



Biodiv Sci ›› 2017, Vol. 25 ›› Issue (6): 638-646. DOI: 10.17520/biods.2017060 cstr: 32101.14.biods.2017060
• Original Papers • Previous Articles Next Articles
Qiujie Zhou1, Yacheng Cai1, Wei Lun Ng1, Wei Wu1, Seping Dai2, Feng Wang3, Renchao Zhou1,*( )
)
Received:2017-02-27
Accepted:2017-04-12
Online:2017-06-20
Published:2017-07-10
Contact:
Zhou Renchao
Qiujie Zhou, Yacheng Cai, Wei Lun Ng, Wei Wu, Seping Dai, Feng Wang, Renchao Zhou. Molecular evidence for natural hybridization between two Melastoma species endemic to Hainan and their widespread congeners[J]. Biodiv Sci, 2017, 25(6): 638-646.

Fig. 1 Morphological comparison between Melastoma penicillatum, M. candidum and their putative hybrid. (A) M. penicillatum, (B) putative hybrid, (C) M. candidum. Fruits of M. penicillatum (D), putative hybrid (E) and M. candidum (F).
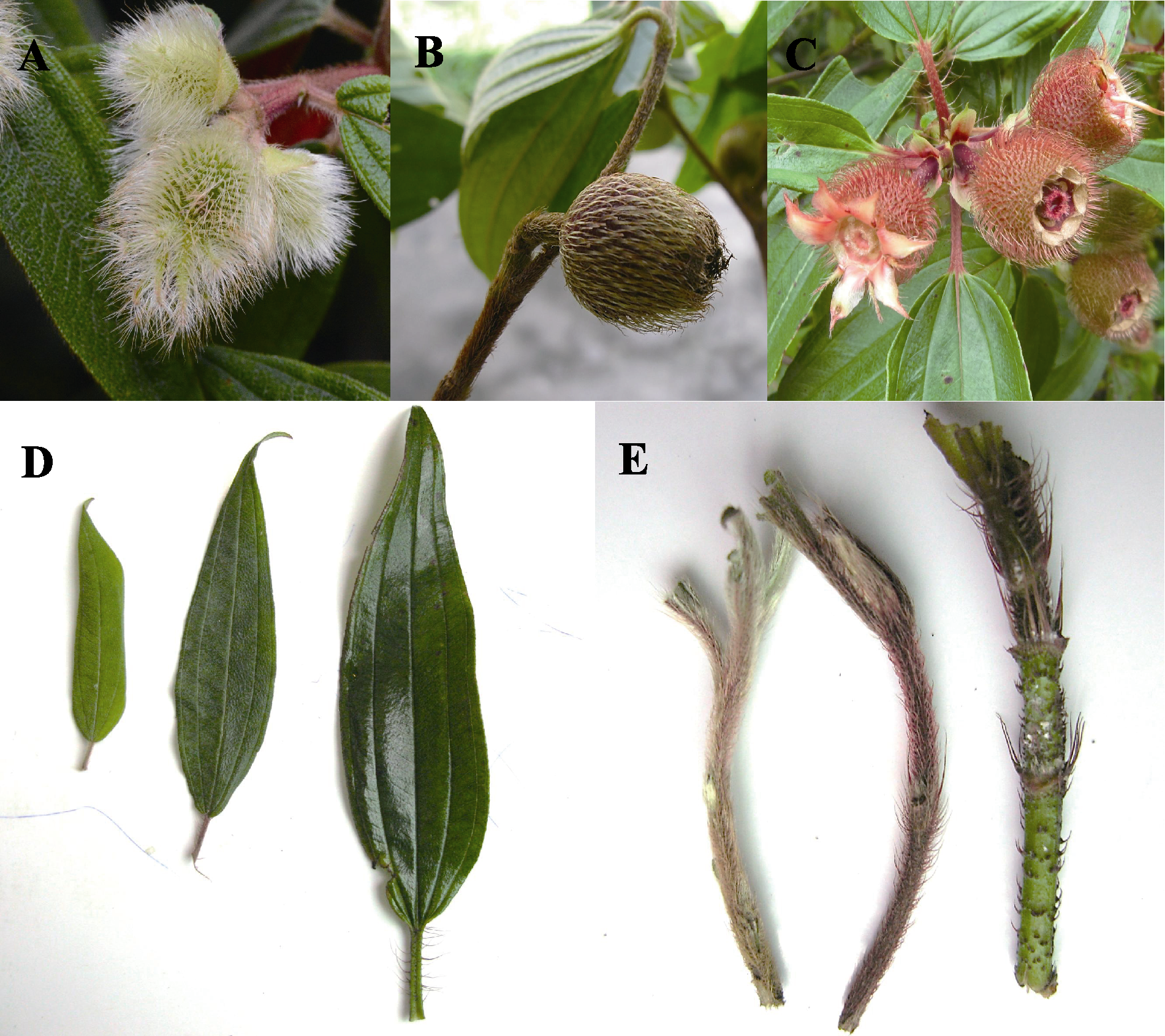

Fig. 2 Morphological comparison between Melastoma dendrisetosum, M. sanguineum and their putative hybrid. Fruits of M. dendrisetosum (A), putative hybrid (B) and M. sanguineum (C). D, Leaves of M. dendrisetosum (left), putative hybrid (middle) and M. sanguineum (right). E, Young twigs of M. dendrisetosum (left), putative hybrid (middle) and M. sanguineum (right).
| 类群 Taxon | 采样地点 Location | 个体数量 Sample size |
|---|---|---|
| 野牡丹 M. candidum (Cf) | 福建龙海 Longhai, Fujian | 19 |
