



Biodiv Sci ›› 2017, Vol. 25 ›› Issue (6): 675-682. DOI: 10.17520/biods.2017042 cstr: 32101.14.biods.2017042
Special Issue: 物种形成与系统进化
• Original Papers • Previous Articles Next Articles
Jiangping Shu1,2, Li Liu1,2, Hui Shen2, Xiling Dai1, Quanxi Wang1,4, Yuehong Yan2,3,4,*( )
)
Received:2017-02-18
Accepted:2017-06-20
Online:2017-06-20
Published:2017-07-10
Contact:
Yan Yuehong
Jiangping Shu, Li Liu, Hui Shen, Xiling Dai, Quanxi Wang, Yuehong Yan. The complex reticulate evolutionary relationships of early terrestrial plants as revealed by phylogenomics analysis[J]. Biodiv Sci, 2017, 25(6): 675-682.
| 物种 Species | 分类 Classification | BUSCO评估结果 BUSCO results |
|---|---|---|
| 福建观音座莲 Angiopteris fokiensis | 真蕨类植物 Monilophytes | C: 66.4% [S: 43.2%, D: 23.2%], F: 5.9%, M: 27.7%, n: 1440 |
| 欧洲云杉 Picea abies | 种子植物 Spermatophytes | C: 34.0% [S: 28.9%, D: 5.1%], F: 7.4%, M: 58.6%, n: 1,440 |
| 江南卷柏 Selaginella moellendorffii | 石松类植物 Lycophytes | C: 63.2% [S: 10.0%, D: 53.2%], F: 4.7%, M: 32.1%, n: 1,440 |
| 小立碗藓 Physcomitrella patens | 苔藓植物 Bryophytes | C: 70.1% [S: 46.0%, D: 24.1%], F: 2.6%, M: 27.3%, n: 1,440 |
| 莱茵衣藻 Chlamydomonas reintmrdtii | 藻类植物 Thallophytes | C: 18.8% [S: 17.9%, D: 0.9%], F: 1.7%, M: 79.5%, n: 1,440 |
Table 1 The assessment results of assembly completeness of transcriptome and genome
| 物种 Species | 分类 Classification | BUSCO评估结果 BUSCO results |
|---|---|---|
| 福建观音座莲 Angiopteris fokiensis | 真蕨类植物 Monilophytes | C: 66.4% [S: 43.2%, D: 23.2%], F: 5.9%, M: 27.7%, n: 1440 |
| 欧洲云杉 Picea abies | 种子植物 Spermatophytes | C: 34.0% [S: 28.9%, D: 5.1%], F: 7.4%, M: 58.6%, n: 1,440 |
| 江南卷柏 Selaginella moellendorffii | 石松类植物 Lycophytes | C: 63.2% [S: 10.0%, D: 53.2%], F: 4.7%, M: 32.1%, n: 1,440 |
| 小立碗藓 Physcomitrella patens | 苔藓植物 Bryophytes | C: 70.1% [S: 46.0%, D: 24.1%], F: 2.6%, M: 27.3%, n: 1,440 |
| 莱茵衣藻 Chlamydomonas reintmrdtii | 藻类植物 Thallophytes | C: 18.8% [S: 17.9%, D: 0.9%], F: 1.7%, M: 79.5%, n: 1,440 |
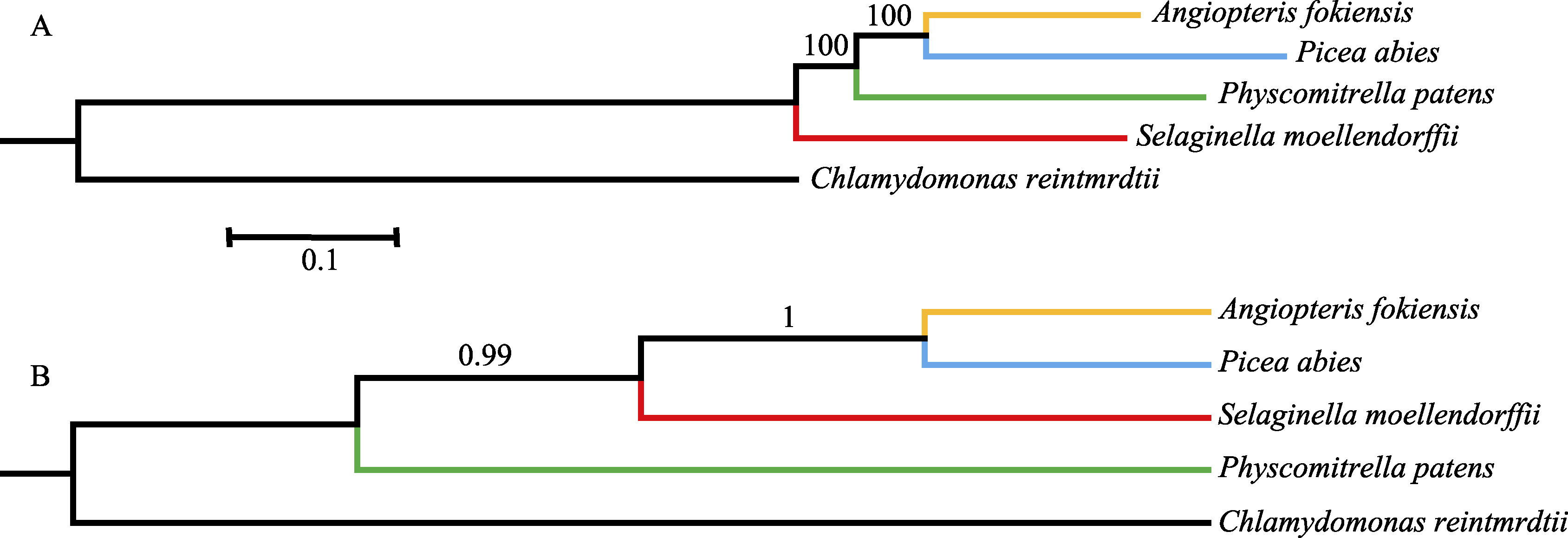
Fig. 1 The phylogenetic trees based on concatenation and coalescence methods. (A) The maximum likelihood tree based on concatenation method; (B) The species tree based on coalescence method.
