



生物多样性 ›› 2012, Vol. 20 ›› Issue (2): 184-192. DOI: 10.3724/SP.J.1003.2012.09211 cstr: 32101.14.SP.J.1003.2012.09211
收稿日期:2011-11-22
接受日期:2012-02-28
出版日期:2012-03-20
发布日期:2012-04-09
通讯作者:
吴海龙
作者简介:* E-mail: whlong@mail.ahnu.edu.cn基金资助:
Fang Li, Yilin Shu, Hailong Wu*( )
)
Received:2011-11-22
Accepted:2012-02-28
Online:2012-03-20
Published:2012-04-09
Contact:
Hailong Wu
摘要:
两栖类正经历全球范围内的种群衰退, 很多两栖动物集群灭绝事件与环境病原体(如壶菌(Batrachochytrium dendrobatidis)的侵扰有关。MHC基因的表达产物在有颌脊椎动物免疫应答过程中起关键作用, 其多态性通常与动物对疾病的抗性或易感性密切相关, 因而被认为是研究动物适应性进化的最佳候选基因之一。本文对中国特有的无尾两栖动物凹耳蛙(Odorrana tormota)MHC II类B基因多态性进行初步研究。首先, 利用1对通用引物扩增出凹耳蛙MHC II类B基因exon2长约180 bp的DNA片段。在此基础上, 利用ligation-mediated PCR进一步获取侧翼未知序列, 序列拼接后长2,030 bp, 包含exon2以及intron1和intron2的部分序列。基于上述序列设计出凹耳蛙B基因exon2特异性引物(IIQ1BU/IIQ1BD), 对该物种黄山种群32个样品进行PCR扩增和克隆测序, 共获得34个不同的等位基因, 等位基因序列核苷酸和氨基酸变异位点的比例分别为16.17%(33/204)和26.87%(18/67), 大多数氨基酸变异位点位于推测的抗原结合位点(antigen binding sites, ABS)。每个样品包含2-5个等位基因, 结合等位基因序列特征以及cDNA表达分析结果, 推测凹耳蛙至少拥有3个可表达的B基因座位。与文献报道的蛙科其他物种比较后发现, 尽管凹耳蛙目前的分布区非常狭窄, 但其MHC II类 B基因多态性明显高于蛙科其他动物。等位基因碱基替换模式提示凹耳蛙MHC II类B基因曾经历过强烈的正选择作用, ABS区的dN值显著大于dS (P<0.05), PAML软件包CODEML程序中不同模型的似然比检测(likelihood rate test)结果同样支持上述推论, 贝叶斯经验贝叶斯路径(Bayesian Empirical Bayes)共检测出5个显著受正选择作用的氨基酸位点。贝叶斯系统树的拓扑结构显示, 无尾两栖类不同科的等位基因分别形成单系群, 但蛙科不同属的等位基因未能形成单系群, 蛙属绿池蛙(Rana clamitans)的1个等位基因与臭蛙属凹耳蛙的部分等位基因享有共同的谱系关系, 提示蛙科不同属间的B基因存在跨种多态性。
李方, 疏义林, 吴海龙 (2012) 凹耳蛙MHC II类B基因第二外显子多态性分析. 生物多样性, 20, 184-192. DOI: 10.3724/SP.J.1003.2012.09211.
Fang Li, Yilin Shu, Hailong Wu (2012) Polymorphism of exon 2 of MHC Class II B gene in the Chinese concave- eared torrent frog (Odorrana tormota). Biodiversity Science, 20, 184-192. DOI: 10.3724/SP.J.1003.2012.09211.