| 野牡丹 M. candidum (Ch) | 海南吊罗山 Diaoluoshan, Hainan | 20 |
| 毛菍 M. sanguineum (S) | 海南吊罗山 Diaoluoshan, Hainan | 20 |
| 紫毛野牡丹 M. penicillatum (P) | 海南吊罗山 Diaoluoshan, Hainan | 20 |
| 枝毛野牡丹 M. dendrisetosum (D) | 海南吊罗山 Diaoluoshan, Hainan | 20 |
| 紫毛野牡丹与野牡丹的嫌疑杂种 Putative hybrid between P and C | 海南吊罗山 Diaoluoshan, Hainan | 1 |
| 枝毛野牡丹与毛菍的嫌疑杂种 Putative hybrid between D and S | 海南吊罗山 Diaoluoshan, Hainan | 4 |
Table 1 Sampling details of four species of Melastoma and two putative hybrids used in this study
| 类群 Taxon | 采样地点 Location | 个体数量 Sample size |
|---|---|---|
| 野牡丹 M. candidum (Cf) | 福建龙海 Longhai, Fujian | 19 |
| 野牡丹 M. candidum (Ch) | 海南吊罗山 Diaoluoshan, Hainan | 20 |
| 毛菍 M. sanguineum (S) | 海南吊罗山 Diaoluoshan, Hainan | 20 |
| 紫毛野牡丹 M. penicillatum (P) | 海南吊罗山 Diaoluoshan, Hainan | 20 |
| 枝毛野牡丹 M. dendrisetosum (D) | 海南吊罗山 Diaoluoshan, Hainan | 20 |
| 紫毛野牡丹与野牡丹的嫌疑杂种 Putative hybrid between P and C | 海南吊罗山 Diaoluoshan, Hainan | 1 |
| 枝毛野牡丹与毛菍的嫌疑杂种 Putative hybrid between D and S | 海南吊罗山 Diaoluoshan, Hainan | 4 |
| 类群 Taxon | cam | tpi | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 292 | 244 | 204 | 225 | 258 | 272 | 315 | 340 | 568 | |
| 野牡丹 Melastoma candidum (C) | A | G | T | T | T | G | A | T | C |
| 紫毛野牡丹 M. penicillatum (P) | G | A | A | C | C | A | G | C | T |
| 紫毛野牡丹和野牡丹的嫌疑杂种 Putative hybrid between P and C | R | R | W | Y | Y | R | R | Y | Y |