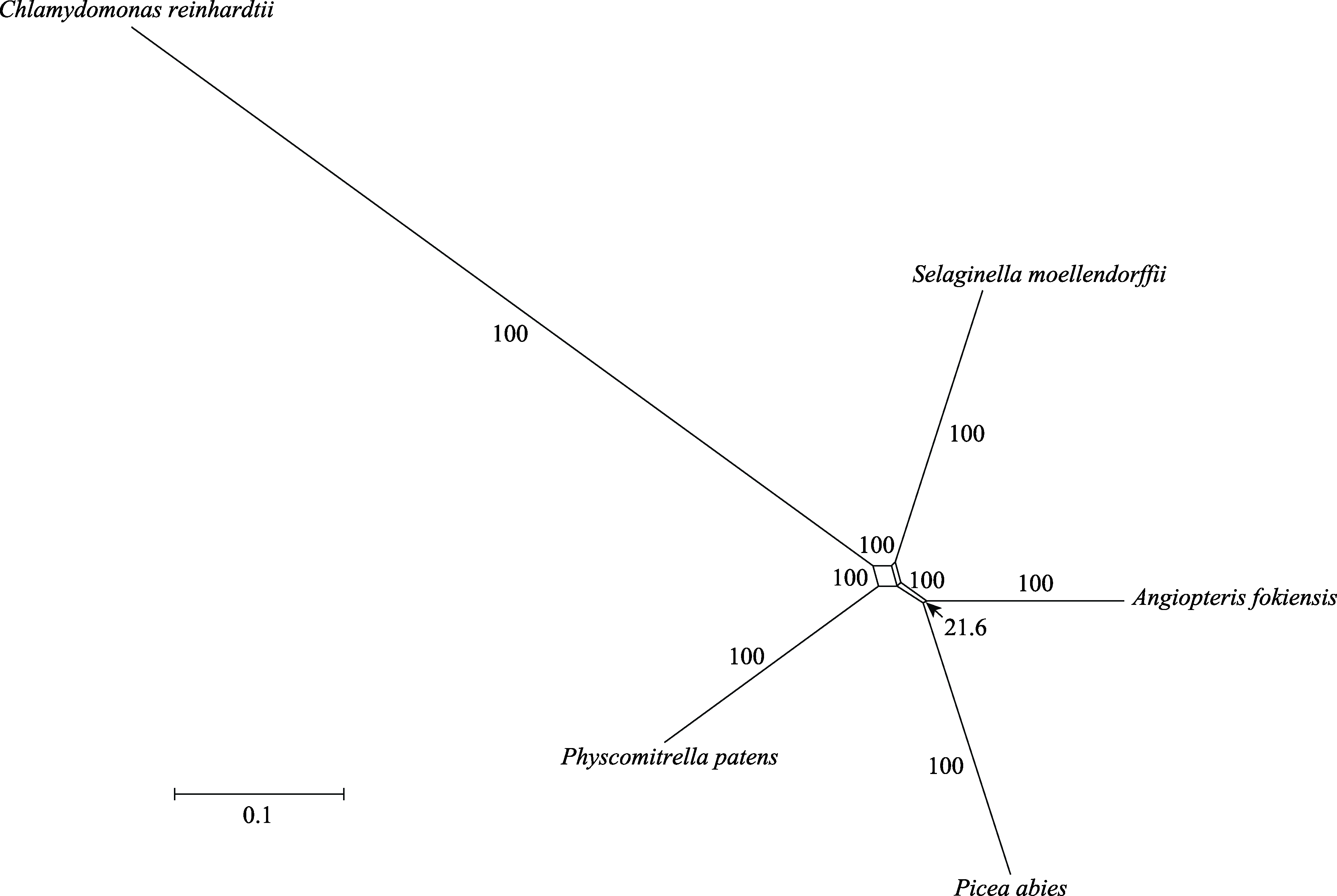
Fig. 3 The phylogenetic network of early land plants based on 1,668 genes. The numbers mean the bootstrap support of each split branch. In addition to the shortest split branch (arrow), the bootstrap support of other split branches is 100%. The parallel split branches are the same type of split branch.
| [1] | Arnold CA (2013) An Introduction to Paleobotany. Read Books Ltd., New York. |
| [2] | Bandelt HJ, Forster P, Röhl A (1999) Median-joining networks for inferring intraspecific phylogenies. Molecular Biology and Evolution, 16, 37-48. |
| [3] | Bohlin K (1901) Utkast till de gröna algernas och arkegoniaternas fylogeni. Almqvist & Wiksells Boktryckeri AB, Uppsala. |
| [4] | Bower FO (1935) Primitive land plants. Science, 81, 537-539. |
| [5] | Bryant D, Moulton V (2004) Neighbor-net: an agglomerative method for the construction of phylogenetic networks. Molecular Biology and Evolution, 21, 255-265. |
| [6] | Campbell D (1899) Lectures on the Evolution of Plants. Kessinger Publishing, London. |
| [7] | Church AH (1919) Thalassiophyta and the Subaerial Transmigration. Oxford University Press, Oxford. |
| [8] | Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Molecular Ecology, 9, 1657-1659. |
| [9] | Cranston KA, Hurwitz B, Ware D, Stein L, Wing RA (2009) Species trees from highly incongruent gene trees in rice. Systematic Biology, 58, 489-500. |
| [10] | Darriba D, Taboada GL, Doallo R, Posada D (2011) ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics, 27, 1164-1165. |
| [11] | Degnan JH, Rosenberg NA (2009) Gene tree discordance, phylogenetic inference and the multispecies coalescent. Trends in Ecology & Evolution, 24, 332-340. |
| [12] | Delwiche CF, Palmer JD (1996) Rampant horizontal transfer and duplication of Rubisco genes in eubacteria and plastids. Molecular Biology and Evolution, 13, 873-882. |
| [13] | Doolittle WF (1999) Phylogenetic classification and the universal tree. Science, 284, 2124-2128. |
| [14] | Dunn CW, Hejnol A, Matus DQ, Pang K, Browne WE, Smith SA, Seaver E, Rouse GW, Obst M, Edgecombe GD (2008) Broad phylogenomic sampling improves resolution of the animal tree of life. Nature, 452, 745-749. |