| 引物 Primer | 序列 Sequence (5′-3′) |
|---|---|
| X1 | CGTTGGATATACGATCTGGAGG |
| X2 | GTACTTCGACAGTGATAAGGGATT |
| S1 | GGGTTTCTGTTCCAGGAGTCAG |
| S2 | AGCACTTGGTCTCCCCAGCTCGGT |
| IIQ1BU | GTACGGAGGATATCAGGTTTC |
| IIQ1BD | CAGGTGCAGTATATTCCTTCAT |
表1 本文用于分析凹耳蛙MHC II类B基因变异的引物序列
Table 1 Sequences of the primers used in the study of MHC II B genes variation in the Chinese concave-eared torrent frog
| 引物 Primer | 序列 Sequence (5′-3′) |
|---|---|
| X1 | CGTTGGATATACGATCTGGAGG |
| X2 | GTACTTCGACAGTGATAAGGGATT |
| S1 | GGGTTTCTGTTCCAGGAGTCAG |
| S2 | AGCACTTGGTCTCCCCAGCTCGGT |
| IIQ1BU | GTACGGAGGATATCAGGTTTC |
| IIQ1BD | CAGGTGCAGTATATTCCTTCAT |

图1 凹耳蛙MHC II类等位基因的氨基酸序列对位排列。* 根据Brown等(1993)的方法界定的抗原结合位点(ABS), # 参照Tong等(2006)的方法界定的ABS位点。
Fig. 1 Amino acid alignment of MHC II alleles of the Chinese concave-eared torrent frog. Sites involved in putative antigen binding sites are indicated by * (according to Brown et al., 1993) or by # (according to Tong et al., 2006), respectively.
| 位点 Sites | 同义替换率dN | 非同义替换率dS | dN/dS | Z检验P值 P(Z-test) |
|---|---|---|---|---|
| All | 0.047(0.014) | 0.050(0.018) | 0.710 | 0.909 |
| ABS ( | 0.116(0.037) | 0.046(0.029) | 1.081 | 0.028 |
| Non-ABS ( | 0.020(0.011) | 0.051(0.025) | 0.510 | 0.075 |
| ABS ( | 0.167(0.056) | 0.081(0.066) | 1.480 | 0.032 |
| Non-ABS ( | 0.025(0.011) | 0.044(0.018) | 0.491 | 0.176 |
表2 凹耳蛙MHC II类B基因exon2序列、ABS(putative antigen binding sites)区、非ABS区平均非同义替换率(rates of nonsynonymous substitutions per nonsynonymous site, dN)、同义替换率(rates of synonymous substitutions per synonymous site, dS)以及dN/dS比值。
Table 2 The average rates of nonsynonymous substitutions per nonsynonymous site (dN) and synonymous substitutions per synonymous site (dS) as well as the resulting ratio dN/dS for the sequenced section, the ABS (putative antigen binding sites) and non-ABS of MHC II B genes exon 2 of the Chinese concave-eared torrent frog.
| 位点 Sites | 同义替换率dN | 非同义替换率dS | dN/dS | Z检验P值 P(Z-test) |
|---|---|---|---|---|
| All | 0.047(0.014) | 0.050(0.018) | 0.710 | 0.909 |
| ABS ( | 0.116(0.037) | 0.046(0.029) | 1.081 | 0.028 |