Table 2 The base composition of the differentially fixed sites at the cam and tpi genes between Melastoma penicillatum and M. candidum in their putative hybrid (R = A + G; Y = C + T; W = T + A)
| 类群 Taxon | cam | tpi | |||||||
|---|---|---|---|---|---|---|---|---|---|
| 292 | 244 | 204 | 225 | 258 | 272 | 315 | 340 | 568 | |
| 野牡丹 Melastoma candidum (C) | A | G | T | T | T | G | A | T | C |
| 紫毛野牡丹 M. penicillatum (P) | G | A | A | C | C | A | G | C | T |
| 紫毛野牡丹和野牡丹的嫌疑杂种 Putative hybrid between P and C | R | R | W | Y | Y | R | R | Y | Y |
| 类群 Taxon | cam | chi | gbss | tpi | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 190 | 329 | 661 | 79 | 258 | 315 | 414 | 80 | 240 | 286 | |
| 毛菍 M. sanguineum (S) | T | C | G | C | A | G | G | G | T | A |
| 枝毛野牡丹 M. dendrisetosum (D) | C | T | A | T | C | A | C | A | C | T |
| 枝毛野牡丹和毛菍的嫌疑杂种 Putative hybrid between D and S | Y | Y | R | Y | M | R | S | R | Y | W |
Table 3 The base composition of the differentially fixed sites at the four nuclear genes between Melastoma dendrisetosum and M. sanguineum in their putative hybrid (R = A + G; Y = C + T; W = T + A; M = A + C; S = C + G)
| 类群 Taxon | cam | chi | gbss | tpi | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 190 | 329 | 661 | 79 | 258 | 315 | 414 | 80 | 240 | 286 | |
