasim Z, Susila H, Ahn JH (2021) Evolution and functional diversification of FLOWERING LOCUS T/TERMINAL FLOWER 1 family genes in plants. Seminars in Cell & Developmental Biology, 109, 20-30. |
| [19] |
Jin YH, Liu F, Huang W, Sun Q, Huang XZ (2019) Identification of reliable reference genes for qRT-PCR in the ephemeral plant Arabidopsis pumila based on full-length transcriptome data. Scientific Reports, 9, 8408.
DOI URL |
| [20] |
Kagale S, Koh C, Nixon J, Bollina V, Clarke WE, Tuteja R, Spillane C, Robinson SJ, Links MG, Clarke C, Higgins EE, Huebert T, Sharpe AG, Parkin IAP (2014) The emerging biofuel crop Camelina sativa retains a highly undifferentiated hexaploid genome structure. Nature Communications, 5, 3706.
DOI URL |
| [21] |
Karlgren A, Gyllenstrand N, Källman T, Sundström JF, Moore D, Lascoux M, Lagercrantz U (2011) Evolution of the PEBP gene family in plants: Functional diversification in seed plant evolution. Plant Physiology, 156, 1967-1977.
DOI PMID |
| [22] |
Li Q, Fan CM, Zhang XM, Wang X, Wu FQ, Hu RB, Fu YF (2014) Identification of a soybean MOTHER OF FT AND TFL1 homolog involved in regulation of seed germination. PLoS ONE, 9, e99642.
DOI URL |
| [23] | Li XC, Kang KC, Huang XZ, Fan YB, Song MM, Huang YJ, Ding JJ (2020) Genome-wide identification, phylogenetic analysis and expression profiling of the MKK gene family in Arabidopsis pumila. Hereditas (Beijing), 42, 403-421. (in Chinese with English abstract) |
| [李晓翠, 康凯程, 黄先忠, 范永斌, 宋苗苗, 黄韵杰, 丁佳佳 (2020) 小拟南芥MKK基因家族全基因组鉴定及进化和表达分析. 遗传, 42, 403-421.] | |
| [24] |
Liu S, Liu Y, Yang X, Tong C, Edwards D, Parkin IAP, Zhao M, Ma J, Yu J, Huang S, Wang X, Wang J, Lu K, Fang Z, Bancroft I, Yang TJ, Hu Q, Wang X, Yue Z, Li H, Yang L, Wu J, Zhou Q, Wang W, King GJ, Pires JC, Lu C, Wu Z, Sampath P, Wang Z, Guo H, Pan S, Yang L, Min J, Zhang D, Jin D, Li W, Belcram H, Tu J, Guan M, Qi C, Du D, Li J, Jiang L, Batley J, Sharpe AG, Park BS, Ruperao P, Cheng F, Waminal NE, Huang Y, Dong C, Wang L, Li J, Hu Z, Zhuang M, Huang Y, Huang J, Shi J, Mei D, Liu J, Lee TH, Wang J, Jin H, Li Z, Li X, Zhang J, Xiao L, Zhou Y, Liu Z, Liu X, Qin R, Tang X, Liu W, Wang Y, Zhang Y, Lee J, Kim HH, Denoeud F, Xu X, Liang X, Hua W, Wang X, Wang J, Chalhoub B, Paterson AH (2014) The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nature Communications, 5, 3930.
DOI URL |
| [25] |
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods, 25, 402-408.
DOI PMID |
| [26] |
Marks RA, Hotaling S, Frandsen PB, VanBuren R (2021) Representation and participation across 20 years of plant genome sequencing. Nature Plants, 7, 1571-1578.
DOI PMID |
| [27] |
Nakamura S, Abe F, Kawahigashi H, Nakazono K, Tagiri A, Matsumoto T, Utsugi S, Ogawa T, Handa H, Ishida H, Mori M, Kawaura K, Ogihara Y, Miura H (2011) A wheat homolog of MOTHER OF FT AND TFL1 acts in the regulation of germination. Plant Cell, 23, 3215-3229.
DOI URL |
| [28] |
Qiao X, Li M, Li LT, Yin H, Wu JY, Zhang SL (2015) Genome-wide identification and comparative analysis of the heat shock transcription factor family in Chinese white pear (Pyrus bretschneideri) and five other Rosaceae species. BMC Plant Biology, 15, 12.
DOI PMID |
| [29] |
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017) DnaSP 6: DNA sequence polymorphism analysis of large data sets. Molecular Biology and Evolution, 34, 3299-3302.
DOI URL |
| [30] |
Song XM, Ma X, Li CJ, Hu JJ, Yang QH, Wang T, Wang L, Wang JP, Guo D, Ge WN, Wang ZY, Li MM, Wang QM, Ren TZ, Feng SY, Wang LX, Zhang WM, Wang XY (2018) Comprehensive analyses of the BES1 gene family in Brassica napus and examination of their evolutionary pattern in representative species. BMC Genomics, 19, 346.
DOI URL |
| [31] |
Sun YQ, Shang LG, Zhu QH, Fan LJ, Guo LB (2022) Twenty years of plant genome sequencing: Achievements and challenges. Trends in Plant Science, 27, 391-401.
DOI URL |
| [32] |
Susila H, Jurić S, Liu L, Gawarecka K, Chung KS, Jin S, Kim SJ, Nasim Z, Youn G, Suh MC, Yu H, Ahn JH (2021) Florigen sequestration in cellular membranes modulates temperature-responsive flowering. Science, 373, 1137-1142.
DOI URL |
| [33] |
Tamaki S, Matsuo S, Wong HL, Yokoi S, Shimamoto K (2007) Hd3a protein is a mobile flowering signal in rice. Science, 316, 1033-1036.