| Non-ABS ( | 0.020(0.011) | 0.051(0.025) | 0.510 | 0.075 |
| ABS ( | 0.167(0.056) | 0.081(0.066) | 1.480 | 0.032 |
| Non-ABS ( | 0.025(0.011) | 0.044(0.018) | 0.491 | 0.176 |
| 模型 Model | 似然值 Likelihood | 估计参数 Estimate parameters | (LRT) 2△L | 受正选择位点 Positively selected sites |
|---|---|---|---|---|
| M1 | -655.027 | p0= 0.69482 p1=0.30518 ω0=0.02484 ω1=1.00000 | 52.322 (P<0.001) | None |
| M2 | -628.866 | p0=0.48425 p1=0.43958 p2=0.07617 ω0=0.00000 ω1=1.00000 ω2=15.77201 | 1T**, 10Y**, 34W**, 43Q*, 59Y** | |
| M7 | -655.106 | p= 0.01273 q= 0.02500 | 52.454 (P<0.001) | None |
| M8 | -628.879 | p0=0.92380 p=0.00503 q=0.00516 (p1= 0.07620) ω= 16.20861 | 1T**, 10Y**, 33S, 34W**, 43Q*, 59Y** |
表3 不同密码子进化模型对凹耳蛙MHC II类 B基因外显子2吻合度检验的参数与似然值
Table 3 Summary of parameter estimates and likelihood values of different models of codon evolution for exon 2 of MHC II B gene in the Chinese concave-eared torrent frog
| 模型 Model | 似然值 Likelihood | 估计参数 Estimate parameters | (LRT) 2△L | 受正选择位点 Positively selected sites |
|---|---|---|---|---|
| M1 | -655.027 | p0= 0.69482 p1=0.30518 ω0=0.02484 ω1=1.00000 | 52.322 (P<0.001) | None |
| M2 | -628.866 | p0=0.48425 p1=0.43958 p2=0.07617 ω0=0.00000 ω1=1.00000 ω2=15.77201 | 1T**, 10Y**, 34W**, 43Q*, 59Y** | |
| M7 | -655.106 | p= 0.01273 q= 0.02500 | 52.454 (P<0.001) | None |
| M8 | -628.879 | p0=0.92380 p=0.00503 q=0.00516 (p1= 0.07620) ω= 16.20861 | 1T**, 10Y**, 33S, 34W**, 43Q*, 59Y** |
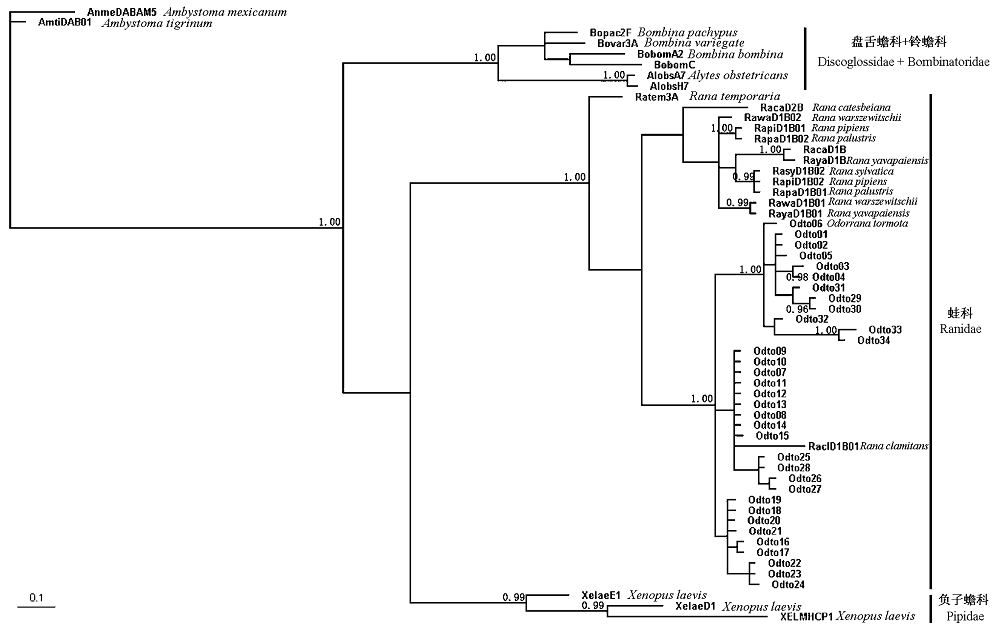
图2 利用贝叶斯方法基于MHC II类B基因外显子2核苷酸序列构建部分无尾两栖类的系统树, 以有尾类的虎纹顿口螈和墨西哥蝾螈的同源序列作为外类群。仅大于0.95的后验概率标在系统树的分支上。
Fig. 2 Phylogenetic relationship of anuran MHC class II B exon 2 nucleotide sequences constructed with Bayesian inference with two outgroup sequences from the caudate amphibian, Ambystoma tigrinum and Ambystoma mexicanum. Posterior probabilities above 0.95 are indicated on branches.
| [1] |
Alcaide M, Edwards SV, Negro JJ (2007) Characterization, polymorphism, and evolution of MHC class II B genes in birds of prey. Journal of Molecular Evolution, 65, 541-554.