| 毛菍 M. sanguineum (S) | T | C | G | C | A | G | G | G | T | A |
| 枝毛野牡丹 M. dendrisetosum (D) | C | T | A | T | C | A | C | A | C | T |
| 枝毛野牡丹和毛菍的嫌疑杂种 Putative hybrid between D and S | Y | Y | R | Y | M | R | S | R | Y | W |
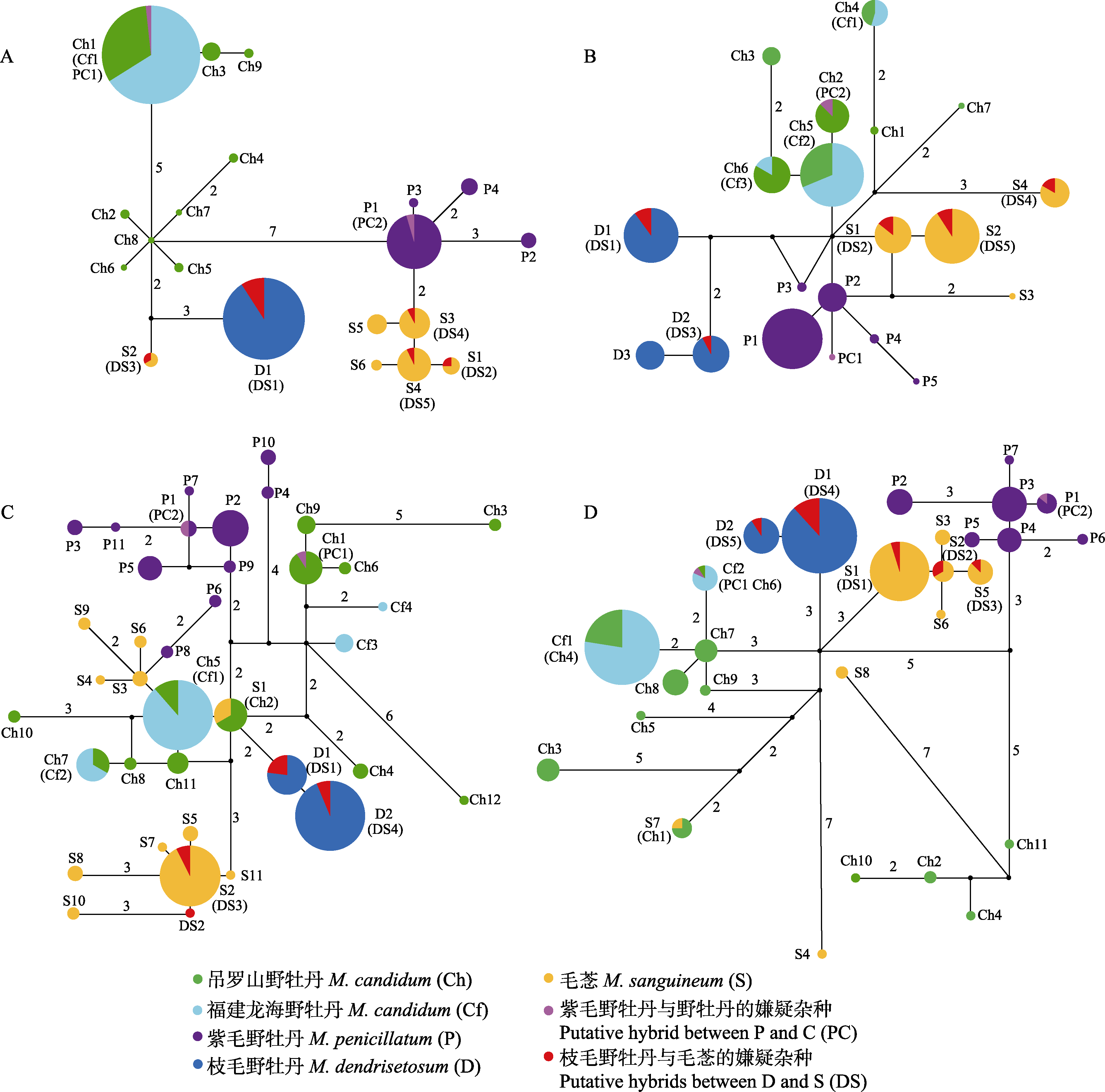
Fig. 3 Median-joining networks of tpi (A), gbss (B), chi (C) and cam (D) genes of the four Melastoma species and the two putative hybrids. The numbers around the connecting lines between haplotypes represent the number of mutational steps between them, while those without numbers represent only one mutational step. Small black circles represent hypothetical or unsampled haplotypes.
| 类群 Taxon | 个体编号 Sample ID | tpi | cam | chi | gbss |
|---|---|---|---|---|---|
| 紫毛野牡丹和野牡丹的嫌疑杂种 Putative hybrid (PC) | PC | PC1PC2 | PC1PC2 | PC1PC2 | PC1 PC2 |
| 枝毛野牡丹和毛菍的嫌疑杂种 Putative hybrid (DS) | DS_1 | DS1DS2 | DS1DS4 | DS1 DS2 | DS1DS2 |
| DS_2 | DS1DS3 | DS2DS4 | DS1DS3 | DS3DS4 | |
| DS_3 | DS1DS4 | DS3DS5 | DS3DS4 | DS1DS2 | |
| DS_4 | DS1DS5 | DS2DS4 | DS1DS3 | DS1DS5 |
Table 4 Genotypes of two putative hybrids of Melastoma at four nuclear genes (tpi, cam, chi and gbss). Haplotypes with single, double, wavy, or dotted underlines have identical sequences with those of M. candidum, M. penicillatum, M. sanguineum, and M. dendrisetosum, respectively. Haplotypes without underline are unique to the putative hybrids.
