植物开花时间的遗传调控通路研究进展
- 1.北京师范大学生命科学学院, 生物多样性与生态工程教育部重点实验室, 北京 100875
2.北京师范大学, 生命科学与技术国家级实验教学示范中心, 北京 100875
收稿日期: 2020-09-21
录用日期: 2020-11-30
网络出版日期: 2021-02-01
基金资助
国家自然科学基金(31700326);国家自然科学基金(31770253)
Advances in the genetic regulating pathways of plant flowering time
- 1 MOE Key Laboratory for Biodiversity Science and Ecological Engineering, College of Life Sciences, Beijing Normal University, Beijing 100875
2 National Demonstration Center for Experimental Life Sciences and Biotechnology Education, Beijing Normal University, Beijing 100875
Received date: 2020-09-21
Accepted date: 2020-11-30
Online published: 2021-02-01
摘要
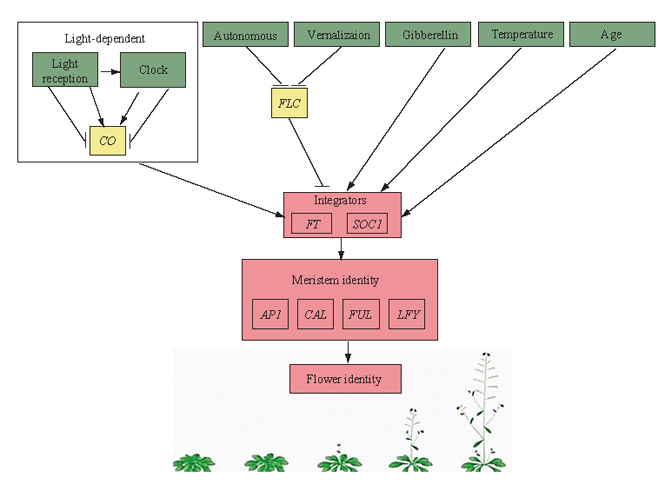
开花时间对植物的繁殖成功至关重要。广泛分布的物种经常发生开花时间的分化, 从而能够更好地适应不同的环境条件。为了探索植物开花行为发生适应性分化的分子机制, 首先要明确调控开花行为的遗传通路。本文梳理了植物各类群调控开花时间的遗传通路, 以期为开花时间适应性分化的分子机制研究提供依据。 植物从营养生长向繁殖转变时, 其开花行为主要受到光照、温度、水分等外界环境因子和赤霉素等内在因素的影响。通过对模式植物拟南芥(Arabidopsis thaliana)和其他类群的研究, 总结出了调控植物开花时间的6条通路, 包括日照长度和光质影响开花的光依赖通路, 长时间冷暴露后促进植物开花的春化通路, 高温或低温环境影响开花的温度通路, 以及赤霉素通路、年龄通路和自主通路3条内部调节过程。植物开花时间调控的6条上游通路信号传递到下游的开花整合基因FT(FLOWERING LOCUS T)和SOC1(SUPPRESSOR OF OVEREXPRESSION OF CONSTANS 1), 整合基因将这些复杂的调节因子整合后进一步传递到下游花分生组织, 从而启动开花。此外, 非编码RNA、转座子对开花时间的调控也具有重要作用。部分遗传通路被证实在植物适应环境的过程中起到了重要作用。目前对植物开花调控的研究已经有一百多年历史, 理论相对成熟。然而, 仍然存在许多具有争议和未解决的问题, 如开花基因的表达方式、开花行为的特殊调控机制、开花时间变异的适应性意义等等, 需要更进一步的研究。
本文引用格式
杨小凤 , 李小蒙 , 廖万金 . 植物开花时间的遗传调控通路研究进展[J]. 生物多样性, 2021 , 29(6) : 825 -842 . DOI: 10.17520/biods.2020370
Abstract
Aims: Flowering time is critical for plant reproductive success. Plant species, especially which are widely distributed, are observed to exhibit significant differentiation in flowering time to maximize their fitness. In order to explore the molecular mechanism of adaptive differentiation of flowering time in plants, it is essential to clarify the regulating pathways of flowering. This paper summarizes the genetic pathways that regulate flowering time in various groups of plants, which can provide important reference for molecular mechanism research of adaptive differentiation of flowering time.
Progresses: As a vital transition from vegetative growth to reproduction in plants, the flowering time has been reported to be regulated by both extrinsic and intrinsic factors. Based on researches of the model plant Arabidopsis thaliana and other taxa, six pathways that regulate flowering time have been identified. The light-dependent pathway, vernalization pathway, and temperature pathway refer to the regulation of flowering time in response to day length or quality of light, a long period of cold exposure, and external high or low temperature, respectively. While the gibberellin pathway, the age pathway and the autonomic pathway are the endogenous pathways. Signals from the six upstream flowering regulation pathways are transmitted to the downstream flowering integratorsFT (FLOWERING LOCUS T) andSOC1(SUPPRESSOR OF OVEREXPRESSION OF CONSTANS 1), assembled by these two integrators, and transmitted to the downstream flowering meristem where flowering initiates. In addition, most recent studies have shown that non-coding RNA and transposable elements also play important roles in the regulation of flowering time.
Prospect: The researches on the regulation of plant flowering have studied more than a hundred years ago, forming many mature theories. However, there are still many controversial and unresolved problems, such as the expression ways of flowering genes, the special regulatory mechanism of flowering, and the adaptive significance of differentiated flowering time, which require further study.