DOI URL PMID |
| [2] |
Babik W, Pabijan M, Radwan J (2008) Contrasting patterns of variation in MHC loci in the Alpine newt. Molecular Ecology, 17, 2339-2355.
DOI URL PMID |
| [3] |
Bai CM, Garner TWJ, Li YM (2010) First evidence of Batrachochytrium dendrobatidis in China: discovery of chytridiomycosis in introduced American bullfrogs and native amphibians in the Yunnan Province, China. EcoHealth, 7, 127-134.
URL PMID |
| [4] |
Bos DH, DeWoody JA (2005) Molecular characterization of major histocompatibility complex class II alleles in wild tiger salamanders (Ambystoma tigrinum). Immunogenetics, 57, 775-781.
URL PMID |
| [5] |
Bos DH, Waldman B (2006) Evolution by recombination and transspecies polymorphism in the MHC class I gene of Xenopus laevis. Molecular Biology and Evolution, 23, 137-143.
URL PMID |
| [6] |
Brown JH, Jardetzky TS, Gorga JC, Stern LJ, Urban RG, Strominger JL, Wiley DC (1993) Three-dimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature, 364, 33-39.
URL PMID |
| [7] | Chen BH (陈璧辉) (1991) The Amphibian and Reptilian Fauna of Anhui (安徽两栖爬行动物志). Anhui Science and Technology Publishing House, Hefei. (in Chinese) |
| [8] | Cottage A, Yang AP, Maunders H, de Lacy RC, Ramsay NA (2001) Identification of DNA sequences flanking T-DNA insertions by PCR-walking. Plant Molecular Biology Reporter, 19, 321-327. |
| [9] | Daszak P, Cunningham AA, Hyatt AD (2003) Infectious disease and amphibian population decline. Diversity and Distributions, 9, 141-150. |
| [10] | Fei L (费梁), Hu SQ (胡淑琴), Ye CY (叶昌媛), Huang YZ (黄永昭) (2009) Fauna Sinica: Amphibia (Vol. 2. Anura) (中国动物志: 两栖纲(中卷无尾目)). Science Press, Beijing. (in Chinese) |
| [11] |
Feng AS, Narins PM, Xu CH, Lin WY, Yu ZL, Qiu Q, Xu ZM, Shen JX (2006) Ultrasonic communication in frogs. Nature, 440, 333-336.
DOI URL PMID |
| [12] |
Feng AS, Narins PM, Xu CH (2002) Vocal acrobatics in a Chinese frog, Amolops tormotus. Naturwissenschaften, 89, 352-356.
DOI URL PMID |
| [13] |
Harf R, Sommer S (2005) Association between major histocompatibility complex class II DRB alleles and parasite load in the hairy-footed gerbil, Gerbillurus paeba, in the southern Kalahari. Molecular Ecology, 14, 85-91.
DOI URL PMID |
| [14] |
Hauswaldt JS, Stuckas H, Pfautsch S, Tiedemann R (2007) Molecular characterization of MHC class II in a nonmodel anuran species, the fire-bellied toad Bombina bombina. Immunogenetics, 59, 479-491.
URL PMID |
| [15] |
Hedrick PW, Parker KM (1998) MHC variation in the endangered gila topminnow. Evolution, 52, 194-199.
DOI URL PMID |
| [16] |
Hovhannisyan Z, Weiss A, Martin A, Wiesner M, Tollefsen S, Yoshida K, Ciszewski C, Curran SA, Murray JA, David CS, Sollid LM, Koning F, Teyton L, Jabri B (2008) The role of HLA-DQ8 β57 polymorphism in the anti-gluten T-cell response in coeliac disease. Nature, 456, 534-538.
URL PMID |
| [17] |
Huelsenbeck JP, Ronquist F (2001) MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics, 17, 754-755.