| 类群 Taxon | 个体编号 Sample ID | tpi | cam | chi | gbss |
|---|---|---|---|---|---|
| 紫毛野牡丹和野牡丹的嫌疑杂种 Putative hybrid (PC) | PC | PC1PC2 | PC1PC2 | PC1PC2 | PC1 PC2 |
| 枝毛野牡丹和毛菍的嫌疑杂种 Putative hybrid (DS) | DS_1 | DS1DS2 | DS1DS4 | DS1 DS2 | DS1DS2 |
| DS_2 | DS1DS3 | DS2DS4 | DS1DS3 | DS3DS4 | |
| DS_3 | DS1DS4 | DS3DS5 | DS3DS4 | DS1DS2 | |
| DS_4 | DS1DS5 | DS2DS4 | DS1DS3 | DS1DS5 |
| [1] | Abbott R, Albach D, Ansell S, Arntzen JW, Baird SJE, Bierne N, Boughman J, Brelsford A, Buerkle CA, Buggs R, Dieckmann U, Eroukhmanoff F, Grill A, Cahan SH, Hermansen JS, Hewitt G, Hudson AG, Jiggins C, Jones J, Keller B, Marczewski T, Mallet J, Martinez-Rodriguez P, Möst M, Mullen S, Nichols R, Nolte AW, Parisod C, Pfennig K, Rice AM, Ritchie MG, Seifert B, Smadja CM, Stelkens R, Szymura JM, Väinölä R, Wolf JBW, Zinner D, Butlin RK (2013) Hybridization and speciation. Journal of Evolutionary Biology, 26, 229-246. |
| [2] | Arnold ML (1997) xNatural Hybridization and Evolution. Oxford University Press, Oxford. |
| [3] | Arnold ML (2006) Evolution Through Genetic Exchange. Oxford University Press, Oxford. |
| [4] | Arnold ML, Martin NH (2009) Adaptation by introgression. Journal of Biology, 8, 82. |
| [5] | Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution, 16, 37-48. |
| [6] | Chao LF, Chen YY, Wang SQ, Liu T, Wu W, Dai SP, Wang F, Fan Q, Shi SH, Zhou RC (2014) One species or two? Multilocus analysis of nucleotide variation of Melastoma penicillatum and Melastoma sanguineum (Melastomataceae) in Hainan, China. Biochemical Systematics and Ecology, 55, 275-282. |
| [7] | Chen J, Renner SS (2007) Melastomataceae. In: Flora of China (ed. Editorial Committee of Flora of China), Vol. 13, pp. 360-399. Science Press, Beijing and Missouri Botanical Garden, St Louis. |
| [8] | Chen J (1983) A study of Melastoma (Melastomataceae) in China. Journal of South China Agricultural College, 4(1), 31-36. |
| (in Chinese) [陈介 (1983) 中国野牡丹科野牡丹属植物的研究. 华南农学院学报, 4(1), 31-36.] | |
| [9] | Chen J (1984) Melastomataceae. In: Flora Reipublicae Popularis Sinicae (ed. Editorial Committee of Flora Reipublicae Popularis Sinicae, Chinese Academy of Sciences), Tomus, 53(1), pp. 152-161. Science Press, Beijing. |
| (in Chinese) [陈介 (1984) 野牡丹科. 见: 中国植物志 (中国科学院中国植物志编辑委员会编), 53(1), 152-161. 科学出版社, 北京.] | |
| [10] | Dai SP, Wu W, Zhang RS, Liu T, Chen YY, Shi SH, Zhou RC (2012) Molecular evidence for hybrid origin of Melastoma intermedium. Biochemical Systematics and Ecology, 41, 136-141. |
| [11] | Doyle JJ (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin, 19, 11-15. |
| [12] | Grant PR, Grant BR (2014) Speciation undone. Nature, 507, 178-179. |
| [13] | Gross CL (1993) The breeding system and pollinators of Melastoma affine (Melastomataceae): a pioneer shrub in tropical Australia. Biotropica, 25, 468-474. |
| [14] | Huxel GR (1999) Rapid displacement of native species by invasive species: effects of hybridization. Biological Conservation, 89, 143-152. |
| [15] | Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25, 1451-1452. |