Key words: flowering gene; flowering time; fitness; genetic regulation; genetic pathway
参考文献
| [1] | Abe M, Kosaka S, Shibuta M, Nagata K, Uemura T, Nakano A, Kaya H (2019) Transient activity of the florigen complex during the floral transition in Arabidopsis thaliana . Development, 146, dev171504. |
| [2] | Abou-Elwafa SF, Büttner B, Chia T, Schulze-Buxloh G, Hohmann U, Mutasa-G?ttgens E, Jung C, Müller AE (2011) Conservation and divergence of autonomous pathway genes in the flowering regulatory network of Beta vulgaris . Journal of Experimental Botany, 62,3359-3374. |
| [3] | Achard P, Herr A, Baulcombe DC, Harberd NP (2004) Modulation of floral development by a gibberellin-regulated microRNA. Development, 131,3357-3365. |
| [4] | Alexandre CM, Hennig L (2008) FLC or not FLC: The other side of vernalization . Journal of Experimental Botany, 59,1127-1135. |
| [5] | Alonso-Blanco C, Aarts MGM, Bentsink L, Keurentjes JJB, Reymond M, Vreugdenhil D, Koornneef M (2009) What has natural variation taught us about plant development, physiology, and adaptation? The Plant Cell, 21,1877-1896. |
| [6] | Andrés F, Coupland G (2012) The genetic basis of flowering responses to seasonal cues. Nature Reviews Genetics, 13,627-639. |
| [7] | Andrés F, Porri A, Torti S, Mateos J, Romera-Branchat M, Garcia-Martinez JL, Fornara F, Gregis V, Kater MM, Coupland G (2014) SHORT VEGETATIVE PHASE reduces gibberellin biosynthesis at the Arabidopsis shoot apex to regulate the floral transition. Proceedings of the National Academy of Sciences, USA, 111,E2760-E2769. |
| [8] | Balasubramanian S, Sureshkumar S, Lempe J, Weigel D (2006) Potent induction of Arabidopsis thaliana flowering by elevated growth temperature . PLoS Genetics, 2,e106. |
| [9] | Banta S,and Heyland MD. (2011) The genetics and evolution of flowering time variation in plants: Identifying genes that control a key life history transition. In: Mechanisms of life history evolution (eds Flatt T, Heyland A), pp. 114-126. Oxford University Press Inc., New York. |
| [10] | Bao SJ, Hua CM, Huang GQ, Cheng P, Gong XM, Shen LS, Yu H (2019) Molecular basis of natural variation in photoperiodic flowering responses. Developmental Cell, 50,90-101. |
| [11] | Barrett SCH, Colautti RI, Eckert CG (2008) Plant reproductive systems and evolution during biological invasion. Molecular Ecology, 17,373-383. |
| [12] | B?urle I, Dean C (2008) Differential interactions of the autonomous pathway RRM proteins and chromatin regulators in the silencing of Arabidopsis targets . PLoS ONE, 3,e2733. |
| [13] | Bernler G, Havelange A, Houssa C, Petitjean A, Lejeune P (1993) Physiological signals that induce flowering. The Plant Cell, 5,1147-1155. |
| [14] | Busch MA, Bomblies K, Weigel D (1999) Activation of a floral homeotic gene in Arabidopsis . Science, 285,585-587. |
| [15] | Campos-Rivero G, Osorio-Montalvo P, Sánchez-Borges R, Us-Camas R, Duarte-Aké F, De-La-Pe?a C (2017) Plant hormone signaling in flowering: An epigenetic point of view. Journal of Plant Physiology, 214,16-27. |
| [16] | Cheng JZ, Zhou YP, Lv TX, Xie CP, Tian CE (2017) Research progress on the autonomous flowering time pathway in Arabidopsis . Physiology and Molecular Biology of Plants, 23,477-485. |
| [17] | Corbesier L, Vincent C, Jang S, Fornara F, Fan QZ, Searle I, Giakountis A, Farrona S, Gissot L, Turnbull C, Coupland G (2007) FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis . Science, 316,1030-1033. |
| [18] | de Lucas M, Davière JM, Rodríguez-Falcón M, Pontin M, Iglesias-Pedraz JM, Lorrain S, Fankhauser C, Blázquez MA, Titarenko E, Prat S (2008) A molecular framework for light and gibberellin control of cell elongation. Nature, 451,480-484. |
| [19] | Doi K, Izawa T, Fuse T, Yamanouchi U, Kubo T, Shimatani Z, Yano M, Yoshimura A (2004) Ehd1, a B-type response regulator in rice, confers short-day promotion of flowering and controls FT-like gene expression independently of Hd1. Genes & Development. 18,926-936. |
| [20] | Elzinga JA, Atlan A, Biere A, Gigord L, Weis AE, Bernasconi G (2007) Time after time: Flowering phenology and biotic interactions. Trends in Ecology & Evolution, 22,432-439. |
| [21] | Fernández V, Takahashi Y, Le Gourrierec J, Coupland G (2016) Photoperiodic and thermosensory pathways interact through CONSTANS to promote flowering at high temperature under short days. The Plant Journal, 86,426-440. |
| [22] | Fornara F, de Montaigu A, Coupland G (2010) SnapShot: Control of flowering in Arabidopsis . Cell, 141,550-550.e2. |
| [23] | Fornara F, Panigrahi KCS, Gissot L, Sauerbrunn N, Rühl M, Jarillo JA, Coupland G (2009) Arabidopsis DOF transcription factors act redundantly to reduce CONSTANS expression and are essential for a photoperiodic flowering response . Developmental Cell, 17,75-86. |
| [24] | Gendall AR, Levy YY, Wilson A, Dean C (2001) The VERNALIZATION 2 gene mediates the epigenetic regulation of vernalization in Arabidopsis . Cell, 107,525-535. |