| [15] | Dunn CW, Howison M, Zapata F (2013) Agalma: an automated phylogenomics workflow. BMC Bioinformatics, 14, 330. |
| [16] | Eames AJ (1936) Morphology of Vascular Plants. McGraw- Hill Book Company, New York. |
| [17] | Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Research, 32, 1792-1797. |
| [18] | Fritsch F (1945) Studies in the comparative morphology of the algae. IV. Algae and archegoniate plants. Annals of Botany, 9, 1-29. |
| [19] | Fritsch FE (1916) The algal ancestry of the higher plants. New Phytologist, 15, 233-249. |
| [20] | Fu L, Niu B, Zhu Z, Wu S, Li W (2012) CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics, 28, 3150-3152. |
| [21] | Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nature Biotechnology, 29, 644-652. |
| [22] | Griffiths RC, Marjoram P (1996) Ancestral inference from samples of DNA sequences with recombination. Journal of Computational Biology, 3, 479-502. |
| [23] | Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Molecular Biology and Evolution, 23, 254-267. |
| [24] | Huson DH, Scornavacca C (2011) A survey of combinatorial methods for phylogenetic networks. Genome Biology and Evolution, 3, 23-35. |
| [25] | Kenrick P, Crane PR (1997) The origin and early evolution of plants on land. Nature, 389, 33-39. |
| [26] | Li CS (1994) Origin of land plants is an important event of life evolution. Bulletin of National Natural Science Foundation of China, (4), 8-14. (in Chinese with English abstract) |
| [李承森 (1994) 生物进化的重大事件——陆地植物的起源及其研究的新进展. 中国科学基金, (4), 8-14.] | |
| [27] | Li L, Stoeckert CJ, Roos DS (2003) OrthoMCL: identification of ortholog groups for eukaryotic genomes. Genome Research, 13, 2178-2189. |
| [28] | Mallet J (2005) Hybridization as an invasion of the genome. Trends in Ecology & Evolution, 20, 229-237. |
| [29] | Mirarab S, Bayzid MS, Warnow T (2016) Evaluating summary methods for multilocus species tree estimation in the presence of incomplete lineage sorting. Systematic Biology, 65, 366-380. |