| [16] | Liu T, Chen YY, Chao LF, Wang SQ, Wu W, Dai SP, Wang F, Fan Q, Zhou RC (2014) Extensive hybridization and introgression between Melastoma candidum and M. sanguineum. PLoS ONE, 9, e96680. |
| [17] | Lu GH, Wu WH, Wang RZ, Li XL, Wang YQ (2009) Division of labor of heteromorphic stamens in Melastoma malabathricum. Biodiversity Science, 17, 174-181. (in Chinese with English abstract) |
| [路国辉, 武文华, 王瑞珍, 李新亮, 王英强 (2009) 野牡丹异型雄蕊的功能分化. 生物多样性, 17, 174-181.] | |
| [18] | Luo ZL, Zhang DX (2005) A review of heteranthery in flowering plants. Journal of Tropical and Subtropical Botany, 13, 536-542. |
| [19] | Luo ZL, Zhang DX, Renner SS (2008) Why two kinds of stamens in buzz-pollinated flowers? Experimental support for Darwin’s division-of-labour hypothesis. Functional Ecology, 22, 794-800. |
| [20] | Luo ZL, Gu L, Zhang DX (2009) Intrafloral differentiation of stamens in heterantherous flowers. Journal of Systematics and Evolution, 47, 43-56. |
| [21] | Mallet J (2005) Hybridization as an invasion of the genome. Trends in Ecology & Evolution, 20, 229-237. |
| [22] | Meyer K (2001) Revision of the Southeast Asian genus Melastoma. Blumea, 46, 351-398. |
| [23] | Moody ML, Les DH (2002) Evidence of hybridity in invasive watermilfoil (Myriophyllum) populations. Proceedings of the National Academy of Sciences, USA, 99, 14867-14871. |
| [24] | Peng DH, Lan SR, Wu SS (2014) Pollination biology and breeding system of Melastoma dendrisetosum. Forest Research, 27(1), 11-16. (in Chinese with English abstract) |
| [彭东辉, 兰思仁, 吴沙沙 (2014) 中国特有种枝毛野牡丹传粉生物学及繁育系统研究. 林业科学研究, 27(1), 11-16.] | |
| [25] | Renner SS, Meyer K (2001) Melastomeae come full circle: biogeographic reconstruction and molecular clock dating. Evolution, 55, 1315-1324. |
| [26] | Rhymer JM, Simberloff D (1996) Extinction by hybridization and introgression. Annual Review of Ecology and Systematics, 27, 83-109. |
| [27] | Rieseberg LH (1997) Hybrid origins of plant species. Annual Review of Ecology and Systematics, 28, 359-389. |
| [28] | Strand AE, Leebens-Mack J, Milligan BG (1997) Nuclear DNA-based markers for plant evolutionary biology. Molecular Ecology, 6, 113-118. |
| [29] | Taberlet P, Gielly L, Pautou G, Bouvet J (1991) Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Molecular Biology, 17, 1105-1109. |
| [30] | Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Research, 25, 4876-4882. |
| [31] | Todesco M, Pascual MA, Owens GL, Ostevik KL, Moyers BT, Hübner S, Heredia SM, Hahn MA, Caseys C, Bock DG, Rieseberg LH (2016) Hybridization and extinction. Evolutionary Applications, 9, 892-908. |
| [32] | Wolf DE, Takebayashi N, Rieseberg LH (2001) Predicting the risk of extinction through hybridization. Conservation Biology, 15, 1039-1053. |
| [1] | Jing Gan Xiangxu Liu Xueming Lu Xing Yue. China's Large Cities in Global Biodiversity Hotspots: Conservation Policies and Optimization Directions [J]. Biodiv Sci, 2025, 33(5): 24529-. |
| [2] | Zixuan Zeng Rui Yang Yue Huang Luyao Chen. Characteristics of bird diversity and environmental relationships in Tsinghua University campus [J]. Biodiv Sci, 2025, 33(5): 24373-. |