| [25] | Golicz AA, Steinfort U, Arya H, Singh MB, Bhalla PL (2020) Analysis of the quinoa genome reveals conservation and divergence of the flowering pathways. Functional & Integrative Genomics, 20,245-258. |
| [26] | Gou JQ, Tang CR, Chen NC, Wang H, Debnath S, Sun L, Flanagan A, Tang YH, Jiang QZ, Allen RD, Wang ZY (2019) SPL7 and SPL8 represent a novel flowering regulation mechanism in switchgrass . New Phytologist, 222,1610-1623. |
| [27] | Guo TT, Mu Q, Wang JY, Vanous AE, Onogi A, Iwata H, Li XR, Yu JM (2020) Dynamic effects of interacting genes underlying rice flowering-time phenotypic plasticity and global adaptation. Genome Research, 30,673-683. |
| [28] | Halliday KJ, Salter MG, Thingnaes E, Whitelam GC (2003) Phytochrome control of flowering is temperature sensitive and correlates with expression of the floral integrator FT . The Plant Journal, 33,875-885. |
| [29] | Hayama R, Agashe B, Luley E, King R, Coupland G (2007) A circadian rhythm set by dusk determines the expression of FT homologs and the short-day photoperiodic flowering response in Pharbitis . The Plant Cell, 19,2988-3000. |
| [30] | Hayama R, Mizoguchi T, Coupland G (2018) Differential effects of light-to-dark transitions on phase setting in circadian expression among clock-controlled genes in Pharbitis nil . Plant Signaling & Behavior, 13,e1473686. |
| [31] | He J, Xu ML, Willmann MR, McCormick K, Hu TQ, Yang L, Starker CG, Voytas DF, Meyers BC, Poethig RS (2018) Threshold-dependent repression of SPL gene expression by miR156/miR157 controls vegetative phase change in Arabidopsis thaliana . PLoS Genetics, 14,e1007337. |
| [32] | Heidari B, Nemie-Feyissa D, Kangasj?rvi S, Lillo C (2013) Antagonistic regulation of flowering time through distinct regulatory subunits of protein phosphatase 2A. PLoS ONE, 8,e67987. |
| [33] | Hemming MN, Walford SA, Fieg S, Dennis ES, Trevaskis B (2012) Identification of high-temperature-responsive genes in cereals. Plant Physiology, 158,1439-1450. |
| [34] | Heo JB, Sung S (2011) Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science, 331,76-79. |
| [35] | Hirakawa H, Sumitomo K, Hisamatsu T, Nagano S, Shirasawa K, Higuchi Y, Kusaba M, Koshioka M, Nakano Y, Yagi M, Yamaguchi H, Taniguchi K, Nakano M, Isobe SN (2019) De novo whole-genome assembly in Chrysanthemum seticuspe, a model species of Chrysanthemums, and its application to genetic and gene discovery analysis . DNA Research, 26,195-203. |
| [36] | Huang FY, Liu TK, Tang J, Duan WK, Hou XL (2019) BcMAF2 activates BcTEM1 and represses flowering in Pak-choi (Brassica rapa ssp. chinensis). Plant Molecular Biology, 100,19-32. |
| [37] | Hyun Y, Richter R, Coupland G (2017) Competence to flower: Age-controlled sensitivity to environmental cues. Plant Physiology, 173,36-46. |
| [38] | Hyun Y, Richter R, Vincent C, Martinez-Gallegos R, Porri A, Coupland G (2016) Multi-layered regulation of SPL15 and cooperation with SOC1integrate endogenous flowering pathways at the Arabidopsis shoot meristem . Developmental Cell, 37,254-266. |
| [39] | Imaizumi T, Schultz TF, Harmon FG, Ho LA, Kay SA (2005) FKF1 F-box protein mediates cyclic degradation of a repressor of CONSTANS in Arabidopsis. Science, 309,293-297. |
| [40] | Ito S, Song YH, Josephson-Day AR, Miller RJ, Breton G, Olmstead RG, Imaizumi T (2012) FLOWERING BHLH transcriptional activators control expression of the photoperiodic flowering regulator CONSTANS in Arabidopsis. Proceedings of the National Academy of Sciences, USA. 109,3582-3587. |
| [41] | Itoh H, Nonoue Y, Yano M, Izawa T (2010) A pair of floral regulators sets critical day length for Hd3a florigen expression in rice . Nature Genetics, 42,635-638. |
| [42] | Jacobsen SE, Olszewski NE, Meyerowitz EM (1998) SPINDLY's role in the gibberellin response pathway. Symposia of the Society for Experimental Biology, 51,73-78. |
| [43] | Jagadish SVK, Bahuguna RN, Djanaguiraman M, Gamuyao R, Prasad PVV, Craufurd PQ (2016) Implications of high temperature and elevated CO 2 on flowering time in plants . Frontiers in Plant Science, 7,913. |
| [44] | Jang S, Marchal V, Panigrahi KCS, Wenkel S, Soppe W, Deng XW, Valverde F, Coupland G (2008) Arabidopsis COP1 shapes the temporal pattern of CO accumulation conferring a photoperiodic flowering response . The EMBO Journal, 27,1277-1288. |
| [45] | Johanson U, West J, Lister C, Michaels S, Amasino R, Dean C (2000) Molecular analysis of FRIGIDA, a major determinant of natural variation in Arabidopsis flowering time . Science, 290,344-347. |
| [46] | Kim JJ, Lee JH, Kim W, Jung HS, Huijser P, Ahn JH (2012) The microRNA156-SQUAMOSA PROMOTER BINDING PROTEIN-LIKE3 module regulates ambient temperature-responsive flowering via FLOWERING LOCUS T in Arabidopsis . Plant Physiology, 159,461-478. |
| [47] | Kim MY, Kang YJ, Lee T, Lee SH (2013) Divergence of flowering-related genes in three legume species. The Plant Genome, 6, plantgenome2013. 03.0008. |
| [48] | Kim S, Soltis PS, Wall K, Soltis DE (2006) Phylogeny and domain evolution in the APETALA2-like gene family . Molecular Biology and Evolution, 23,107-120. |