| [30] | Mirarab S, Reaz R, Bayzid MS, Zimmermann T, Swenson MS, Warnow T (2014) ASTRAL: genome-scale coalescent- based species tree estimation. Bioinformatics, 30, 541-548. |
| [31] | Misof B, Liu S, Meusemann K, Peters RS, Donath A, Mayer C, Frandsen PB, Ware J, Flouri T, Beutel RG (2014) Phylogenomics resolves the timing and pattern of insect evolution. Science, 346, 763-767. |
| [32] | Nakhleh L (2010) Evolutionary phylogenetic networks: models and issues. In: Problem Solving Handbook in Computational Biology and Bioinformatics (eds Lenwood SH, Naren R), pp. 125-158. Springer, New York. |
| [33] | Nakhleh L (2013) Computational approaches to species phylogeny inference and gene tree reconciliation. Trends in Ecology & Evolution, 28, 719-728. |
| [34] | Regier JC, Shultz JW, Zwick A, Hussey A, Ball B, Wetzer R, Martin JW, Cunningham CW (2010) Arthropod relationships revealed by phylogenomic analysis of nuclear protein-coding sequences. Nature, 463, 1079-1083. |
| [35] | Rieseberg LH (1997) Hybrid origins of plant species. Annual Review of Ecology and Systematics, 28, 359-389. |
| [36] | Rubinstein CV, Gerrienne P, de la Puente G, Astini R, Steemans P (2010) Early Middle Ordovician evidence for land plants in Argentina (eastern Gondwana). New Phytologist, 188, 365-369. |
| [37] | Salichos L, Rokas A (2013) Inferring ancient divergences requires genes with strong phylogenetic signals. Nature, 497, 327-331. |
| [38] | Scott DH (1900) Studies in Fossil Botany. Adam & Charles Black, London. |
| [39] | Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics, 31, 3210-3212. |
| [40] | Sneath PH (1975) Cladistic representation of reticulate evolution. Systematic Zoology, 24, 360-368. |
| [41] | Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics, 30, 1312-1313. |
| [42] | Steemans P, Le Hérissé A, Melvin J, Miller MA, Paris F, Verniers J, Wellman CH (2009) Origin and radiation of the earliest vascular land plants. Science, 324, 353. |
| [43] | Sukumaran J, Holder MT (2010) DendroPy: a Python library for phylogenetic computing. Bioinformatics, 26, 1569-1571. |
| [44] | Syvanen M (1985) Cross-species gene transfer, implications for a new theory of evolution. Journal of Theoretical Biology, 112, 333-343. |