URL PMID |
| [18] | Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Mammalian Protein Metabolism (ed. Munro HN), pp. 21-132. Academic Press, New York. |
| [19] |
Kiemnec-Tyburczy KM, Richmond JD, Savage AE, Zamudio KR (2010) Selection, trans-species polymorphism, and locus identification of major histocompatibility complex class IIβ alleles of New World ranid frogs. Immunogenetics, 62, 741-751.
DOI URL PMID |
| [20] | Klein J (1986) Natural History of the Major Histocompatibility Complex, 1st edn. John Wiley and Sons, New York, Chichester, Brisbane, Toronto, Singapore. |
| [21] |
Klein J, Sato A (1998) Birth of the major histocompatibility complex. Scandinavian Journal of Immunology, 47, 199-209.
DOI URL PMID |
| [22] | Langefors Å, Lohm J, Grahn M, Andersen Ø, Von Schantz T (2001) Association between major histocompatibility complex class II B alleles and resistance to Aeromonas salmonicida in Atlantic salmon. Proceedings of the Royal Society B: Biological Sciences, 268, 479-485. |
| [23] |
Laurens V, Chapusot C, del Rosario Ordonez M, Bentrari F, Padros MR, Tournefier A (2001) Axolotl MHC class II beta chain: predominance of one allele and alternative splicing of the beta 1 domain. European Journal of Immunology, 31, 506-515.
DOI URL PMID |
| [24] | Li PP, Lu YY, Li A, Yu LN (2008) The tadpole of a little-known frog, Rana tormotus Wu, 1977. Asiatic Herpetological Research, 11, 71-75. |
| [25] | May S, Beebee TJC (2009) Characterisation of major histocompatibility complex class II alleles in the natterjack toad, Bufo calamita. Conservation Genetics Resources, 1, 415-417. |
| [26] |
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Molecular Biology and Evolution, 3, 418-426.
DOI URL PMID |
| [27] |
Nielsen R, Yang Z (1998) Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics, 148, 929-936.
URL PMID |
| [28] |
Ohta Y, Goetz W, Hossain MZ, Nonaka M, Flajnik MF (2006) Ancestral organization of the MHC revealed in the amphibian Xenopus. The Journal of Immunology, 176, 3674-3685.
DOI URL PMID |
| [29] | Padgett-Flohr GE (2008) Pathogenicity of Batrachochytrium dendrobatidis in two threatened California amphibians: Rana draytonii and Ambystoma californiense. Herpetological Conservation and Biology, 3, 182-191. |
| [30] |
Parham P, Ohta T (1996) Population biology of antigen presentation by MHC class I molecules. Science, 272, 67-74.
URL PMID |
| [31] |
Paterson S, Wilson K, Pemberton JM (1998) Major hist- ocompatibility-complex variation associated with juvenile survival and parasite resistance in a large unmanaged ungulate population (Ovis aries L.). Proceedings of the National Academy of Sciences, USA, 95, 3714-3719.
DOI URL |
| [32] |
Pounds JA, Bustamante MR, Coloma LA, Consuegra JA, Fogden MPL, Foster PN, Marca EL, Masters KL, Merino-Viteri A, Puschendorf R, Ron SR, Sánchez-Azofeifa GA, Still CJ, Young BE (2006) Widespread amphibian extinctions from epidemic disease driven by global warming. Nature, 439, 161-167.
DOI URL PMID |
| [33] |
Richardson DS, Westerdahl H (2003) MHC diversity in two Acrocephalus species: the outbred great reed warble and the inbred Seychelles warbler. Molecular Ecology, 12, 3523-3529.
DOI URL PMID |
| [34] |
Sato A, Figueroa F, O’Huigin C, Steck N, Klein J (1998) Cloning of major histocompatibility complex (Mhc) genes from three spine stickleback, Gasterosteus aculeatus. Molecular Marine Biology and Biotechnology, 7, 221-231.
URL PMID |
| [35] |
Shen JX, Feng AS, Xu ZM, Yu ZL, Arch VS, Yu XJ, Narins PM (2008) Ultrasonic frogs show hyperacute phonotaxis to female courtship calls. Nature, 453, 914-916.