| [49] | Kim W, Ahn HJ, Chiou TJ, Ahn JH (2011) The role of the miR399-PHO2 module in the regulation of flowering time in response to different ambient temperatures in Arabidopsis thaliana . Molecules and Cells, 32,83-88. |
| [50] | Kinmonth-Schultz HA, Tong XR, Lee J, Song YH, Ito S, Kim SH, Imaizumi T (2016) Cool night-time temperatures induce the expression of CONSTANS and FLOWERING LOCUS T to regulate flowering in Arabidopsis . New Phytologist, 211,208-224. |
| [51] | Kinoshita A, Richter R (2020) Genetic and molecular basis of floral induction in Arabidopsis thaliana . Journal of Experimental Botany, 71,2490-2504. |
| [52] | Kitamoto N, Ueno S, Takenaka A, Tsumura Y, Washitani I, Ohsawa R (2006) Effect of flowering phenology on pollen flow distance and the consequences for spatial genetic structure within a population of Primula sieboldii (Primulaceae). American Journal of Botany, 93,226-233. |
| [53] | Kobayashi Y, Weigel D (2007) Move on up, it's time for change—Mobile signals controlling photoperiod-dependent flowering. Genes & Development, 21,2371-2384. |
| [54] | Kojima S, Takahashi Y, Kobayashi Y, Monna L, Sasaki T, Araki T, Yano M (2002) Hd3a, a rice ortholog of the Arabidopsis FT gene, promotes transition to flowering downstream of Hd1under short-day conditions . Plant and Cell Physiology, 43,1096-1105. |
| [55] | Komiya R, Ikegami A, Tamaki S, Yokoi S, Shimamoto K (2008) Hd3a and RFT1 are essential for flowering in rice . Development, 135,767-774. |
| [56] | Kralemann LEM, Scalone R, Andersson L, Hennig L (2018) North European invasion by common ragweed is associated with early flowering and dominant changes in FT/TFL1expression . Journal of Experimental Botany, 69,2647-2658. |
| [57] | Kubota A, Ito S, Shim JS, Johnson RS, Song YH, Breton G, Goralogia GS, Kwon MS, Laboy Cintrón D, Koyama T, Ohme-Takagi M, Pruneda-Paz JL, Kay SA, MacCoss MJ, Imaizumi T (2017) TCP4-dependent induction of CONSTANS transcription requires GIGANTEA in photoperiodic flowering in Arabidopsis . PLoS Genetics. 13,e1006856. |
| [58] | Kumar SV, Lucyshyn D, Jaeger KE, Alós E, Alvey E, Harberd NP, Wigge PA (2012) Transcription factor PIF4 controls the thermosensory activation of flowering . Nature, 484,242-245. |
| [59] | Lazaro A, Mouriz A, Pi?eiro M, Jarillo JA (2015) Red light-mediated degradation of CONSTANS by the E 3 ubiquitin ligase HOS1 regulates photoperiodic flowering in Arabidopsis . The Plant Cell, 27,2437-2454. |
| [60] | Lee BD, Kim MR, Kang MY, Cha JY, Han SH, Nawkar GM, Sakuraba Y, Lee SY, Imaizumi T, McClung CR, Kim WY, Paek NC (2017) The F-box protein FKF1 inhibits dimerization of COP1in the control of photoperiodic flowering . Nature Communications, 8,2259. |
| [61] | Lee H, Yoo SJ, Lee JH, Kim W, Yoo SK, Fitzgerald H, Carrington JC, Ahn JH (2010) Genetic framework for flowering-time regulation by ambient temperature-responsive miRNAs in Arabidopsis . Nucleic Acids Research, 38,3081-3093. |
| [62] | Leeggangers HACF, Rosilio-Brami T, Bigas-Nadal J, Rubin N, van Dijk ADJ, Gonzalez FFNC, Saadon-Shitrit S, Nijveen H, Hilhorst HWM, Immink RGH, Zaccai M (2018) Tulipa gesneriana and Lilium longiflorum PEBP genes and their putative roles in flowering time control . Plant and Cell Physiology, 59,90-106. |
| [63] | Lemoine NP, Doublet D, Salminen JP, Burkepile DE, Parker JD (2017) Responses of plant phenology, growth, defense, and reproduction to interactive effects of warming and insect herbivory. Ecology, 98,1817-1828. |
| [64] | Li C, Li YH, Li YF, Lu HF, Hong HL, Tian Y, Li HY, Zhao T, Zhou XW, Liu J, Zhou XN, Jackson SA, Liu B, Qiu LJ (2020) A domestication-associated gene GmPRR3b regulates the circadian clock and flowering time in soybean . Molecular Plant, 13,745-759. |
| [65] | Li D, Liu C, Shen LS, Wu Y, Chen HY, Robertson M, Helliwell CA, Ito T, Meyerowitz E, Yu H (2008) A repressor complex governs the integration of flowering signals in Arabidopsis . Developmental Cell, 15,110-120. |
| [66] | Li MZ, An FY, Li WY, Ma MD, Feng Y, Zhang X, Guo HW (2016) DELLA proteins interact with FLC to repress flowering transition . Journal of Integrative Plant Biology, 58,642-655. |
| [67] | Li XF, Jia LY, Xu J, Deng XJ, Wang Y, Zhang W, Zhang XP, Fang Q, Zhang DM, Sun Y, Xu L (2013) FT-like NFT1 gene may play a role in flower transition induced by heat accumulation in Narcissus tazetta var. chinensis . Plant and Cell Physiology, 54,270-281. |
| [68] | Li XM, Zhang DY, Liao WJ (2015) The rhythmic expression of genes controlling flowering time in southern and northern populations of invasive Ambrosia artemisiifolia . Journal of Plant Ecology, 8,207-212. |
| [69] | Li ZC, Jiang DH, He YH (2018) FRIGIDA establishes a local chromosomal environment for FLOWERING LOCUS C mRNA production . Nature Plants, 4,836-846. |
| [70] | Liljegren SJ, Gustafson-Brown C, Pinyopich A, Ditta GS, Yanofsky MF (1999) Interactions among APETALA1, LEAFY, and TERMINAL FLOWER1 specify meristem fate . The Plant Cell, 11,1007-1018. |