| [3] | Hao Zhou, Mingyi Wang, Chuge Zhang, Zhishu Xiao, Fang Ouyang. The status and challenges of insect hotels in the conservation of urban solitary bees and wasps diversity [J]. Biodiv Sci, 2025, 33(5): 24472-. |
| [4] | Xiaoyu Zhu, Chenhao Wang, Zhongjun Wang, Yujun Zhang. Research progress and prospect of urban green space biodiversity [J]. Biodiv Sci, 2025, 33(5): 25027-. |
| [5] | Xin Wang, Femgyu Bao. Analysis of the ecological restoration effect of South Dianchi National Wetland Park based on the enhancement of bird diversity [J]. Biodiv Sci, 2025, 33(5): 24531-. |
| [6] | Yue Ming, Peiyao Hao, Lingqian Tan, Xi Zheng. A study on urban biodiversity conservation and enhancement in china based on the concept of green and high-quality development of cities [J]. Biodiv Sci, 2025, 33(5): 24524-. |
| [7] | Murong Yi, Ping Lu, Yong Peng, Yong Tang, Jiuheng Xu, Haoping Yin, Luyang Zhang, Xiaodong Weng, Mingxiao Di, Juan Lei, Chenqi Lu, Rujun Cao, Nianhua Dai, Deyang Zhan, Mei Tong, Zhiming Lou, Yonggang Ding, Jing Chai, Jing Che. Population status and habitat of Critically Endangered Jiangxi giant salamander (Andrias jiangxiensis) [J]. Biodiv Sci, 2025, 33(4): 24145-. |
| [8] | Motong Li, Tuo He, Wei Li, Jing Liao, Yan Zeng. Regulating international trade in wild fauna and flora: An analysis of CITES terminology [J]. Biodiv Sci, 2025, 33(4): 24545-. |
| [9] | Xiaoqiang Lu, Shanshan Dong, Yue Ma, Xu Xu, Feng Qiu, Mingyue Zang, Yaqiong Wan, Luanxin Li, Cigang Yu, Yan Liu. Current status, challenges, and prospects of frontier technologies in biodiversity conservation applications [J]. Biodiv Sci, 2025, 33(4): 24440-. |
| [10] | Guo Yutong, Li Sucui, Wang Zhi, Xie Yan, Yang Xue, Zhou Guangjin, You Chunhe, Zhu Saning, Gao Jixi. Coverage and distribution of national key protected wild species in China’s nature reserves [J]. Biodiv Sci, 2025, 33(3): 24423-. |
| [11] | Zhao Weiyang, Wang Wei, Ma Bingran. Advances and prospects in research on other effective area-based conservation measures (OECMs) [J]. Biodiv Sci, 2025, 33(3): 24525-. |
| [12] | Zhou Zhihua, Jin Xiaohua, Luo Ying, Li Diqiang, Yue Jianbing, Liu Fang, He Tuo, Li Xi, Dong Hui, Luo Peng. Analyses and suggestions on mechanisms of forestry and grassland administrations in China to achieve targets of Kunming-Montreal Global Biodiversity Framework [J]. Biodiv Sci, 2025, 33(3): 24487-. |
| [13] | Liu Li, Zang Mingyue, Ma Yue, Wan Yaqiong, Hu Feilong, Lu Xiaoqiang, Liu Yan. Measures, progress and prospects of central-local cooperation in the implementation of the National Biodiversity Strategy and Action Plan [J]. Biodiv Sci, 2025, 33(3): 24532-. |
| [14] | Yang Song, Jun Liu, Shaolin He, Wei Xu, Chen Cheng, Bo Liu, Jiqing Yu. Mainstreaming path of biodiversity conservation for Chinese energy enterprises [J]. Biodiv Sci, 2025, 33(1): 24345-. |
| [15] | Tuo He, Yan Zeng, Yafang Yin, Kun Zhang, Liangchen Yuan, Hui Dong, Zhihua Zhou. Consolidating the scientific foundation for global wild plant conservation and sustainable trade—Comments on the 27th Meeting of the Plants Committee of CITES [J]. Biodiv Sci, 2024, 32(9): 24390-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2022 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn ![]()