| [45] | Szöllősi GJ, Boussau B, Abby SS, Tannier E, Daubin V (2012) Phylogenetic modeling of lateral gene transfer reconstructs the pattern and relative timing of speciations. Proceedings of the National Academy of Sciences, USA, 109, 17513-17518. |
| [46] | Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Systematic Biology, 56, 564-577. |
| [47] | Tofigh A, Hallett M, Lagergren J (2011) Simultaneous identification of duplications and lateral gene transfers. IEEE/ ACM Transactions on Computational Biology and Bioinformatics (TCBB), 8, 517-535. |
| [48] | Wellman CH, Osterloff PL, Mohiuddin U (2003) Fragments of the earliest land plants. Nature, 425, 282-285. |
| [49] | Wickett NJ, Mirarab S, Nguyen N, Warnow T, Carpenter E, Matasci N, Ayyampalayam S, Barker MS, Burleigh JG, Gitzendanner MA (2014) Phylotranscriptomic analysis of the origin and early diversification of land plants. Proceedings of the National Academy of Sciences, USA, 111, 4859-4868. |
| [50] | Woodhams MD, Lockhart PJ, Holland BR (2016) Simulating and summarizing sources of gene tree incongruence. Genome Biology and Evolution, 8, 1299-1315. |
| [51] | Zimmermann W (1938) Phylogenie. In: Manual of Pteridology (ed. Frans V), pp. 558-618. Springer, New York. |
| [52] | Zou XH, Ge S (2008) Conflicting gene trees and phylogenomics. Journal of Systematics and Evolution, 46, 795-807. (in Chinese with English abstract) |
| [邹新慧, 葛颂 (2008) 基因树冲突与系统发育基因组学研究. 植物分类学报, 46, 795-807.] |
| [1] | Wei Wang, Yang Liu. The current status, problems, and policy suggestions for reconstructing the plant tree of life [J]. Biodiv Sci, 2020, 28(2): 176-188. |
| [2] | Xinyu Du, Jinmei Lu, Dezhu Li. Advances in the evolution of plastid genome structure in lycophytes and ferns [J]. Biodiv Sci, 2019, 27(11): 1172-1183. |
| [3] | Liping Zeng,Ning Zhang,Hong Ma. Advances and challenges in resolving the angiosperm phylogeny [J]. Biodiv Sci, 2014, 22(1): 21-39. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2026 Biodiversity Science
Editorial Office of Biodiversity Science, 20 Nanxincun, Xiangshan, Beijing 100093, China
Tel: 010-62836137, 62836665 E-mail: biodiversity@ibcas.ac.cn