| [71] | Liu FQ, Quesada V, Crevillén P, B?urle I, Swiezewski S, Dean C (2007) The Arabidopsis RNA-binding protein FCA requires a lysine-specific demethylase 1 homolog to downregulate FLC . Molecular Cell, 28,398-407. |
| [72] | Liu KD, Feng SX, Pan YL, Zhong JD, Chen Y, Yuan CC, Li HL (2016) Transcriptome analysis and identification of genes associated with floral transition and flower development in sugar apple ( Annona squamosa L.) . Frontiers in Plant Science, 7,1695. |
| [73] | Liu KP, Yu Y, Dong AW, Shen WH (2017) SET DOMAIN GROUP701encodes a H3K4-methytransferase and regulates multiple key processes of rice plant development . New Phytologist, 215,609-623. |
| [74] | Liu L, Liu C, Hou XL, Xi WY, Shen LS, Tao Z, Wang Y, Yu H (2012) FTIP1is an essential regulator required for florigen transport . PLoS Biology, 10,e1001313. |
| [75] | Liu Y, Hao XY, Lu QH, Zhang WF, Zhang HJ, Wang L, Yang YJ, Xiao B, Wang XC (2020) Genome-wide identification and expression analysis of flowering-related genes reveal putative floral induction and differentiation mechanisms in tea plant (Camellia sinensis). Genomics, 112,2318-2326. |
| [76] | Liu YY, Yang KZ, Wei XX, Wang XQ (2016) Revisiting the phosphatidylethanolamine-binding protein (PEBP) gene family reveals cryptic FLOWERING LOCUS T gene homologs in gymnosperms and sheds new light on functional evolution . New Phytologist, 212,730-744. |
| [77] | Luo M, Tai R, Yu CW, Yang SG, Chen CY, Lin WD, Schmidt W, Wu KQ (2015) Regulation of flowering time by the histone deacetylase HDA5 in Arabidopsis . The Plant Journal, 82,925-936. |
| [78] | Mathieu J, Yant LJ, Mürdter F, Küttner F, Schmid M (2009) Repression of flowering by the miR172 target SMZ . PLoS Biology, 7,e1000148. |
| [79] | Mátyás KK, Heged?s G, Taller J, Farkas E, Decsi K, Kutasy B, Kálmán N, Nagy E, Kolics B, Virág E (2019) Different expression pattern of flowering pathway genes contribute to male or female organ development during floral transition in the monoecious weed Ambrosia artemisiifolia L. (Asteraceae). PeerJ, 7,e7421. |
| [80] | Mawphlang OIL, Kharshiing EV (2017) Photoreceptor mediated plant growth responses: Implications for photoreceptor engineering toward improved performance in crops. Frontiers in Plant Science, 8,1181. |
| [81] | Méndez-Vigo B, Ausín I, Zhu WS, Mollá-Morales A, Balasubramanian S, Alonso-Blanco C (2019) Genetic interactions and molecular evolution of the duplicated genes ICARUS2 and ICARUS1 help Arabidopsis plants adapt to different ambient temperatures . The Plant Cell, 31,1222-1237. |
| [82] | Méndez-Vigo B, Martínez-Zapater JM, Alonso-Blanco C (2013) The flowering repressor SVP underlies a novel Arabidopsis thaliana QTL interacting with the genetic background . PLoS Genetics, 9,e1003289. |
| [83] | Michaels SD, Amasino RM (2001) Loss of FLOWERING LOCUS C activity eliminates the late-flowering phenotype of FRIGIDA and autonomous pathway mutations but not responsiveness to vernalization . The Plant Cell, 13,935-941. |
| [84] | Michaels SD, Ditta G, Gustafson-Brown C, Pelaz S, Yanofsky M, Amasino RM (2003) AGL24 acts as a promoter of flowering in Arabidopsis and is positively regulated by vernalization . The Plant Journal, 33,867-874. |
| [85] | Mitchum MG, Yamaguchi S, Hanada A, Kuwahara A, Yoshioka Y, Kato T, Tabata S, Kamiya Y, Sun TP (2006) Distinct and overlapping roles of two gibberellin 3-oxidases in Arabidopsis development . The Plant Journal, 45,804-818. |
| [86] | Moon J, Suh SS, Lee H, Choi KR, Hong CB, Paek NC, Kim SG, Lee I (2003) The SOC1 MADS-box gene integrates vernalization and gibberellin signals for flowering in Arabidopsis . The Plant Journal, 35,613-623. |
| [87] | Mulekar JJ, Huq E (2015) Arabidopsis casein kinase 2 α 4 subunit regulates various developmental pathways in a functionally overlapping manner . Plant Science, 236,295-303. |
| [88] | Mutasa-G?ttgens E, Hedden P (2009) Gibberellin as a factor in floral regulatory networks. Journal of Experimental Botany, 60,1979-1989. |
| [89] | Nie SS, Li C, Xu L, Wang Y, Huang DQ, Muleke EM, Sun XC, Xie Y, Liu LW (2016) De novo transcriptome analysis in radish ( Raphanus sativus L.) and identification of critical genes involved in bolting and flowering . BMC Genomics, 17,389. |
| [90] | Niu LF, Lu FL, Pei YX, Liu CY, Cao XF (2007) Regulation of flowering time by the protein arginine methyltransferase AtPRMT10 . EMBO Reports, 8,1190-1195. |
| [91] | Niu XM, Xu YC, Li ZW, Bian YT, Hou XH, Chen JF, Zou YP, Jiang J, Wu Q, Ge S, Balasubramanian S, Guo YL (2019) Transposable elements drive rapid phenotypic variation in Capsella rubella. Proceedings of the National Academy of Sciences, USA, 116,6908-6913. |
| [92] | Oda A, Higuchi Y, Hisamatsu T (2020) Constitutive expression of CsGI alters critical night length for flowering by changing the photo-sensitive phase of anti-florigen induction in Chrysanthemum . Plant Science, 293,110417. |
| [93] | Perrella G, Vellutini E, Zioutopoulou A, Patitaki E, Headland LR, Kaiserli E (2020) Let it bloom: Cross-talk between light and flowering signaling in Arabidopsis . Physiologia Plantarum, 169,301-311. |
| [94] | Pin PA, Benlloch R, Bonnet D, Wremerth-Weich E, Kraft T, Gielen JJL, Nilsson O (2010) An antagonistic pair of FT homologs mediates the control of flowering time in sugar beet . Science, 330,1397-1400. |
| [95] | Pin PA, Zhang WY, Vogt SH, Dally N, Büttner B, Schulze-Buxloh G, Jelly NS, Chia TYP, Mutasa-G?ttgens ES, Dohm JC, Himmelbauer H, Weisshaar B, Kraus J, Gielen JJL, Lommel M, Weyens G, Wahl B, Schechert A, Müller AE (2012) The role of a pseudo-response regulator gene in life cycle adaptation and domestication of beet. Current Biology, 22,1095-1101. |
| [96] | Porri A, Torti S, Romera-Branchat M, Coupland G (2012) Spatially distinct regulatory roles for gibberellins in the promotion of flowering of Arabidopsis under long photoperiods . Development, 139,2198-2209. |
| [97] | Posé D, Verhage L, Ott F, Yant L, Mathieu J, Angenent GC, Immink RGH, Schmid M (2013) Temperature-dependent regulation of flowering by antagonistic FLM variants. Nature, 503,414-417. |
| [98] | Putterill J, Laurie R, MacKnight R (2004) It's time to flower: The genetic control of flowering time. BioEssays, 26,363-373. |
| [99] | Reeves PA, He YH, Schmitz RJ, Amasino RM, Panella LW, Richards CM (2007) Evolutionary conservation of the FLOWERING LOCUS C-mediated vernalization response: Evidence from the sugar beet (Beta vulgaris). Genetics, 176,295-307. |
| [100] | Ripoll JJ, Rodríguez-Cazorla E, González-Reig S, Andújar A, Alonso-Cantabrana H, Perez-Amador MA, Carbonell J, Martínez-Laborda A, Vera A (2009) Antagonistic interactions between Arabidopsis K-homology domain genes uncover PEPPER as a positive regulator of the central floral repressor FLOWERING LOCUS C . Developmental Biology, 333,251-262. |
| [101] | Rosloski SM, Jali SS, Balasubramanian S, Weigel D, Grbic V (2010) Natural diversity in flowering responses of Arabidopsis thaliana caused by variation in a tandem gene array . Genetics, 186,263-276. |
| [102] | Ruelens P, de Maagd RA, Proost S, Thei?en G, Geuten K, Kaufmann K (2013) FLOWERING LOCUS C in monocots and the tandem origin of angiosperm-specific MADS-box genes . Nature Communications, 4,2280. |
| [103] | Sanchez SE, Rugnone ML, Kay SA (2020) Light perception: A matter of time. Molecular Plant, 13,363-385. |
| [104] | Sasani S, Hemming MN, Oliver SN, Greenup A, Tavakkol-Afshari R, Mahfoozi S, Poustini K, Sharifi HR, Dennis ES, Peacock WJ, Trevaskis B (2009) The influence of vernalization and daylength on expression of flowering-time genes in the shoot apex and leaves of barley(Hordeum vulgare). Journal of Experimental Botany, 60,2169-2178. |
| [105] | Sawa M, Kay SA (2011) GIGANTEA directly activates Flowering Locus T in Arabidopsis thaliana. Proceedings of the National Academy of Sciences, USA, 108,11698-11703. |
| [106] | Sawa M, Nusinow DA, Kay SA, Imaizumi T (2007) FKF1 and GIGANTEA complex formation is required for day-length measurement in Arabidopsis. Science, 318,261-265. |
| [107] | Schiessl SV, Huettel B, Kuehn D, Reinhardt R, Snowdon RJ (2017) Flowering time gene variation in Brassica species shows evolutionary principles . Frontiers in Plant Science, 8,1742. |
| [108] | Shaw LM, Li CX, Woods DP, Alvarez MA, Lin HQ, Lau MY, Chen A, Dubcovsky J (2020) Epistatic interactions between PHOTOPERIOD1, CONSTANS1and CONSTANS2 modulate the photoperiodic response in wheat . PLoS Genetics, 16,e1008812. |
| [109] | Sheldon CC, Rouse DT, Finnegan EJ, Peacock WJ, Dennis ES (2000) The molecular basis of vernalization: The central role of FLOWERING LOCUS C (FLC) . Proceedings of the National Academy of Sciences, USA, 97,3753-3758. |
| [110] | Shibaya T, Hori K, Ogiso-Tanaka E, Yamanouchi U, Shu K, Kitazawa N, Shomura A, Ando T, Ebana K, Wu JZ, Yamazaki T, Yano M (2016) Hd18, encoding histone acetylase related to Arabidopsis FLOWERING LOCUS D, is involved in the control of flowering time in rice . Plant and Cell Physiology, 57,1828-1838. |
| [111] | Shim JS, Jang G (2020) Environmental signal-dependent regulation of flowering time in rice. International Journal of Molecular Sciences, 21,6155. |
| [112] | Shim JS, Kubota A, Imaizumi T (2017) Circadian clock and photoperiodic flowering in Arabidopsis: CONSTANS is a hub for signal integration . Plant Physiology, 173,5-15. |
| [113] | Shindo C, Aranzana MJ, Lister C, Baxter C, Nicholls C, Nordborg M, Dean C (2005) Role of FRIGIDA and FLOWERING LOCUS C in determining variation in flowering time of Arabidopsis . Plant Physiology, 138,1163-1173. |
| [114] | Simpson GG (2004) The autonomous pathway: Epigenetic and post-transcriptional gene regulation in the control of Arabidopsis flowering time . Current Opinion in Plant Biology, 7,570-574. |
| [115] | Simpson GG, Dean C (2002) Arabidopsis, the Rosetta stone of flowering time? Science, 296,285-289. |
| [116] | Song YH, Estrada DA, Johnson RS, Kim SK, Lee SY, MacCoss MJ, Imaizumi T (2014) Distinct roles of FKF1, GIGANTEA, and ZEITLUPE proteins in the regulation of CONSTANS stability in Arabidopsis photoperiodic flowering. Proceedings of the National Academy of Sciences, USA, 111,17672-17677. |
| [117] | Song YH, Ito S, Imaizumi T (2013) Flowering time regulation: Photoperiod- and temperature-sensing in leaves. Trends in Plant Science, 18,575-583. |
| [118] | Spanudakis E, Jackson S (2014) The role of microRNAs in the control of flowering time. Journal of Experimental Botany, 65,365-380. |
| [119] | Srikanth A, Schmid M (2011) Regulation of flowering time: All roads lead to Rome. Cellular and Molecular Life Sciences, 68,2013-2037. |
| [120] | Steffen A, Elgner M, Staiger D (2019) Regulation of flowering time by the RNA-binding proteins AtGRP7 and AtGRP8. Plant and Cell Physiology, 60,2040-2050. |
| [121] | Sugano S, Andronis C, Ong MS, Green RM, Tobin EM (1999) The protein kinase CK2 is involved in regulation of circadian rhythms in Arabidopsis. Proceedings of the National Academy of Sciences, USA, 96,12362-12366. |
| [122] | Sun H, Zhang WP, Wu YZ, Gao LF, Cui F, Zhao CH, Guo ZA, Jia JZ (2020) The circadian clock gene, TaPRR1, is associated with yield-related traits in wheat ( Triticum aestivum L.) . Frontiers in Plant Science, 11,285. |
| [123] | Sun TP (2010) Gibberellin- GID1- DELLA: A pivotal regulatory module for plant growth and development . Plant Physiology, 154,567-570. |
| [124] | Sung S, Amasino RM (2004) Vernalization in Arabidopsis thaliana is mediated by the PHD finger protein VIN3 . Nature, 427,159-164. |
| [125] | Swiezewski S, Liu FQ, Magusin A, Dean C (2009) Cold-induced silencing by long antisense transcripts of an Arabidopsis polycomb target . Nature, 462,799-802. |
| [126] | Takada S, Goto K (2003) TERMINAL FLOWER2, an Arabidopsis homolog of HETEROCHROMATIN PROTEIN1, counteracts the activation of FLOWERING LOCUS T by CONSTANS in the vascular tissues of leaves to regulate flowering time . The Plant Cell, 15,2856-2865. |
| [127] | Tamaki S, Tsuji H, Matsumoto A, Fujita A, Shimatani Z, Terada R, Sakamoto T, Kurata T, Shimamoto K (2015) FT-like proteins induce transposon silencing in the shoot apex during floral induction in rice. Proceedings of the National Academy of Sciences, USA, 112,E901-E910. |
| [128] | Turner A, Beales J, Faure S, Dunford RP, Laurie DA (2005) The pseudo-response regulator Ppd-H1 provides adaptation to photoperiod in barley . Science, 310,1031-1034. |
| [129] | Valverde F, Mouradov A, Soppe W, Ravenscroft D, Samach A, Coupland G (2004) Photoreceptor regulation of CONSTANS protein in photoperiodic flowering . Science, 303,1003-1006. |
| [130] | Verhage L, Angenent GC, Immink RGH (2014) Research on floral timing by ambient temperature comes into blossom. Trends in Plant Science, 19,583-591. |
| [131] | Vermeulen PJ (2015) On selection for flowering time plasticity in response to density. New Phytologist, 205,429-439. |
| [132] | Wang FF, Nan HY, Chen LY, Fang C, Zhang HY, Su T, Li SC, Cheng Q, Dong LD, Liu BH, Kong FJ, Lu SJ (2019) A new dominant locus, E11, controls early flowering time and maturity in soybean. Molecular Breeding, 39,70. |
| [133] | Wang HP, Pan JJ, Li Y, Lou DJ, Hu YR, Yu DQ (2016) The DELLA-CONSTANS transcription factor cascade integrates gibberellic acid and photoperiod signaling to regulate flowering . Plant Physiology, 172,479-488. |
| [134] | Wang LW, Sun S, Wu TT, Liu LP, Sun XG, Cai YP, Li JC, Jia HC, Yuan S, Chen L, Jiang BJ, Wu CX, Hou WS, Han TF (2020) Natural variation and CRISPR/Cas9-mediated mutation in GmPRR37 affect photoperiodic flowering and contribute to regional adaptation of soybean . Plant Biotechnology Journal, 18,1869-1881. |
| [135] | Wang P, Gong R, Yang Y, Yu SB (2019) Ghd8 controls rice photoperiod sensitivity by forming a complex that interacts with Ghd7 . BMC Plant Biology, 19,462. |
| [136] | Wang SL, Gao J, Xue JQ, Xue YQ, Li DD, Guan YR, Zhang XX (2019) De novo sequencing of tree peony ( Paeonia suffruticosa) transcriptome to identify critical genes involved in flowering and floral organ development . BMC Genomics, 20,572. |
| [137] | Wang YY, Xiong F, Ren QP, Wang XL (2020) Regulation of flowering transition by alternative splicing: The role of the U 2 auxiliary factor. Journal of Experimental Botany, 71,751-758. |
| [138] | Wei JH, Choi H, Jin P, Wu YF, Yoon J, Lee YS, Quan TY, An G (2016) GL2-type homeobox gene Roc4 in rice promotes flowering time preferentially under long days by repressing Ghd7 . Plant Science, 252,133-143. |
| [139] | Weigel D (2012) Natural variation in Arabidopsis: From molecular genetics to ecological genomics . Plant Physiology, 158,2-22. |
| [140] | Xia W, Liu R, Zhang J, Mason AS, Li ZY, Gong SF, Zhong YZ, Dou YJ, Sun XW, Fan HK, Xiao Y (2020) Alternative splicing of flowering time gene FT is associated with halving of time to flowering in coconut . Scientific Reports, 10,11640. |
| [141] | Xing DH, Zhao HW, Xu RQ, Li QQ (2008) Arabidopsis PCFS4, a homologue of yeast polyadenylation factor Pcf11p, regulates FCA alternative processing and promotes flowering time . The Plant Journal, 54,899-910. |
| [142] | Xu F, Li T, Xu PB, Li L, Du SS, Lian HL, Yang HQ (2016) DELLA proteins physically interact with CONSTANS to regulate flowering under long days in Arabidopsis . FEBS Letters, 590,541-549. |
| [143] | Xu ML, Hu TQ, Zhao JF, Park MY, Earley KW, Wu G, Yang L, Poethig RS (2016) Developmental functions of miR156-regulated SQUAMOSA PROMOTER BINDING PROTEIN-LIKE ( SPL) genes in Arabidopsis thaliana . PLoS Genetics, 12,e1006263. |
| [144] | Yadav A, Jayaswal PK, Venkat Raman K, Singh B, Singh NK, Usha K (2020) Transcriptome analysis of flowering genes in mango ( Mangifera indica L.) in relation to floral malformation . Journal of Plant Biochemistry and Biotechnology, 29,193-212. |
| [145] | Yamaguchi A, Abe M (2012) Regulation of reproductive development by non-coding RNA in Arabidopsis: To flower or not to flower . Journal of Plant Research, 125,693-704. |
| [146] | Yan L, Fu D, Li C, Blechl A, Tranquilli G, Bonafede M, Sanchez A, Valarik M, Yasuda S, Dubcovsky J (2006) The wheat and barley vernalization gene VRN3 is an orthologue of FT. Proceedings of the National Academy of Sciences, USA, 103,19581-19586. |
| [147] | Yan L, Loukoianov A, Tranquilli G, Helguera M, Fahima T, Dubcovsky J (2003) Positional cloning of the wheat vernalization gene VRN1. Proceedings of the National Academy of Sciences, USA, 100,6263-6268. |
| [148] | Yan LL, Loukoianov A, Blechl A, Tranquilli G, Ramakrishna W, SanMiguel P, Bennetzen JL, Echenique V, Dubcovsky J (2004) The wheat VRN2 gene is a flowering repressor down-regulated by vernalization . Science, 303,1640-1644. |
| [149] | Yan YY, Shen LS, Chen Y, Bao SJ, Thong ZH, Yu H (2014) A MYB-domain protein EFM mediates flowering responses to environmental cues in Arabidopsis . Developmental Cell, 30,437-448. |
| [150] | Yang FX, Zhu GF, Wei YL, Gao J, Liang G, Peng LY, Lu CQ, Jin JP (2019) Low-temperature-induced changes in the transcriptome reveal a major role of CgSVP genes in regulating flowering of Cymbidium goeringii . BMC Genomics, 20,53. |
| [151] | Yang L, Wang HN, Hou XH, Zou YP, Han TS, Niu XM, Zhang J, Zhao Z, Todesco M, Balasubramanian S, Guo YL (2018) Parallel evolution of common allelic variants confers flowering diversity in Capsella rubella . The Plant Cell, 30,1322-1336. |
| [152] | Yano K, Aya K, Hirano K, Ordonio RL, Ueguchi-Tanaka M, Matsuoka M (2015) Comprehensive gene expression analysis of rice aleurone cells: Probing the existence of an alternative gibberellin receptor. Plant Physiology, 167,531-544. |
| [153] | Yano M, Katayose Y, Ashikari M, Yamanouchi U, Monna L, Fuse T, Baba T, Yamamoto K, Umehara Y, Nagamura Y, Sasaki T (2000) Hd1, a major photoperiod sensitivity quantitative trait locus in rice, is closely related to the Arabidopsis flowering time gene CONSTANS. The Plant Cell, 12,2473-2484. |
| [154] | Yanovsky MJ, Kay SA (2001) Signaling networks in the plant circadian system. Current Opinion in Plant Biology, 4,429-435. |
| [155] | Yi H, Li XN, Lee SH, Nou IS, Lim YP, Hur Y (2017) Natural variation in CIRCADIAN CLOCK ASSOCIATED 1 is associated with flowering time in Brassica rapa . Genome, 60,402-413. |
| [156] | Yun H, Hyun Y, Kang MJ, Noh YS, Noh B, Choi Y (2011) Identification of regulators required for the reactivation of FLOWERING LOCUS C during Arabidopsis reproduction . Planta, 234,1237-1250. |
| [157] | Zentella R, Sui N, Barnhill B, Hsieh WP, Hu JH, Shabanowitz J, Boyce M, Olszewski NE, Zhou P, Hunt DF, Sun TP (2017) The Arabidopsis O-fucosyltransferase SPINDLY activates nuclear growth repressor DELLA . Nature Chemical Biology, 13,479-485. |
| [158] | Zhang L, Jiang YY, Zhu YM, Su WB, Long T, Huang TQ, Peng JR, Yu H, Lin SQ, Gao YS (2019) Functional characterization of GI and CO homologs from Eriobotrya deflexa Nakai forma koshunensis . Plant Cell Reports, 38,533-543. |
| [159] | Zhang SW, Gottschalk C, Nocker S (2019) Genetic mechanisms in the repression of flowering by gibberellins in apple ( Malus × domestica Borkh.) . BMC Genomics, 20,747. |
| [160] | Zhao ML, Ni J, Chen MS, Xu ZF (2019) Ectopic expression of Jatropha curcas TREHALOSE-6-PHOSPHATE PHOSPHATASE J causes late-flowering and heterostylous phenotypes in Arabidopsis but not in Jatropha . International Journal of Molecular Sciences, 20,2165. |
| [161] | Zhong XH, Du JM, Hale CJ, Gallego-Bartolome J, Feng SH, Vashisht AA, Chory J, Wohlschlegel JA, Patel DJ, Jacobsen SE (2014) Molecular mechanism of action of plant DRM de novo DNA methyltransferases. Cell, 157,1050-1060. |
| [162] | Zhu QH, Helliwell CA (2011) Regulation of flowering time and floral patterning by miR172 . Journal of Experimental Botany, 62,487-495. |
| [163] | Zhu Y, Liu L, Shen LS, Yu H (2016) NaKR1 regulates long-distance movement of FLOWERING LOCUS T in Arabidopsis . Nature Plants, 2,16075. |
| [164] | Zicola J, Liu LY, T?nzler P, Turck F (2019) Targeted DNA methylation represses two enhancers of FLOWERING LOCUS T in Arabidopsis thaliana . Nature Plants, 5,300-307. |
| [165] | Zuo ZC, Liu HT, Liu B, Liu XM, Lin CT (2011) Blue light-dependent interaction of CRY2with SPA1 regulates COP1 activity and floral initiation in Arabidopsis . Current Biology, 21,841-847. |