URL PMID |
| [36] |
Shen JX, Xu ZM, Yu ZL, Wang SA, Zheng DZ, Fan SC (2011) Ultrasonic frogs show extraordinary sex differences in auditory frequency sensitivity. Nature Communications, 2, 342. doi: 10.1038/ncomms1339.
URL PMID |
| [37] |
Su X, Wu XB, Yan P, Cao SY, Hu YL (2007) Rearrangement of a mitochondrial tRNA gene of the concave-eared torrent frog, Amolops tormotus. Gene, 394, 25-34.
DOI URL PMID |
| [38] |
Tamura K, Dudley J, Nei M, Kumar S (2007) MEGA 4: Molecular Evolutionary Genetics Analysis (MEGA4) software version 4.0. Molecular Biology and Evolution, 24, 1596-1599.
DOI URL PMID |
| [39] |
Teacher AGF, Garner TWJ, Nichols RA (2009) Evidence for directional selection at a novel major histocompatibility class I marker in wild common frogs (Rana temporaria) exposed to a viral pathogen (Ranavirus). PLoS ONE, 4, e4616.
DOI URL PMID |
| [40] |
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research, 25, 4876-4882.
URL PMID |
| [41] |
Tong JC, Bramson J, Kanduc D, Chow S, Sinha AA, Ranganathan S (2006) Modeling the bound conformation of pemphigus vulgaris-associated peptides to MHC Class II DR and DQ alleles. Immunome Research, 2, 1.
DOI URL PMID |
| [42] | Westerdahl H, Waldenstrom J, Hansson B, Hasselquist D, von Schantz T, Bensch S (2005) Associations between malaria and MHC genes in a migratory songbird. Proceedings of the Royal Society B: Biological Sciences, 272, 1511-1518. |
| [43] |
Wong WSW, Yang ZH, Goldman N, Nielsen R (2004) Accuracy and power of statistical methods for detecting adaptive evolution in protein coding sequences and for identifying positively selected sites. Genetics, 168, 1041-1051.
URL PMID |
| [44] | Yan JC, Luo TL, Wu HL (2011) Development and charac- terization of twelve polymorphic microsatellite loci for the Chinese concave-eared frog (Odorrana tormota). Conser- vation Genetics Resource, 3, 225-227. |
| [45] |
Yang ZH (1997) PAML: a program package for phylogenetic analysis by maximum likelihood. Computer Applications in the Biosciences, 13, 555-556.
URL PMID |
| [46] |
Zeisset I, Beebee TJC (2009) Molecular characterization of major histocompatibility complex class II alleles in the common frog, Rana temporaria. Molecular Ecology Resources, 9, 738-745.
URL PMID |
| [1] | 毛锐锐, 沈拓, 李慧, 田琳楚, 谭海蓉, 卢李荣, 吴小刚, 范宗骥, 伍国仪, 李杰, 吴勇, 朱弼成, 肖治术. 广东车八岭国家级自然保护区无尾两栖类动物鸣声特征数据集[J]. 生物多样性, 2024, 32(10): 24356-. |
| [2] | 金彦君, 赵龙辉, 覃远玉, 汪继超. 海南国家公园霸王岭片区无尾两栖类鸣声多样性: 基于自动录音技术[J]. 生物多样性, 2023, 31(1): 22360-. |
| [3] | 宋云枫, 陈传武, 王彦平. 中国两栖类生活史和生态学特征数据集[J]. 生物多样性, 2022, 30(3): 22053-. |
| [4] | 蒋志刚, 罗振华. 物种受威胁状况评估: 研究进展与中国的案例[J]. 生物多样性, 2012, 20(5): 612-622. |
| [5] | 李成, 戴强, 王跃招, 顾海军, 刘志君. 四川无尾两栖类的繁殖模式[J]. 生物多样性, 2005, 13(4): 290-297. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
备案号:京ICP备16067583号-7
Copyright © 2026 版权所有 《生物多样性》编辑部
地址: 北京香山南辛村20号, 邮编:100093
电话: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn
![]()