DOI URL |
| [34] |
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular evolutionary genetics analysis version 6.0. Molecular Biology and Evolution, 30, 2725-2729.
DOI URL |
| [35] |
Taoka KI, Ohki I, Tsuji H, Furuita K, Hayashi K, Yanase T, Yamaguchi M, Nakashima C, Purwestri YA, Tamaki S, Ogaki Y, Shimada C, Nakagawa A, Kojima C, Shimamoto K (2011) 14-3-3 proteins act as intracellular receptors for rice Hd3a florigen. Nature, 476, 332-335.
DOI URL |
| [36] |
Thomas BC, Pedersen B, Freeling M (2006) Following tetraploidy in an Arabidopsis ancestor, genes were removed preferentially from one homeolog leaving clusters enriched in dose-sensitive genes. Genome Research, 16, 934-946.
PMID |
| [37] | Thompson JD, Gibson TJ, Higgins DG (2002) Multiple sequence alignment using ClustalW and ClustalX. Current Protocols in Bioinformatics, Chapter 2, Unit 2. 3. |
| [38] | Wang XY, Gowik U, Tang HB, Bowers JE, Westhoff P, Paterson AH (2009) Comparative genomic analysis of C 4 photosynthetic pathway evolution in grasses. Genome Biology, 10, R68. |
| [39] |
Wang X, Wang H, Wang J, Sun R, Wu J, Liu S, Bai Y, Mun JH, Bancroft I, Cheng F, Huang S, Li X, Hua W, Wang J, Wang X, Freeling M, Pires JC, Paterson AH, Chalhoub B, Wang B, Hayward A, Sharpe AG, Park BS, Weisshaar B, Liu B, Li B, Liu B, Tong C, Song C, Duran C, Peng C, Geng C, Koh C, Lin C, Edwards D, Mu D, Shen D, Soumpourou E, Li F, Fraser F, Conant G, Lassalle G, King GJ, Bonnema G, Tang H, Wang H, Belcram H, Zhou H, Hirakawa H, Abe H, Guo H, Wang H, Jin H, Parkin IAP, Batley J, Kim JS, Just J, Li J, Xu J, Deng J, Kim JA, Li J, Yu J, Meng J, Wang J, Min J, Poulain J, Wang J, Hatakeyama K, Wu K, Wang L, Fang L, Trick M, Links MG, Zhao M, Jin M, Ramchiary N, Drou N, Berkman PJ, Cai Q, Huang Q, Li R, Tabata S, Cheng S, Zhang S, Zhang S, Huang S, Sato S, Sun S, Kwon SJ, Choi SR, Lee TH, Fan W, Zhao X, Tan X, Xu X, Wang Y, Qiu Y, Yin Y, Li Y, Du Y, Liao Y, Lim Y, Narusaka Y, Wang Y, Wang Z, Li Z, Wang Z, Xiong Z, Zhang Z (2011) The genome of the mesopolyploid crop species Brassica rapa. Nature Genetics, 43, 1035-1039.
DOI URL |
| [40] |
Wang YP, Tang HB, Debarry JD, Tan X, Li JP, Wang XY, Lee TH, Jin HZ, Marler B, Guo H, Kissinger JC, Paterson AH (2012) MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Research, 40, e49.
DOI URL |
| [41] |
Wigge PA, Kim MC, Jaeger KE, Busch W, Schmid M, Lohmann JU, Weigel D (2005) Integration of spatial and temporal information during floral induction in Arabidopsis. Science, 309, 1056-1059.
DOI URL |
| [42] |
Xi WY, Liu C, Hou XL, Yu H (2010) MOTHER OF FT AND TFL1 regulates seed germination through a negative feedback loop modulating ABA signaling in Arabidopsis. Plant Cell, 22, 1733-1748.
DOI URL |
| [43] | Yadav CB, Bonthala VS, Muthamilarasan M, Pandey G, Khan Y, Prasad M (2015) Genome-wide development of transposable elements-based markers in foxtail millet and construction of an integrated database. DNA Research, 22, 79-90. |
| [44] |
Yamaguchi A, Kobayashi Y, Goto K, Abe M, Araki T (2005) TWIN SISTER OF FT (TSF) acts as a floral pathway integrator redundantly with FT. Plant and Cell Physiology, 46, 1175-1189.
PMID |
| [45] |
Yu XL, Liu H, Sang N, Li YF, Zhang TT, Sun J, Huang XZ (2019) Identification of cotton MOTHER OF FT AND TFL1 homologs, GhMFT1 and GhMFT2, involved in seed germination. PLoS ONE, 14, e0215771.
DOI URL |
| [46] |
Zhang B, Li C, Li Y, Yu H (2020) Mobile TERMINAL FLOWER1 determines seed size in Arabidopsis. Nature Plants, 6, 1146-1157.
DOI PMID |
| [47] |
Zhang J, Peng HW, Xia FC, Wang W (2021) A comparison of seed plants’ polyploids between the Qinghai-Tibet Plateau alpine and the Pan-Arctic regions. Biodiversity Science, 29, 1470-1480. (in Chinese with English abstract)
DOI |
|
[张军, 彭焕文, 夏富才, 王伟 (2021) 青藏高原高山区和泛北极地区种子植物多倍体比较. 生物多样性, 29, 1470-1480.]
DOI |
|
| [48] |
Zhou HS, Qi KJ, Liu X, Yin H, Wang P, Chen JQ, Wu JY, Zhang SL (2016) Genome-wide identification and comparative analysis of the cation proton antiporters family in pear and four other Rosaceae species. Molecular Genetics and Genomics, 291, 1727-1742.
DOI URL |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn